Dengue genetic divergence generates within-serotype antigenic variation, but serotypes dominate evolutionary dynamics
Sidney M. Bell1,2, Leah Katzelnick3,4, Trevor Bedford1
1Vaccine and Infectious Disease Division, Fred Hutchinson Cancer Research Center, Seattle, WA, USA, 2Molecular and Cell Biology Program, University of Washington, Seattle, WA, USA, 3Division of Infectious Diseases and Vaccinology, School of Public Health, University of California, Berkeley, Berkeley, CA, USA, 4Department of Biology, University of Florida, Gainesville, FL, USA
Abstract
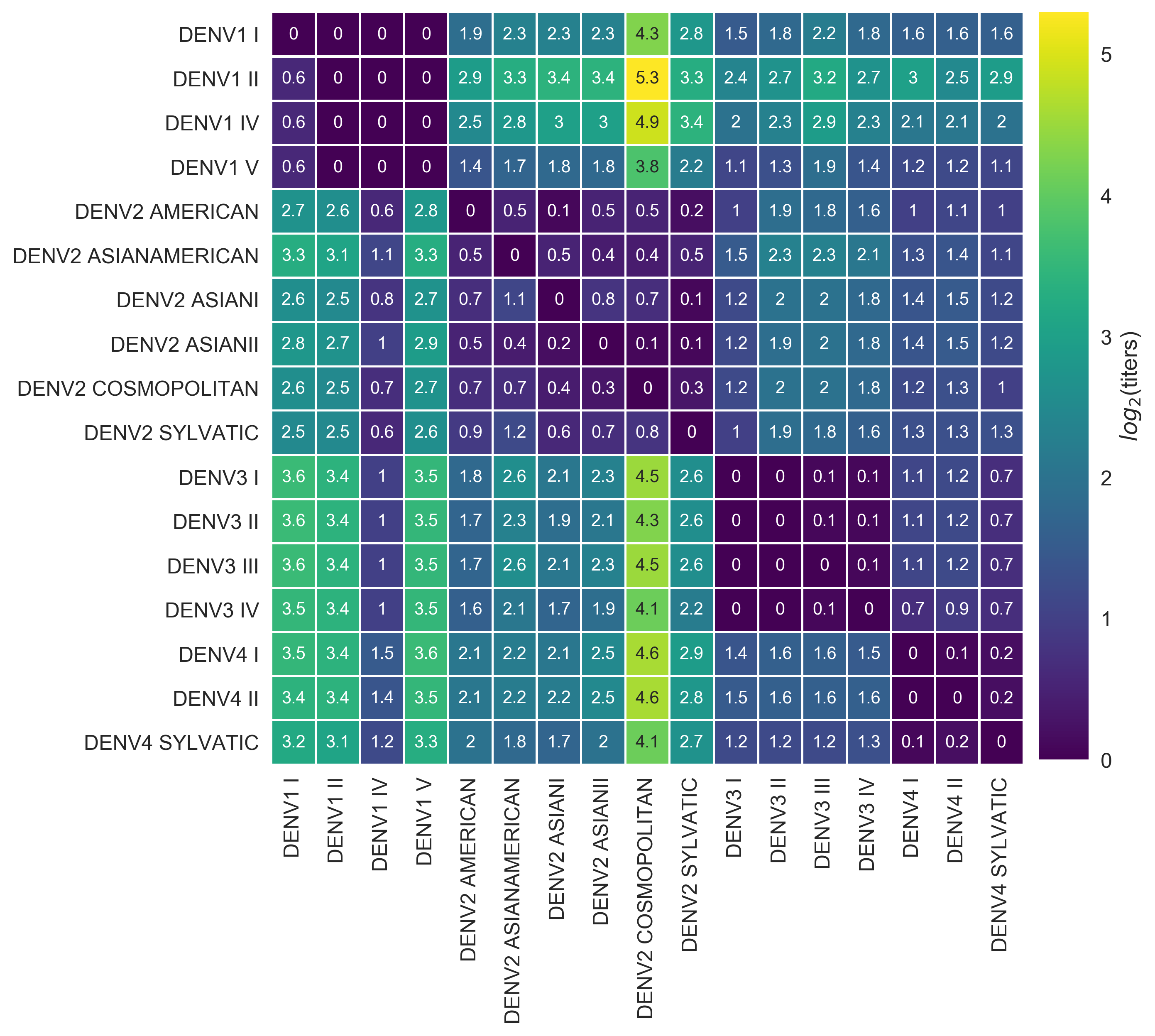
Dengue virus (DENV) exists as four genetically distinct serotypes, each of which is historically assumed to be antigenically uniform. However, recent analyses suggest that antigenic heterogeneity may exist within each serotype, but its source, extent and impact remain unclear. Here, we construct a sequence-based model to directly map antigenic change to underlying genetic divergence. We identify 49 specific substitutions and four colinear substitution clusters that robustly predict dengue antigenic relationships. We report moderate antigenic diversity within each serotype, resulting in variation in genotype-specific patterns of heterotypic cross-neutralization. We also quantify the impact of antigenic variation on real-world DENV population dynamics, and find that serotype-level antigenic fitness is a dominant driver of dengue clade turnover. These results provide a more nuanced understanding of the relationship between dengue genetic and antigenic evolution, and quantify the effect of antigenic fitness on dengue evolutionary dynamics.
Install
Everything is Python 2.7 based. Python packages that are required can be installed via:
cd dengue-antigenic-dynamics/
pip install -r requirements
Analysis outline
- Run the titer model via augur (repackaged portion of the Nextstrain pipeline) to build a viral phylogeny, assign antigenic change to specific mutations, and infer clade frequencies.
- Run the fitness model to quantify population immunity over time, predict clade frequencies, and assess performance.
- Use the visualization notebooks to explore results and recreate all the figures from the paper.
Citation
All contents including manuscript text and source code are copyright 2016-2019 Sidney Bell, Leah Katzelnick and Trevor Bedford. All manuscript text (files ending in .tex) are licensed under Creative Commons Attribution 4.0 and all code (files ending in .py and .ipynb) is licensed under an MIT License.