Trunk vs side branch rate models
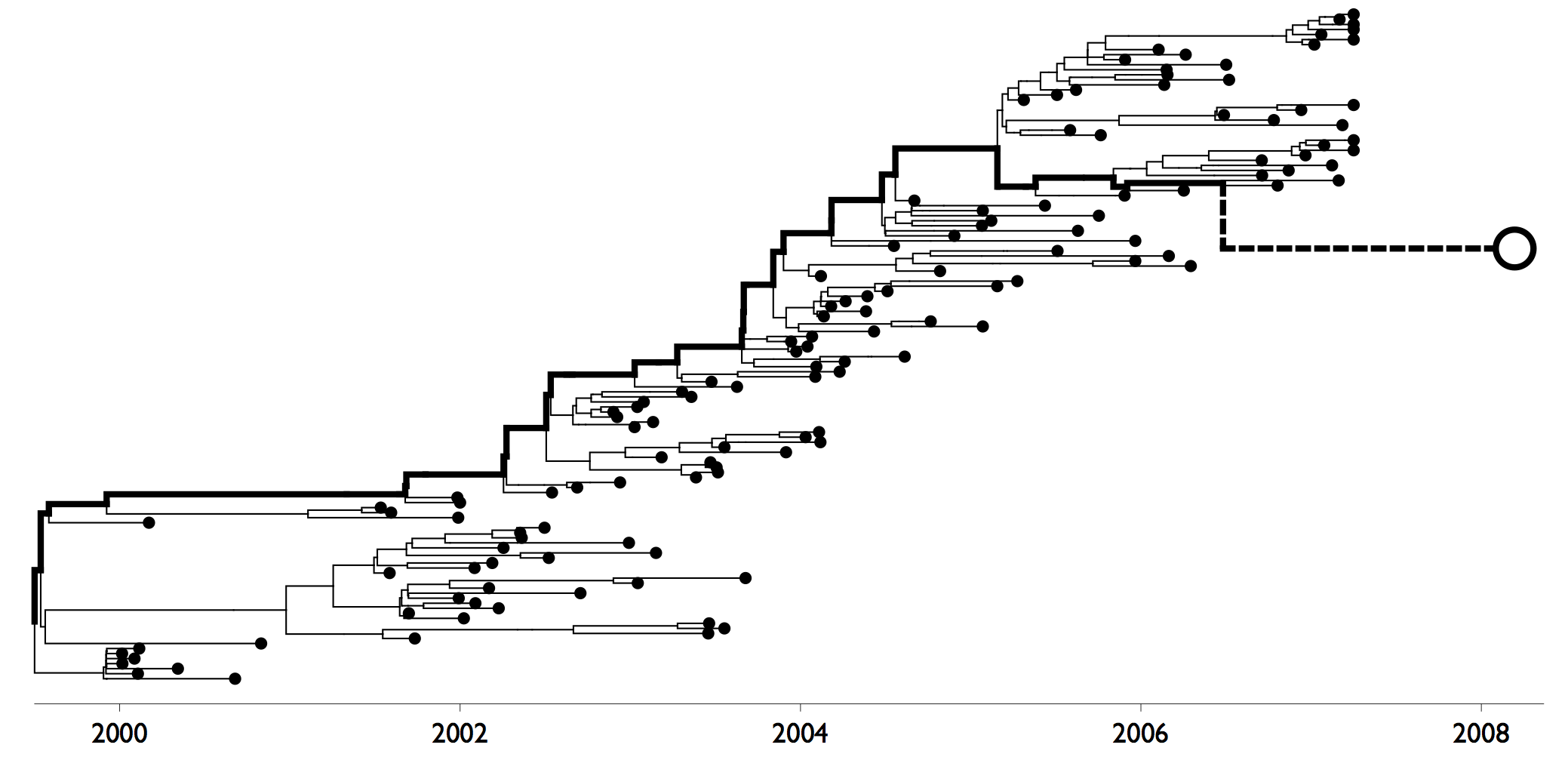
Want a model of an influenza tree where the trunk evolves differently than side branches. This is reflected by different branchRateModels for branches that descend from a single marked tip (trunk branches) and all other branches on the tree (side branches). Trunk branches are scaled by a parameter λ and side branches remain at 1. The rate model thus includes the overall rate μ, so that trunk branches have rate λ × μ and side branches have rate μ.
This branch rate model can be applied to different partitions on the same tree.
We can have a sequence partition from non-epitope sites where the trunk should evolve more slowly, a sequence partition from epitope sites where the trunk should evolve more quickly and a continuous trait partition from the antigenic MDS where the trunk should diffuse more quickly.
Tip to trunk mapping
Want to select a single strain from the set of contemporaneous strains that is most trunk-like in its evolution. Here, we pick an index for the initial stem taxon from this set and operate on this index to choose new stems over the MCMC.