Fitness flux in SARS-CoV-2 and influenza H3N2
Trevor Bedford
Fred Hutchinson Cancer Center / Howard Hughes Medical Institute
3 Mar 2026
VRC Tuesday Seminar
NIH Vaccine Research Center
Slides at: bedford.io/talks
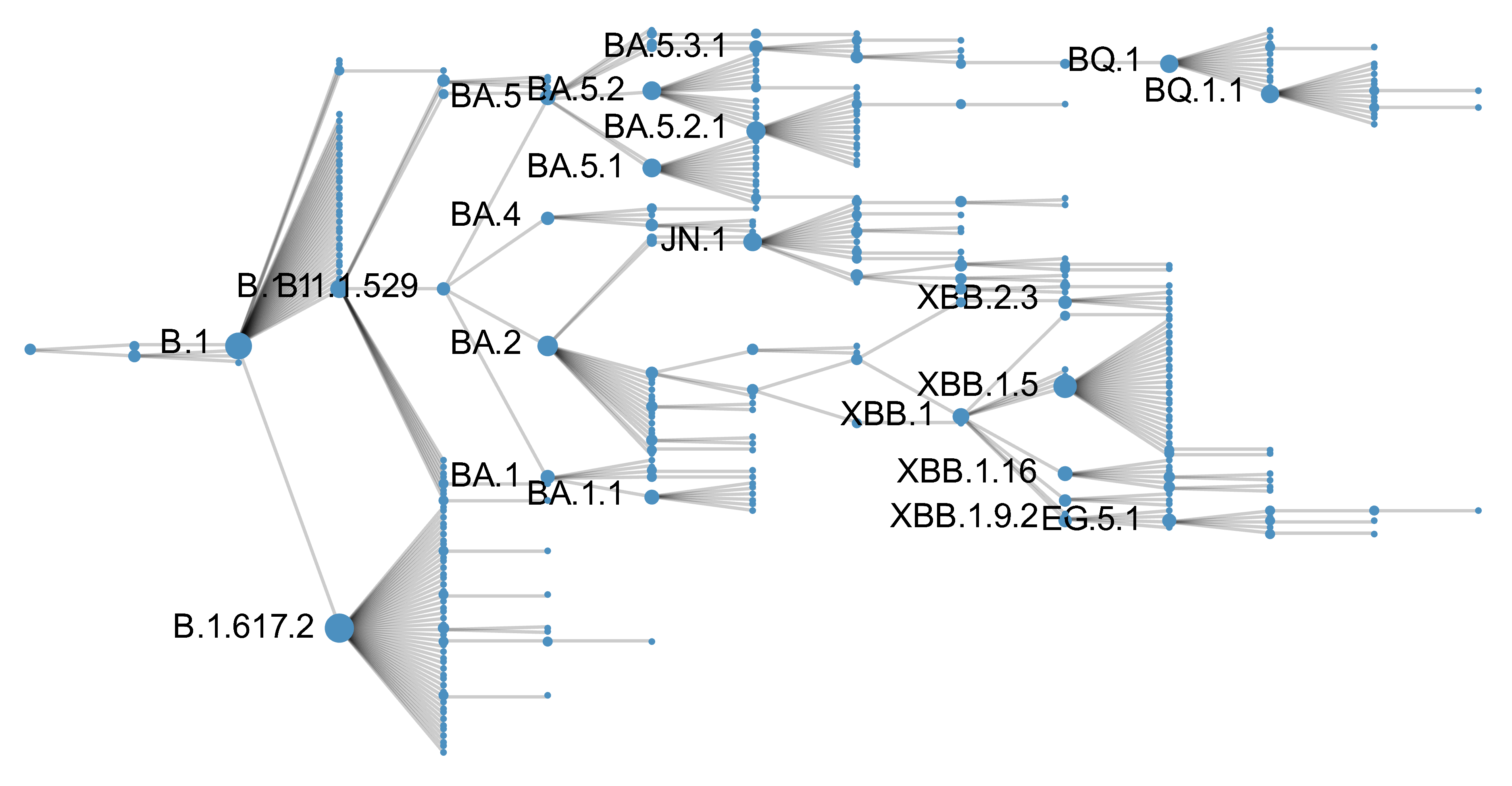
Genetic relationships of SARS-CoV-2 from 2020 to present
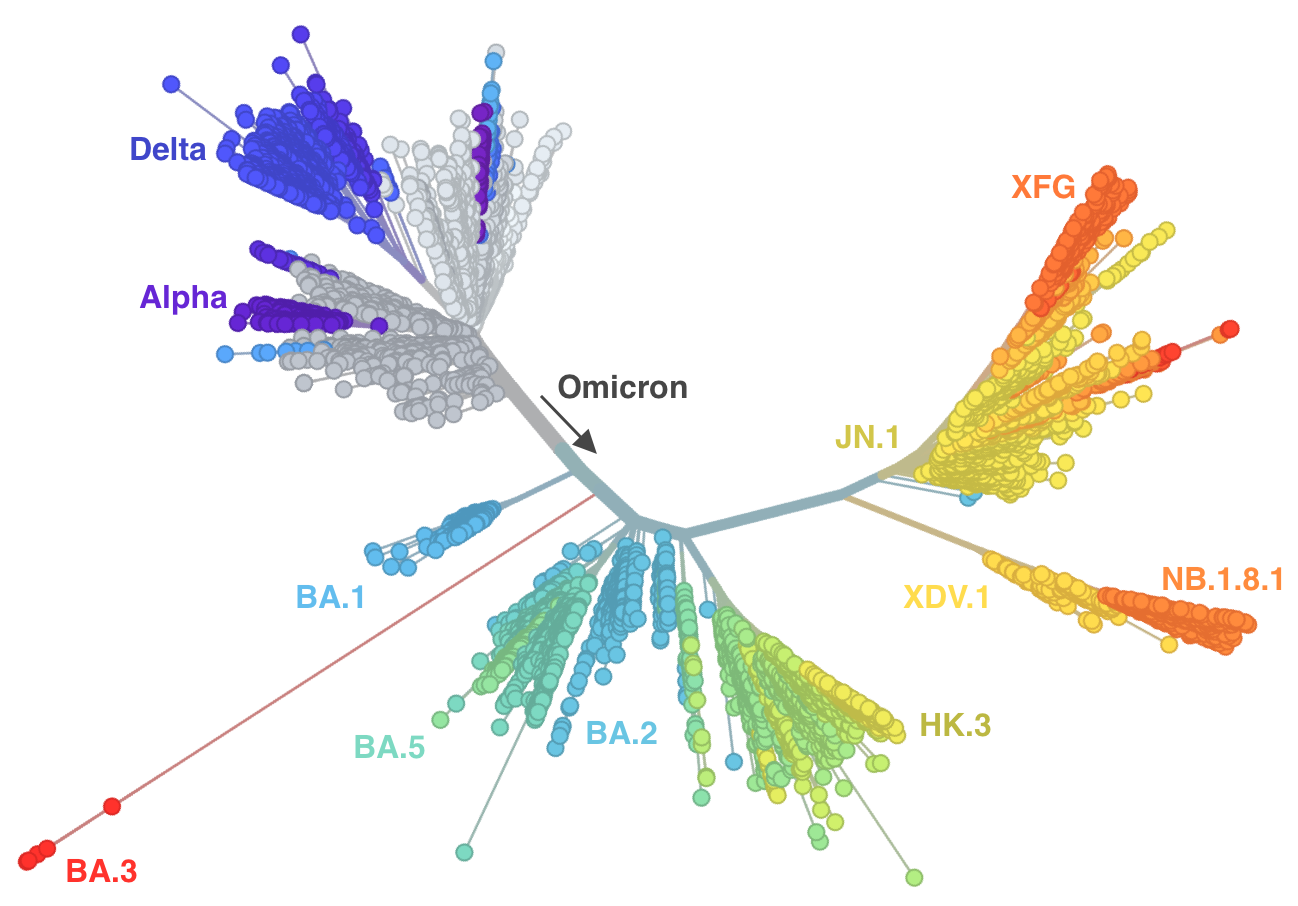 nextstrain.org
nextstrain.org
Rapid displacement of existing diversity by emerging variants
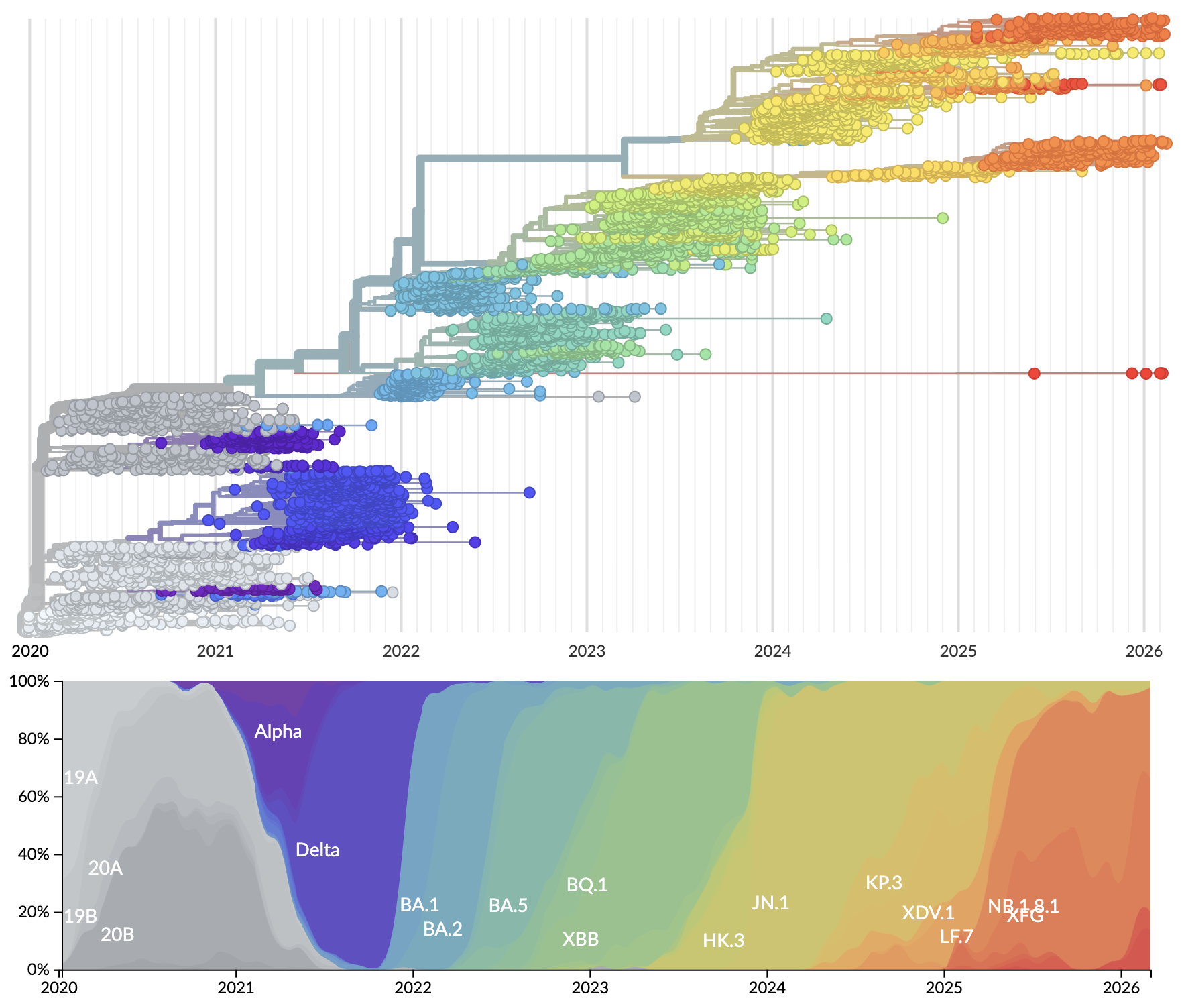
Mutations in S1 domain of spike protein driving displacement
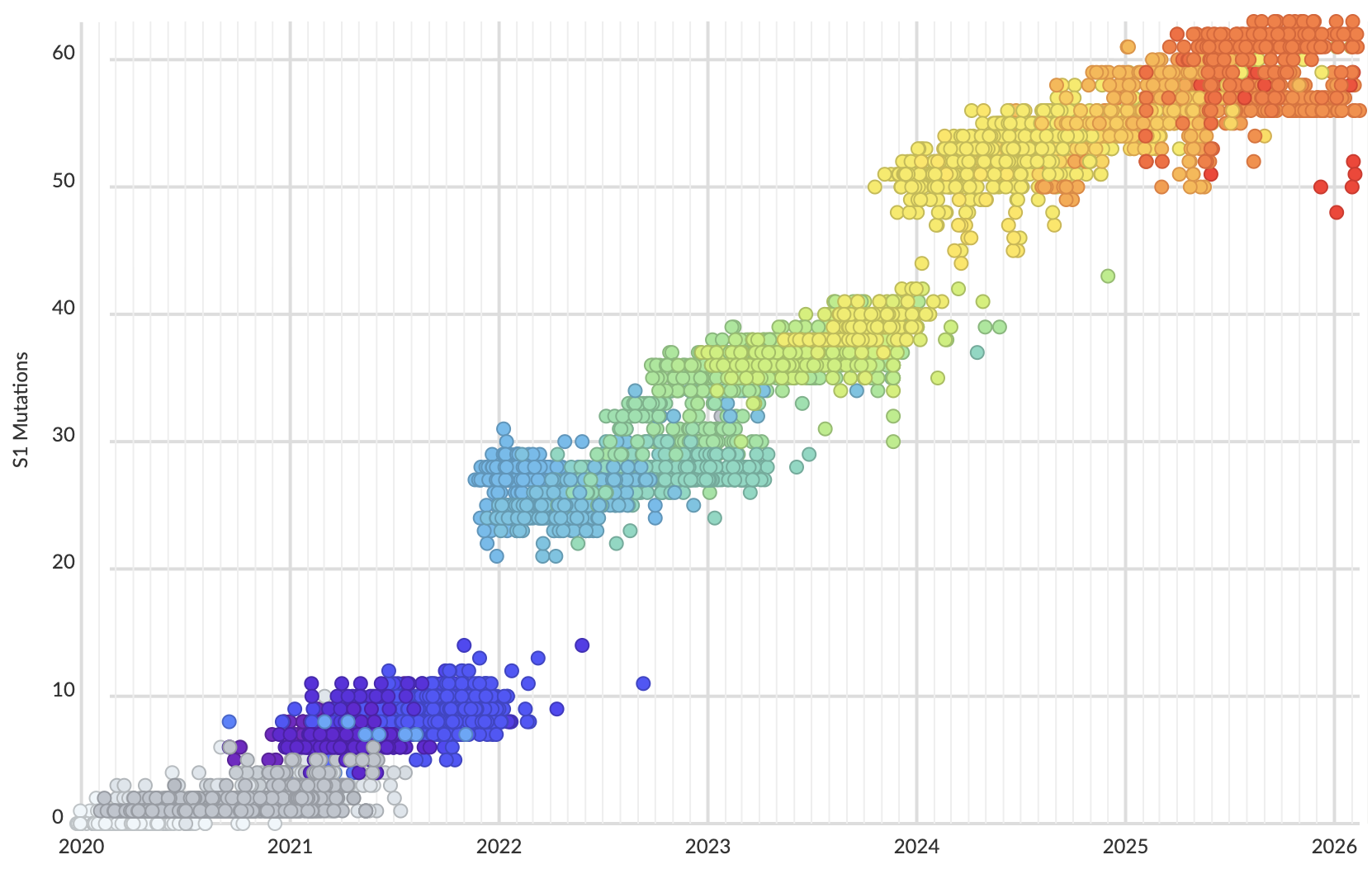 nextstrain.org
nextstrain.org
SARS-CoV-2 evolution fast relative to previous endemic viruses
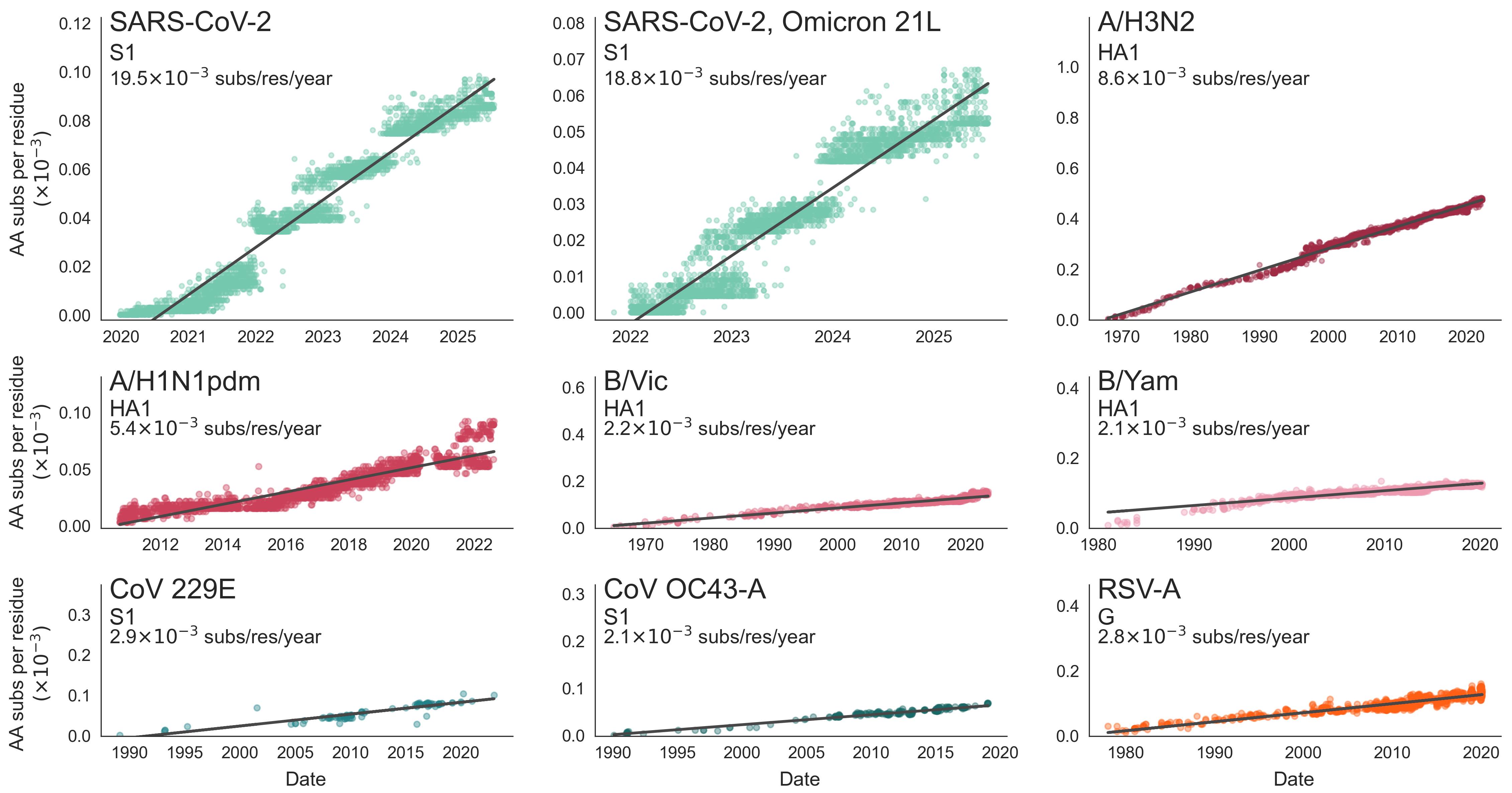
This talk
- Frequency dynamics
- Fitness flux
- Mutational fitness effects
Frequency dynamics
Multinomial logistic regression (MLR)
Simple haploid population genetic model is equivalent to statistical multinomial logistic regression
The frequency $x_i(t)$ of variant $i$ at time $t$ is determined by its initial frequency $p_i$ along with its fitness $f_i$ following
$$x_i(t) = \frac{p_i \, \mathrm{exp}(f_i \, t)}{\sum_j p_j \, \mathrm{exp}(f_j \, t) }$$
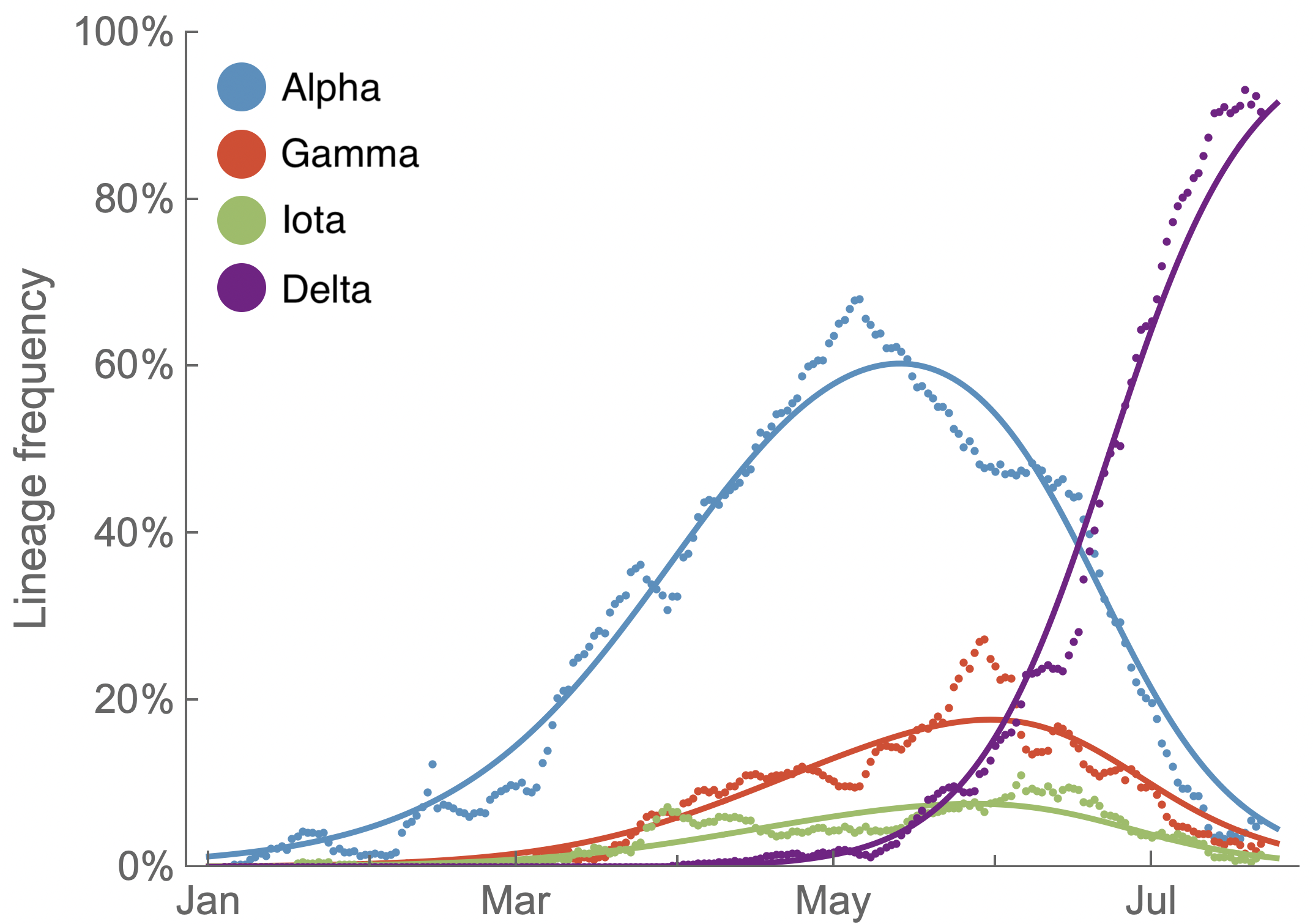
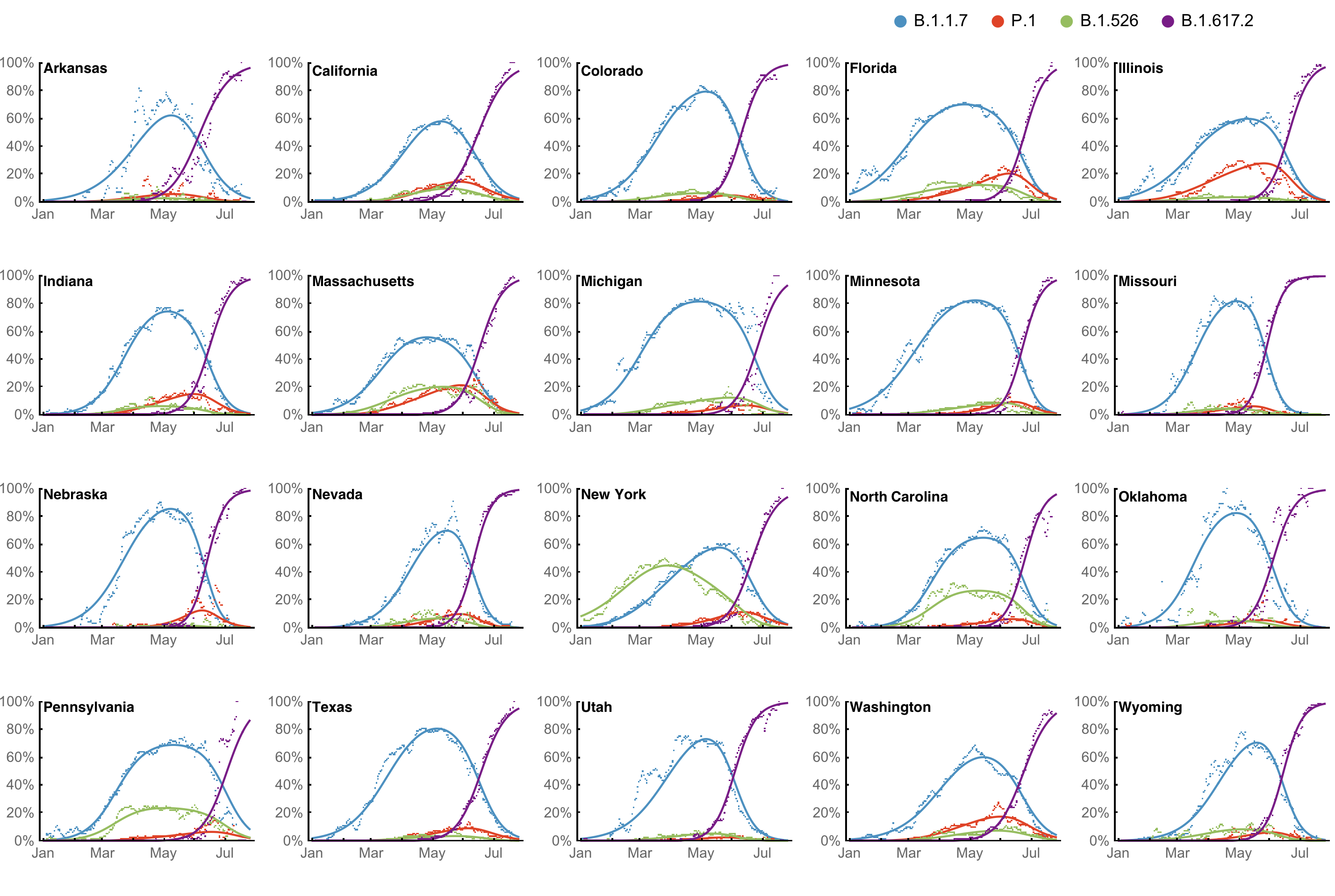
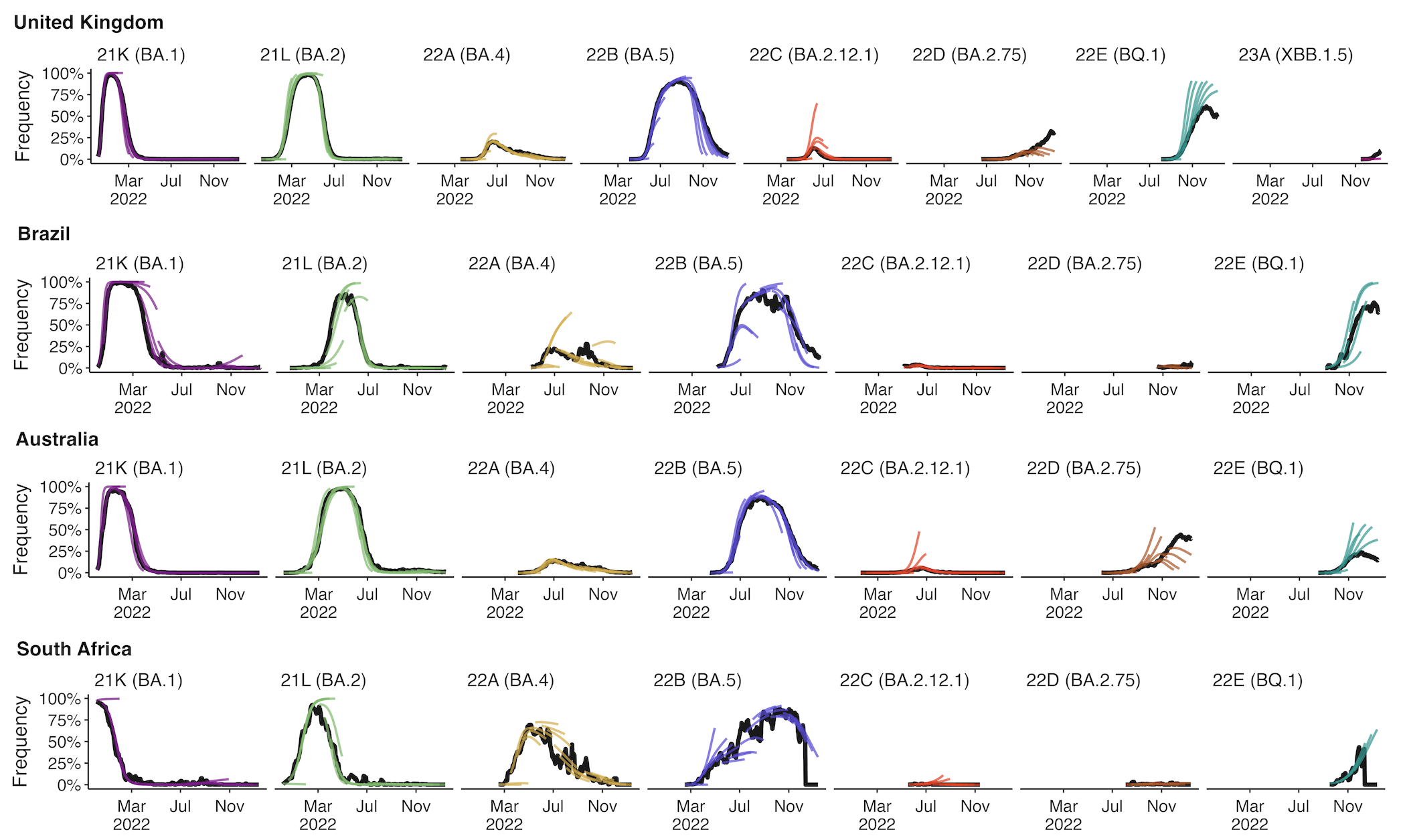
Multinomial logistic regression fits variant frequencies well
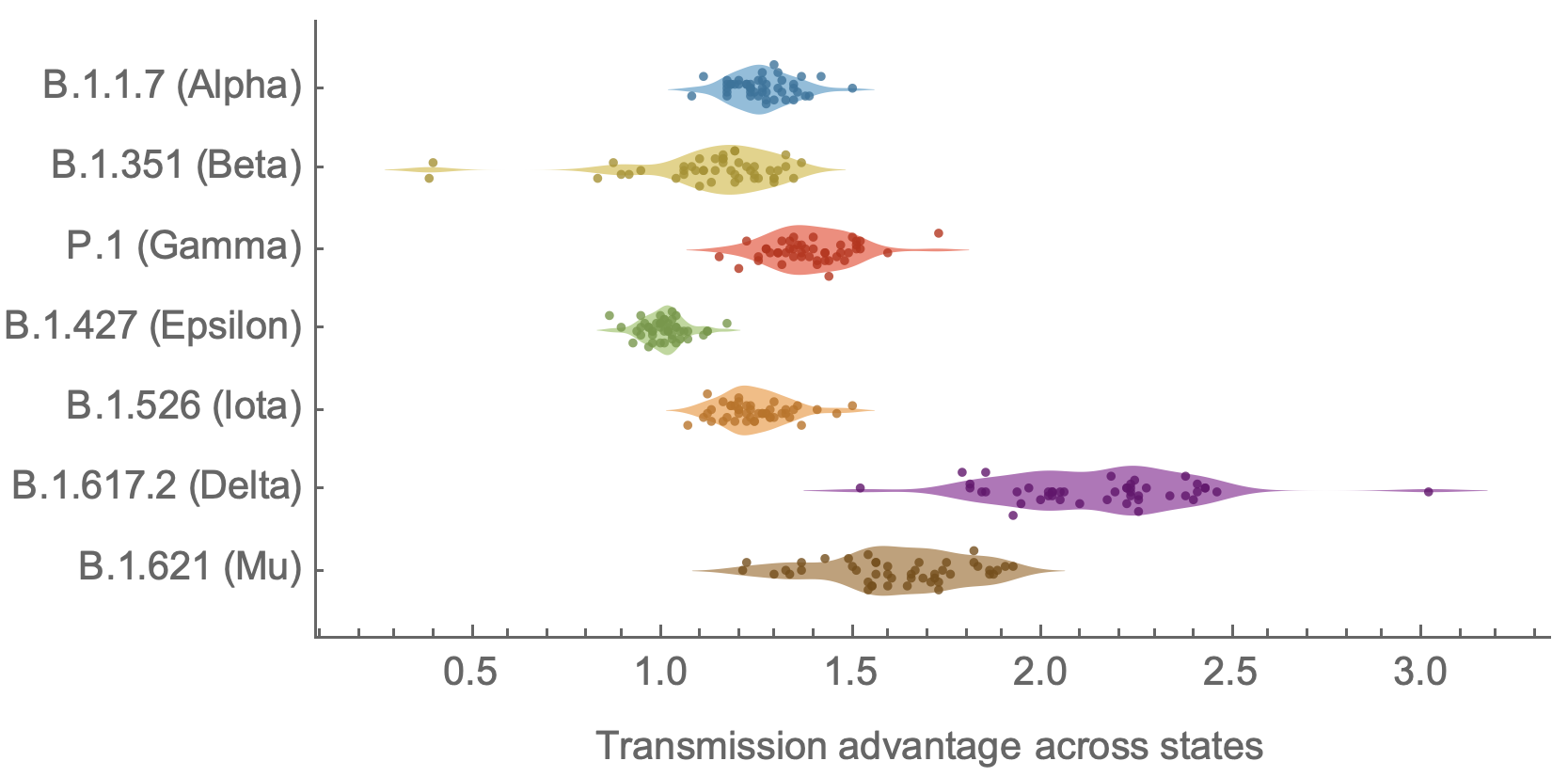
Original VOC viruses had substantially increased transmissibility
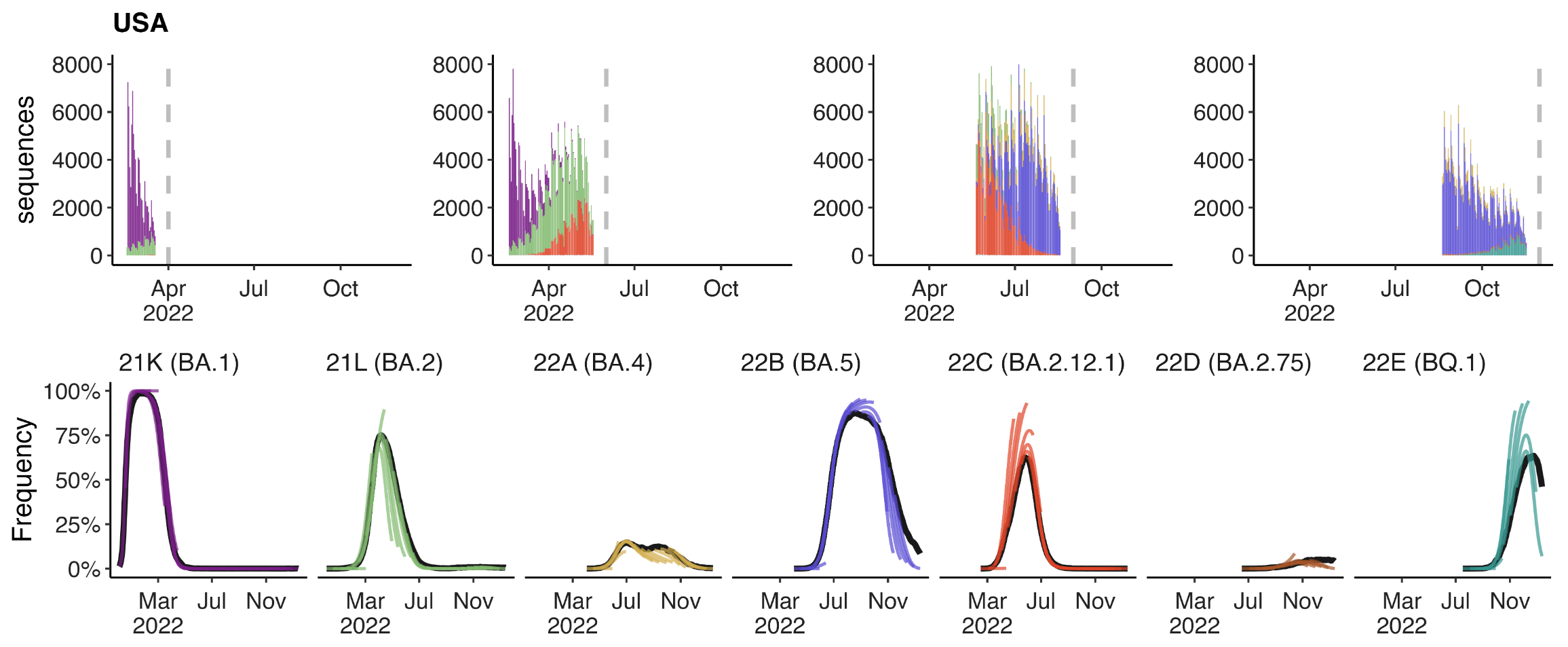
Assessing MLR models for short-term frequency forecasting
Retrospective projections twice monthly during 2022
+30 day short-term forecasts across different countries
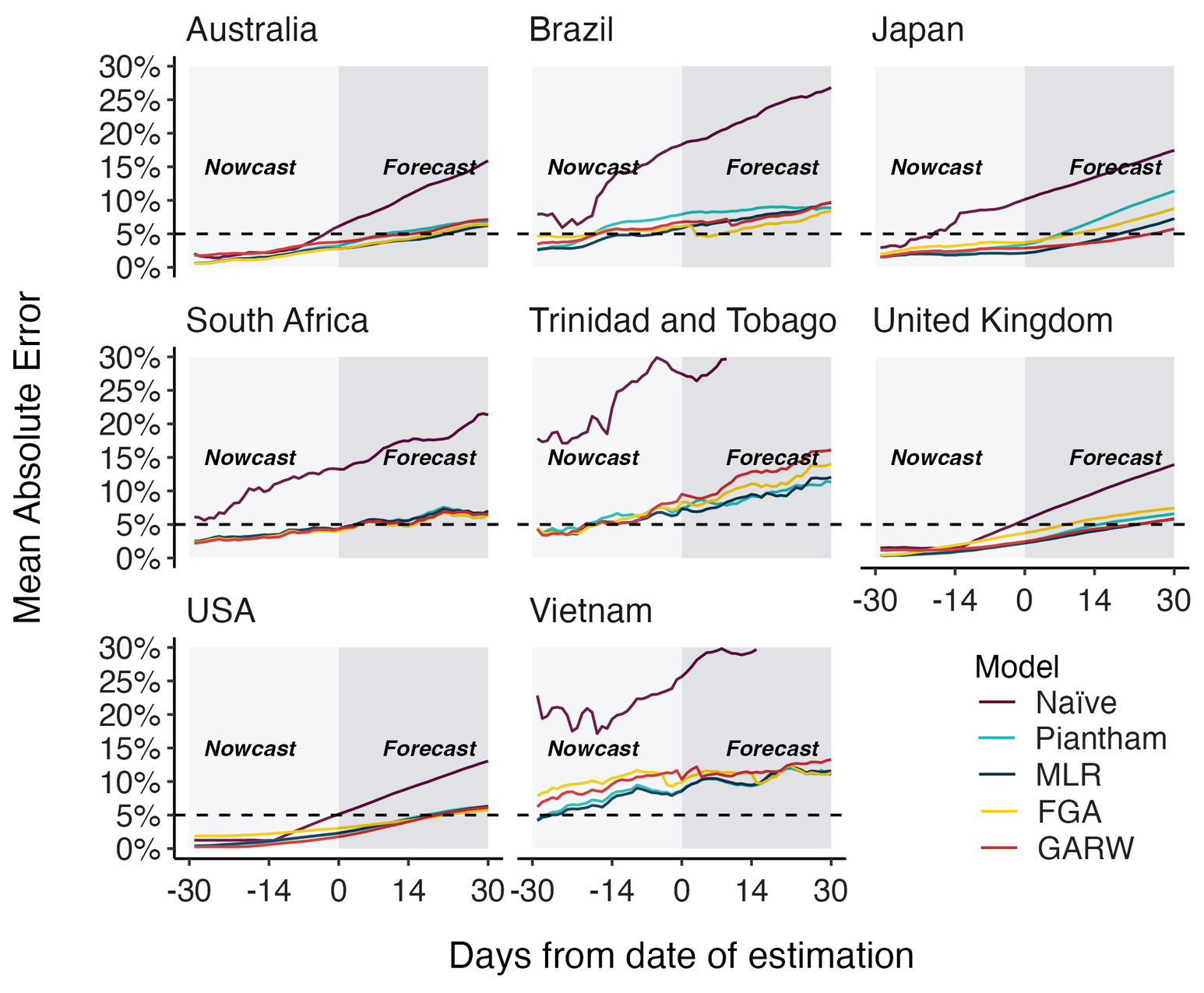
MLR models generate accurate short-term forecasts
30 days out, countries range from 5 to 15% mean absolute error
Correlates with data availability (median number of sequences available from the previous 30 days):
- USA
- ~45k sequences
- Australia
- ~4k sequences
- South Africa
- 170 sequences
- Vietnam
- 30 sequences
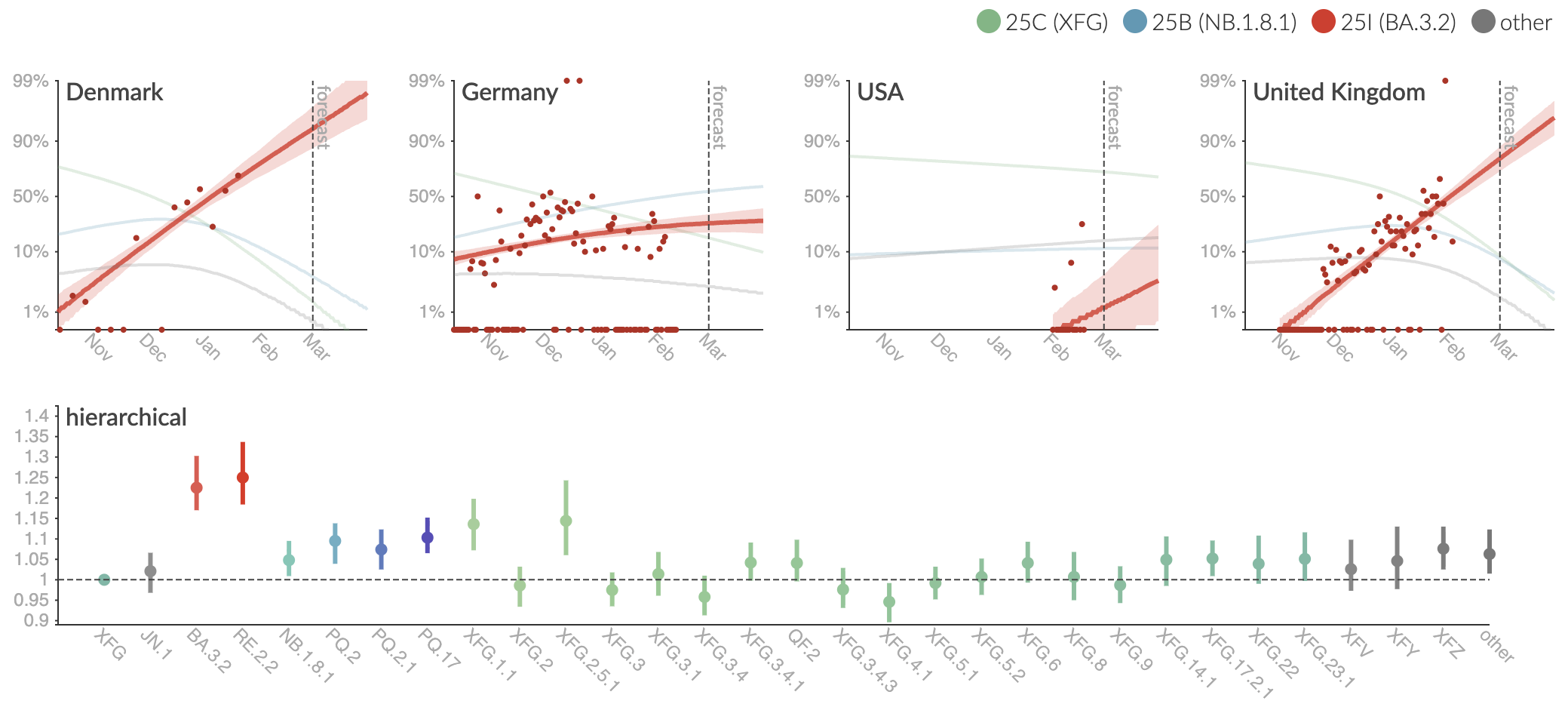
Clade and lineage forecasts continuously updated
BA.3 has been slowly growing, though does not yet have much of a toehold in the US
Fitness flux
Fitness flux is the rate of change of mean population fitness
With variant frequency $x_i(t)$ and constant variant fitness $f_i$
Mean population fitness $\bar{f}(t) = \sum_i x_i(t) \, f_i$ Fitness flux $\phi(t) = \Delta \bar{f}(t) / \Delta t$
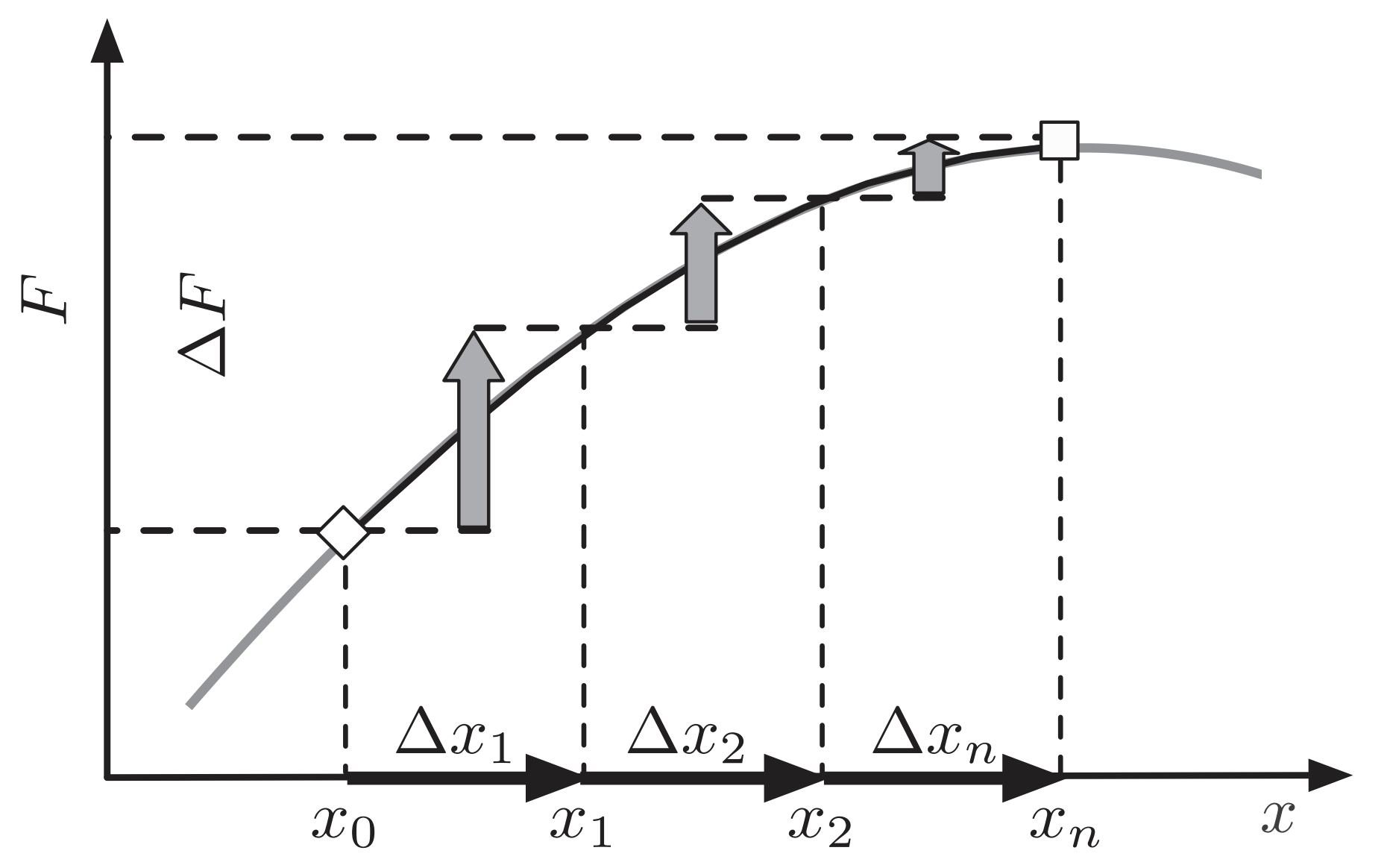 Mustonen and Lässig. 2010. PNAS.
Mustonen and Lässig. 2010. PNAS.
Clade-level frequency dynamics and MLR fits in sliding windows
Constant clade fitness within each window, USA data only, ignores within-clade fitness variation
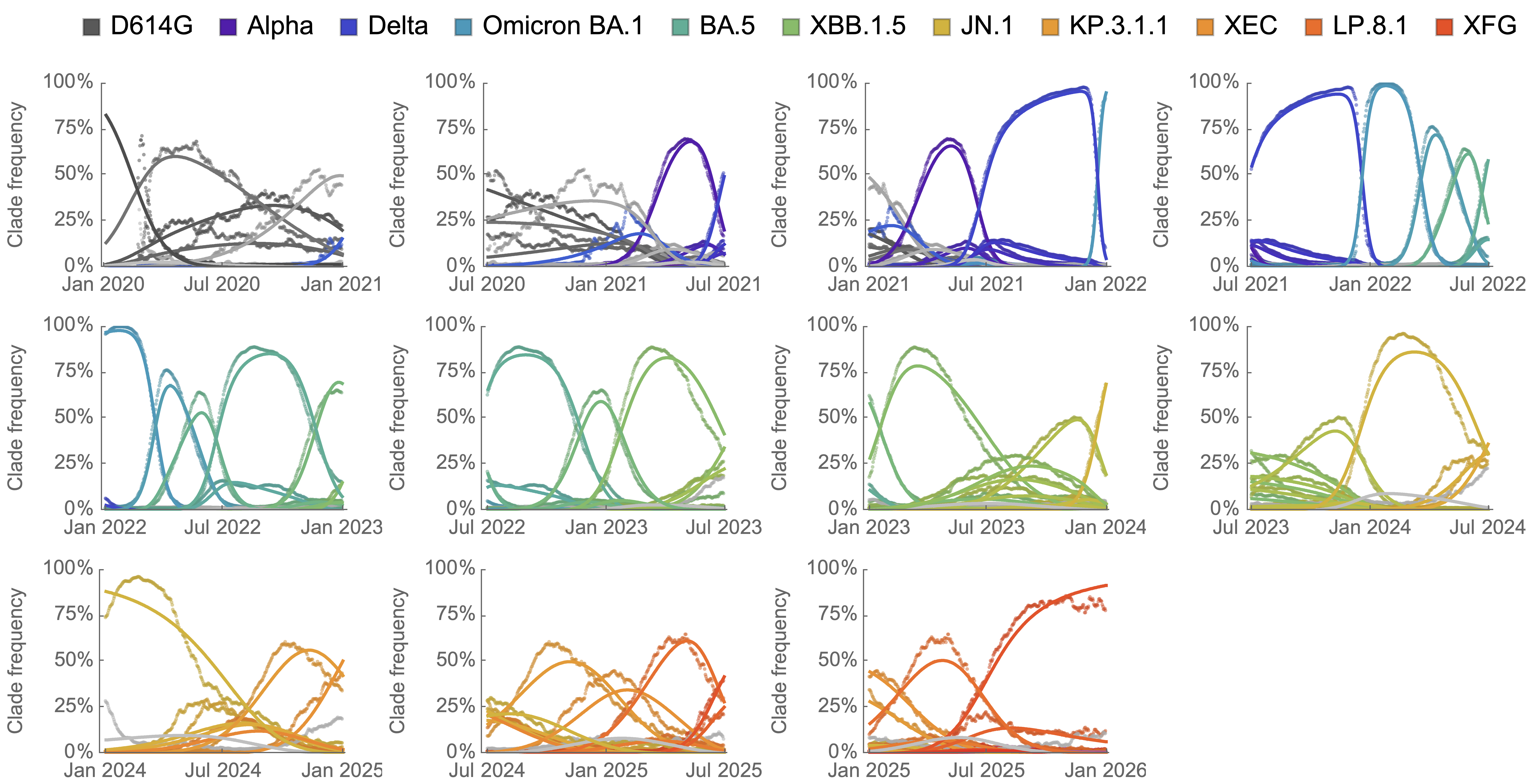
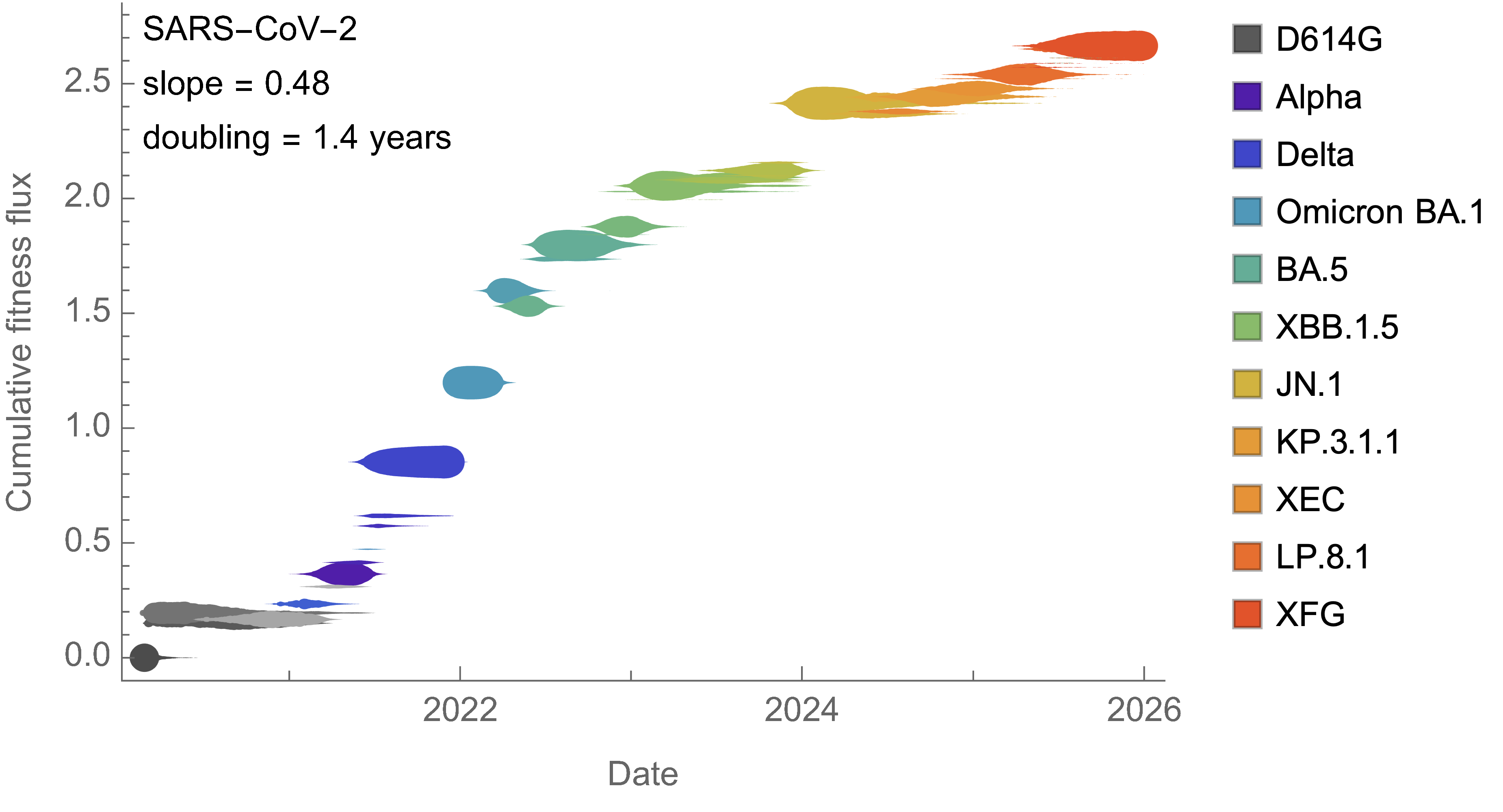
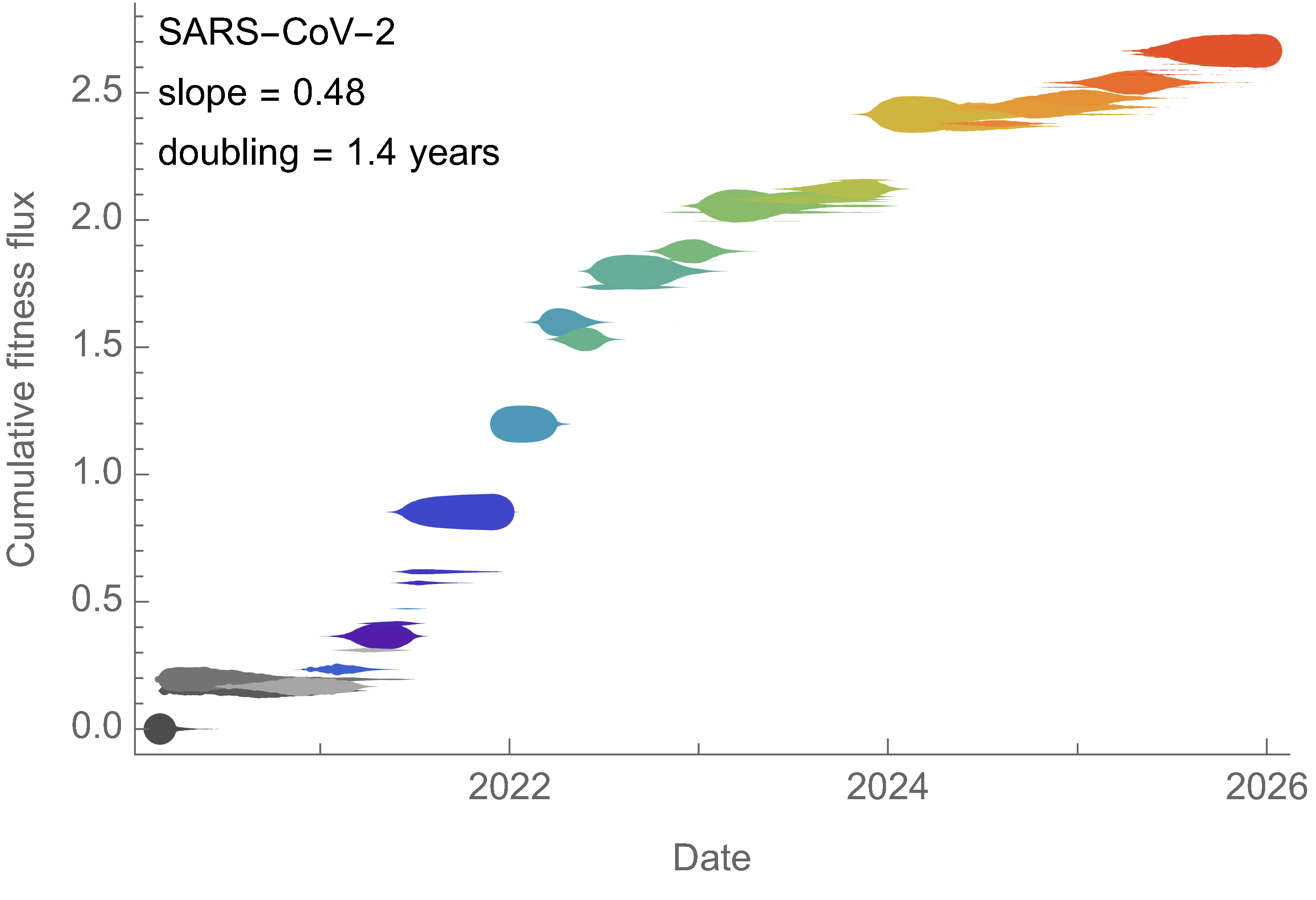
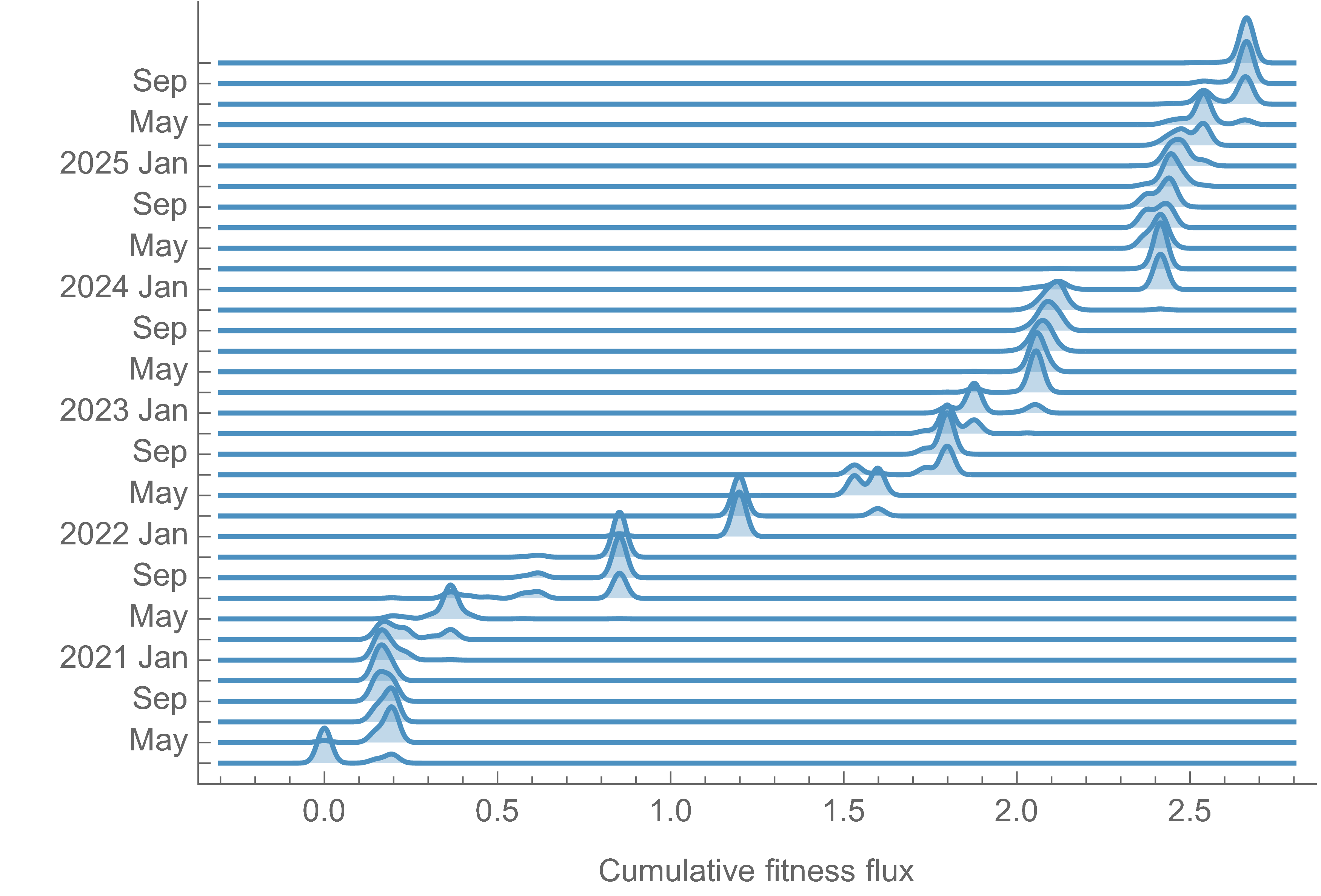
Charting fitness of SARS-CoV-2 variants through time
Initially doubled in fitness every year, but has slowed since 2024
Line thickness is proportional to variant frequency, 44 total variants
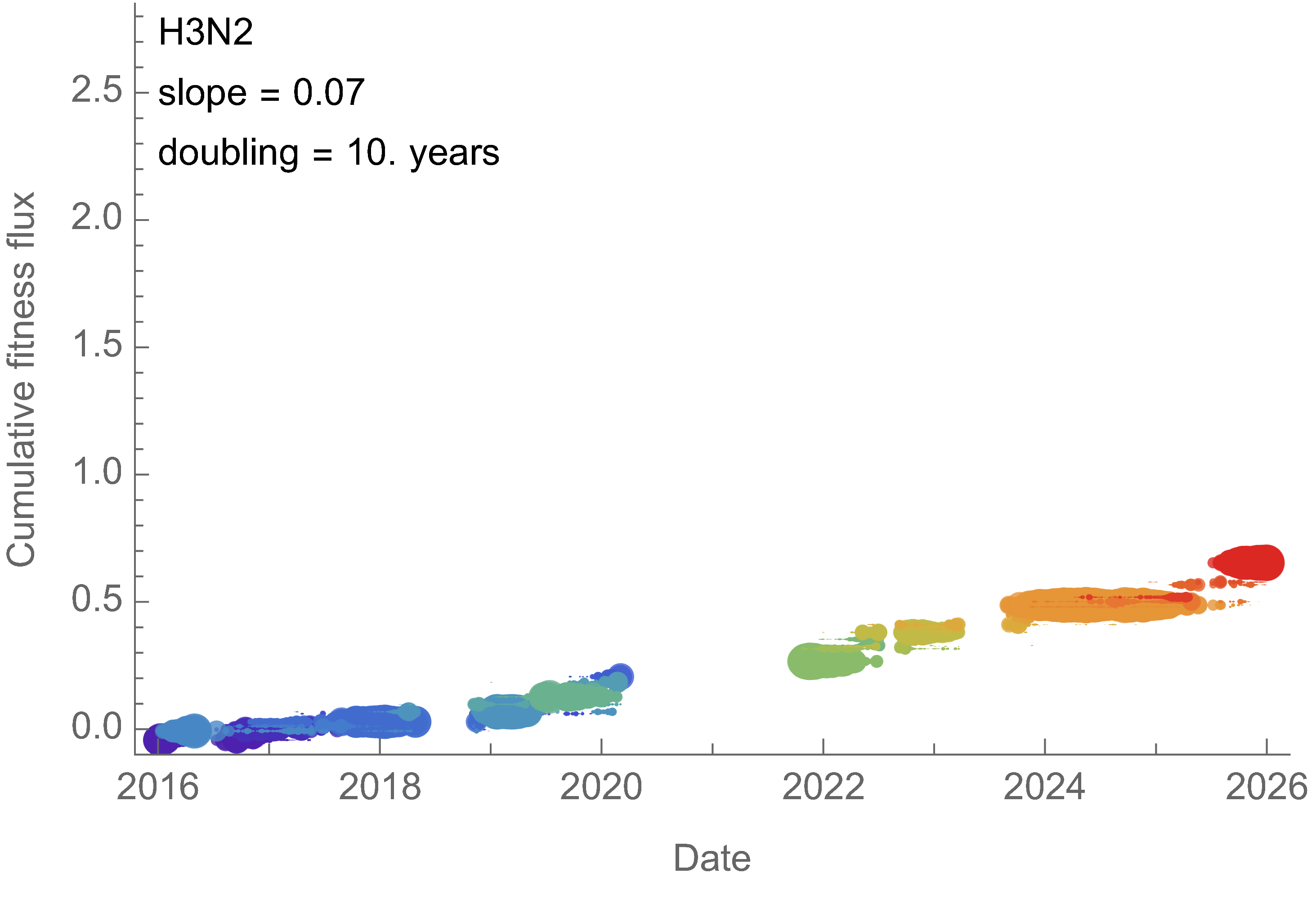
Analogous frequency dynamics and MLR fits for H3N2
Constant clade fitness within each window, USA data only, ignores within-clade fitness variation
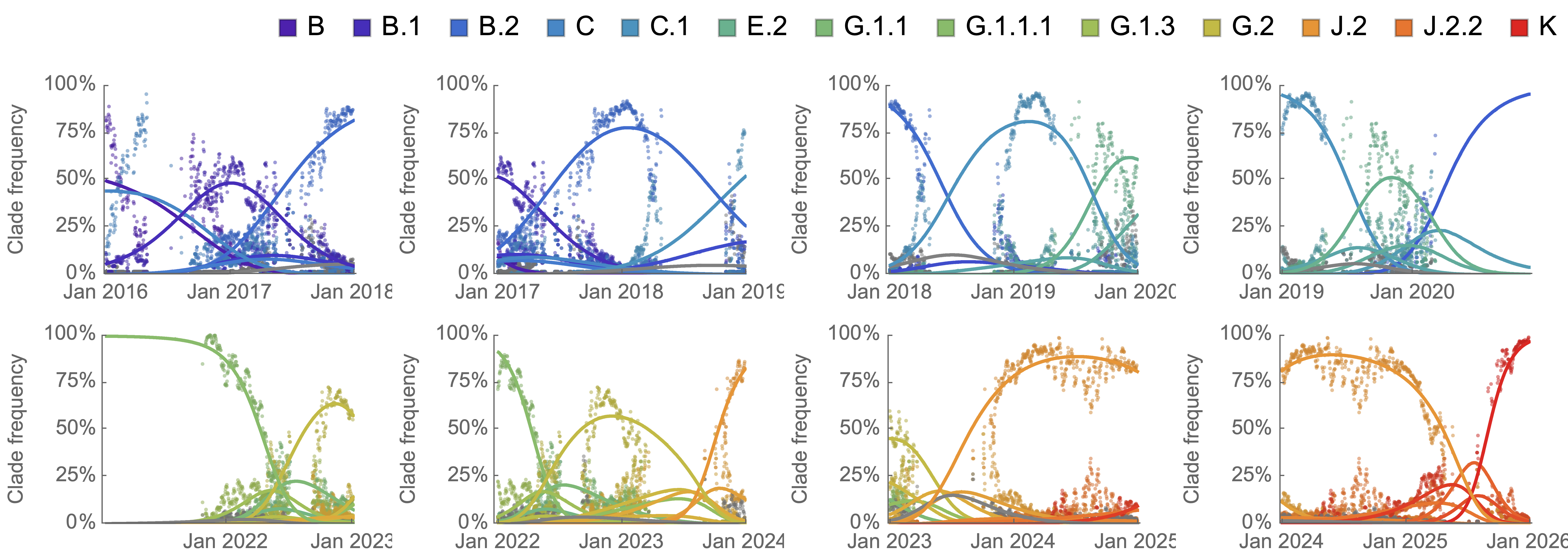
H3N2 roughly doubled in fitness every 10 years
Line thickness is proportional to variant frequency, 32 total variants
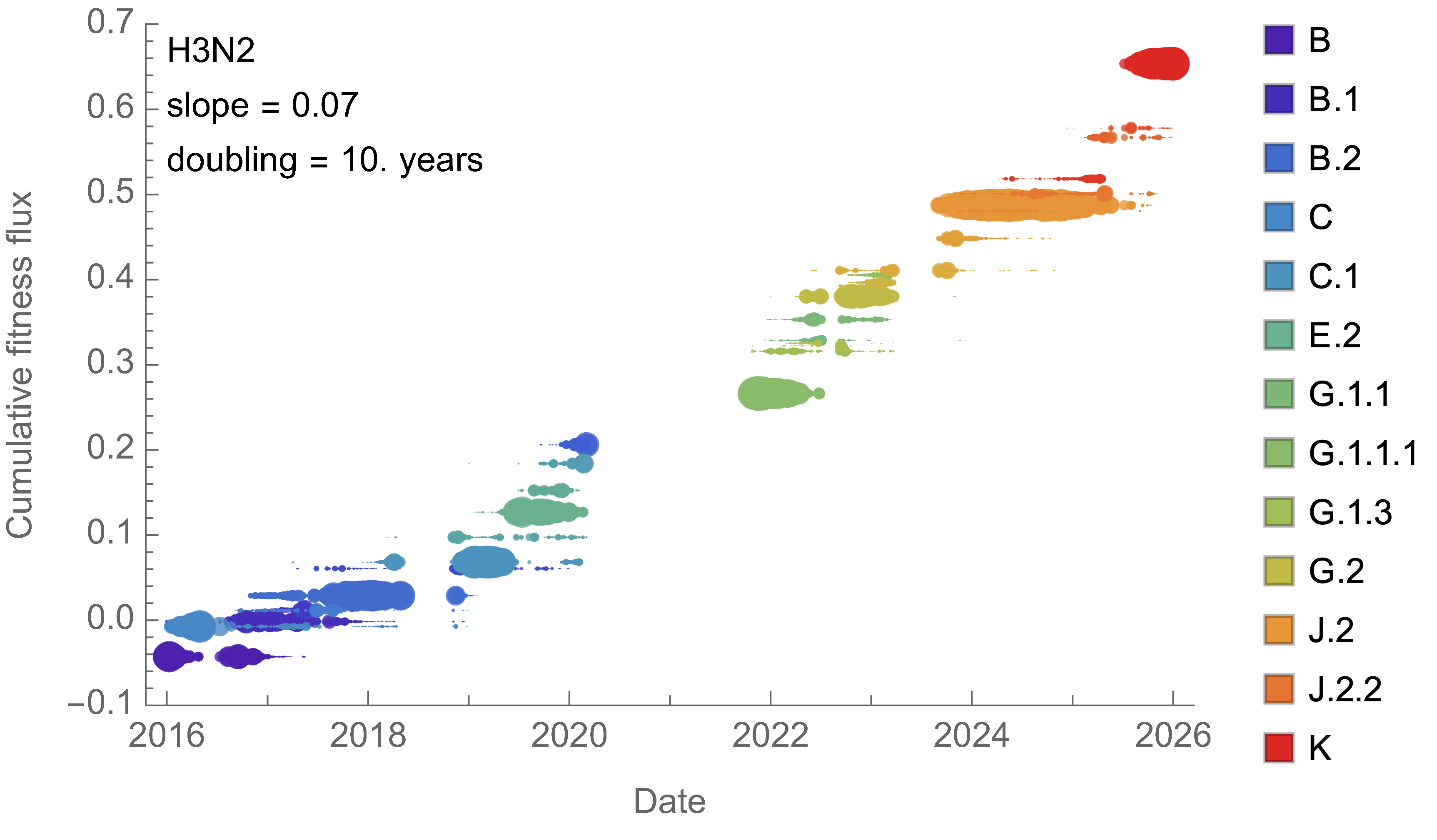
Dramatically faster fitness flux of SARS-CoV-2
Traveling fitness waves
Multistrain models produce traveling waves in antigenic space
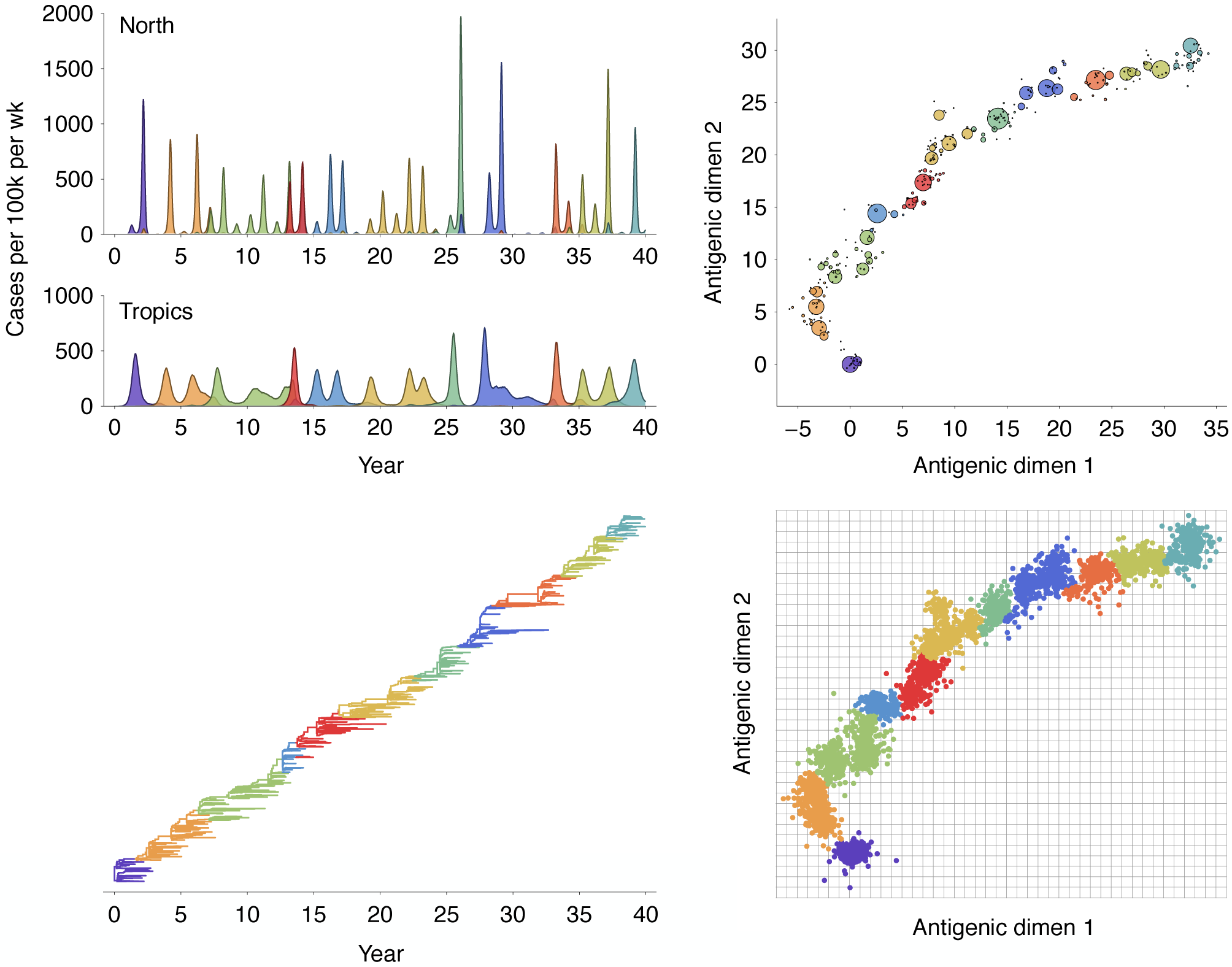 Bedford, Rambaut and Pascual. 2012. BMC Biol.
Bedford, Rambaut and Pascual. 2012. BMC Biol.
Many mutations of small effect create traveling fitness waves
Richard Neher and others have analytically characterized these waves
Diffusion constant $D = \mu \, \langle \delta^2 \rangle/2$, where the average $\langle \ldots \rangle$ is over the distribution of mutational effects $K(\delta)$
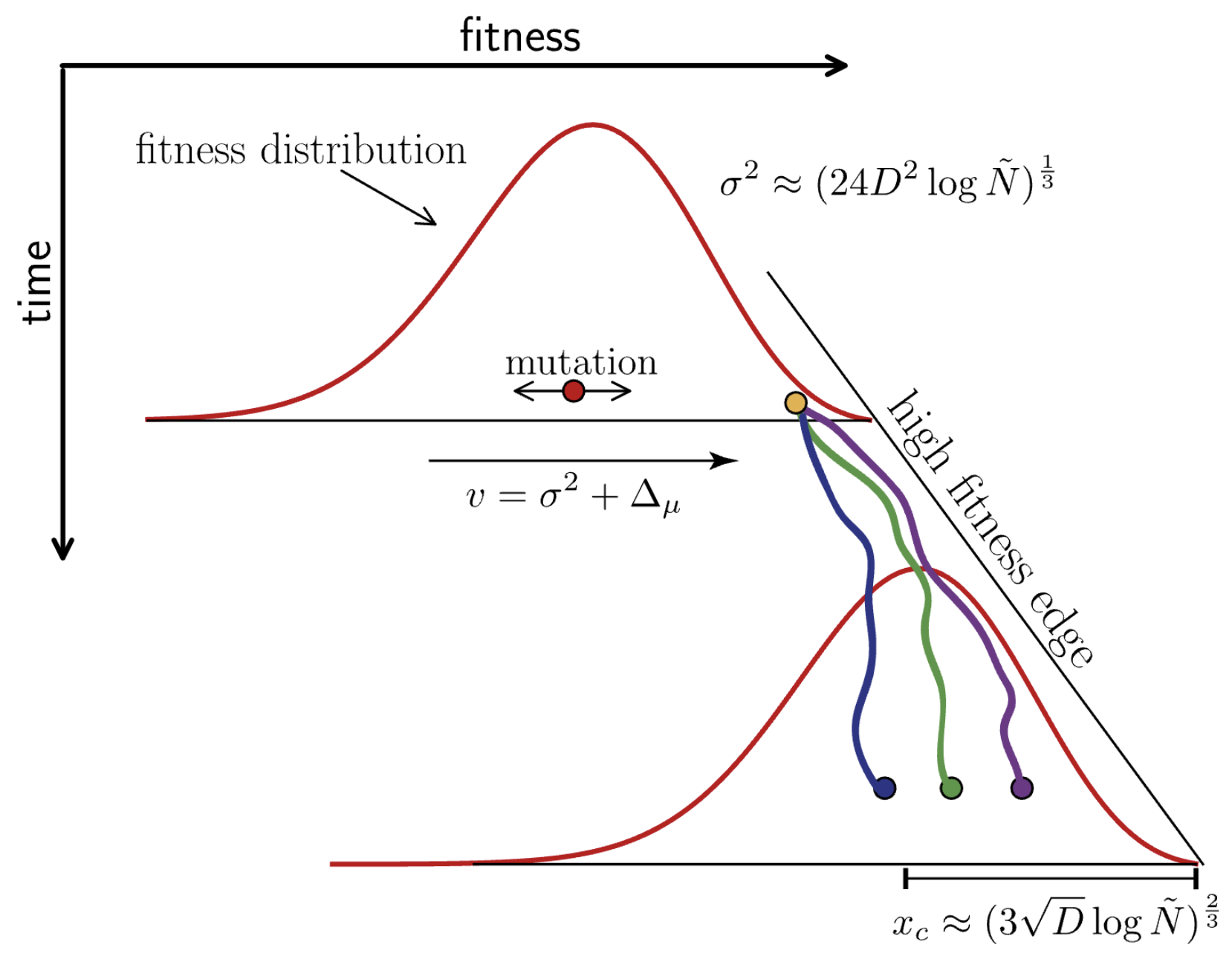 Neher and Hallatschek. 2013. PNAS.
Neher and Hallatschek. 2013. PNAS.
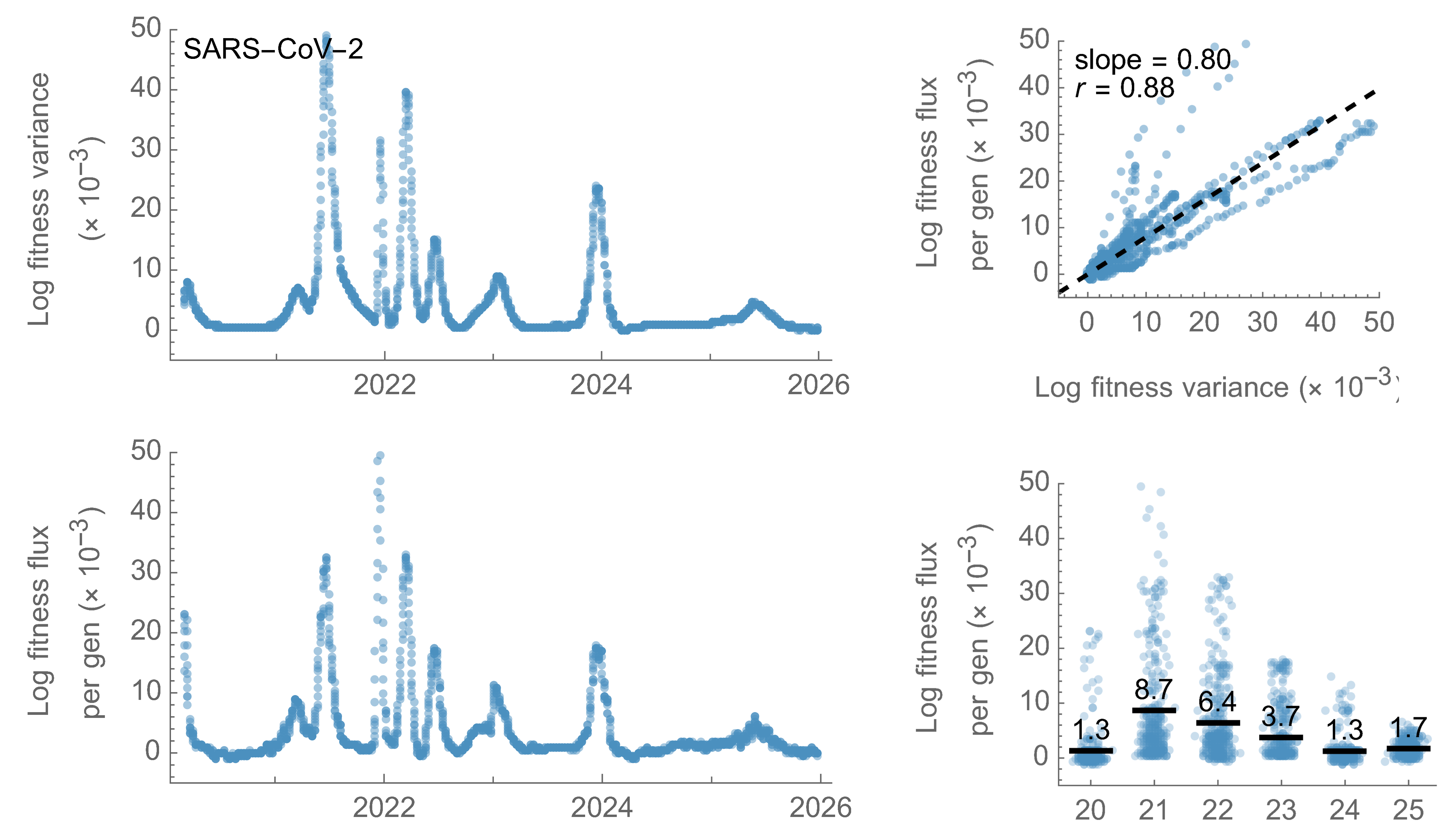
Using empirical frequencies and MLR fitness to characterize fitness wave in SARS-CoV-2
SARS-CoV-2 fitness variance correlates well with fitness flux
H3N2 fitness variance also correlates well with fitness flux
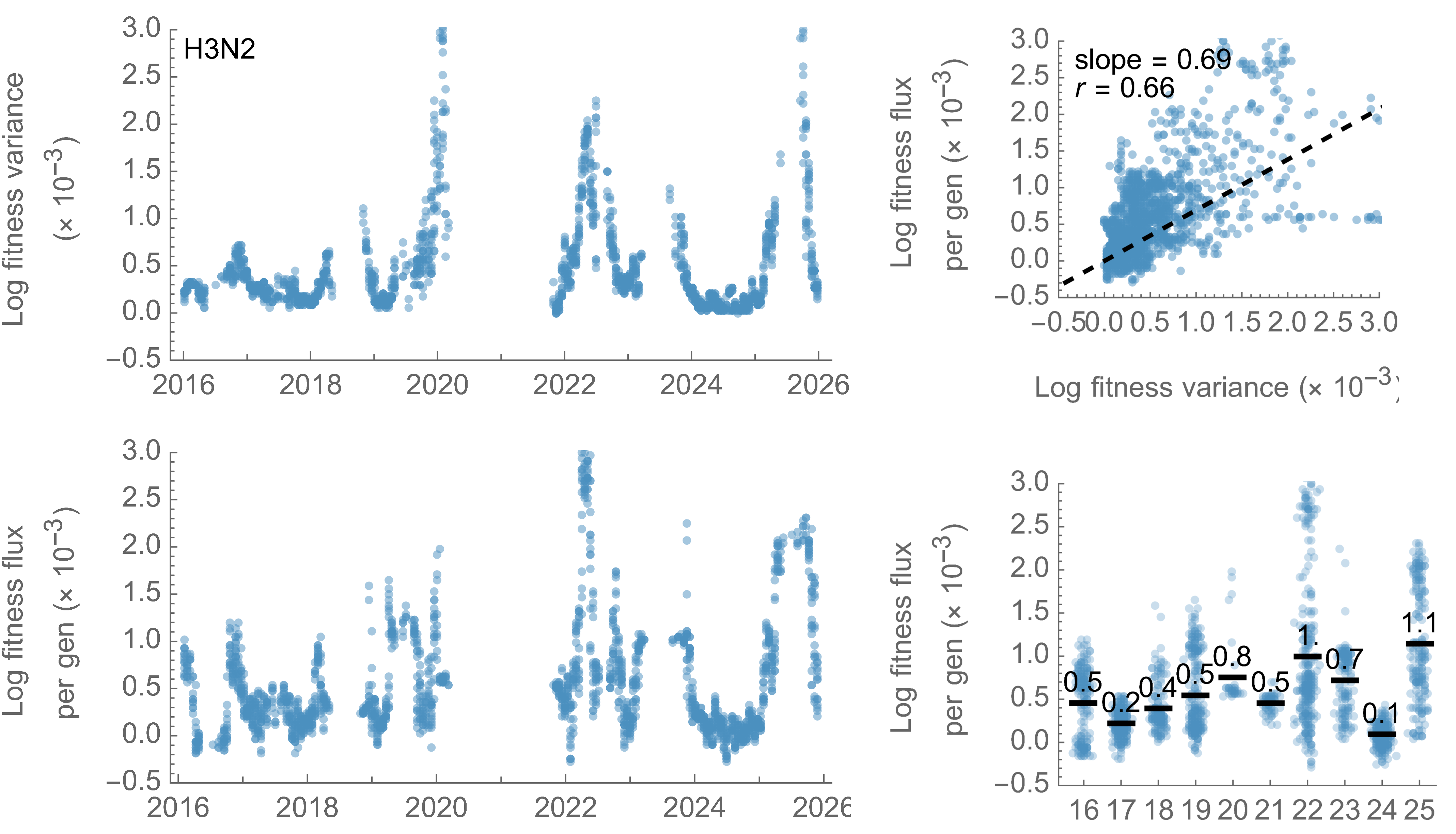
This is a specific example of Fisher's fundamental theorem
"The rate of increase in fitness of any organism at any time is equal to
its genetic variance in fitness at that time," ie
$$\frac{d\bar{f}}{dt} = Var(f)$$
Mutational fitness effects
Analyze MLR fitness between parent/child lineages
Expand to 367 Pango lineages with at least 1000 sequence counts in the US from 2020 to 2025
Similar concept to Obermeyer et al
Most Pango branches have 0-1 spike mutations and change log fitness by ±0.1
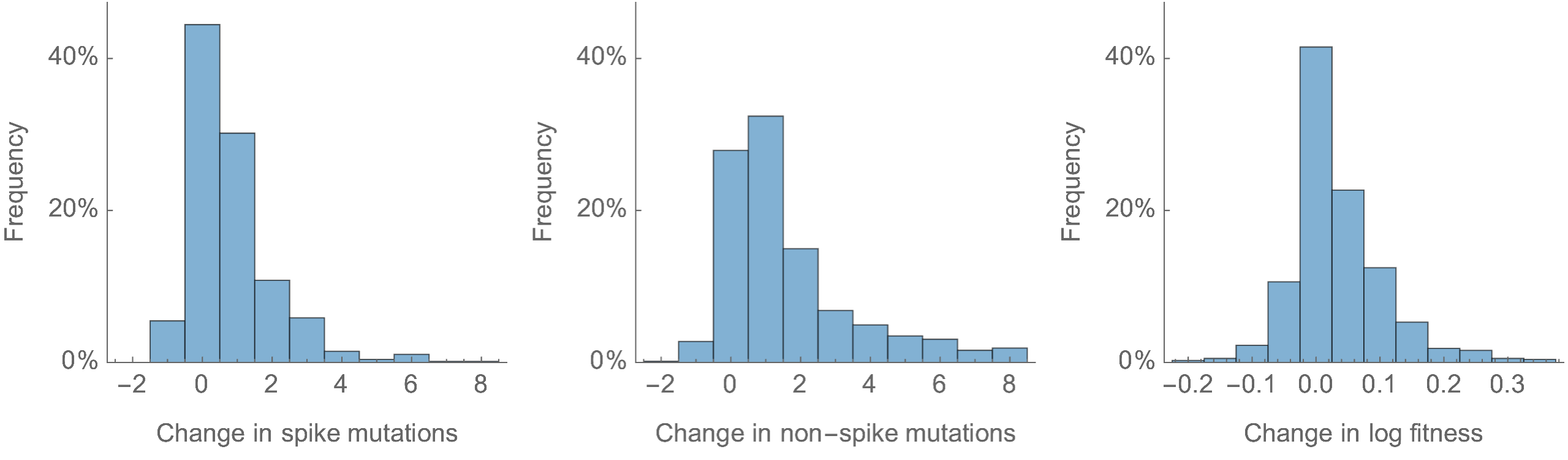
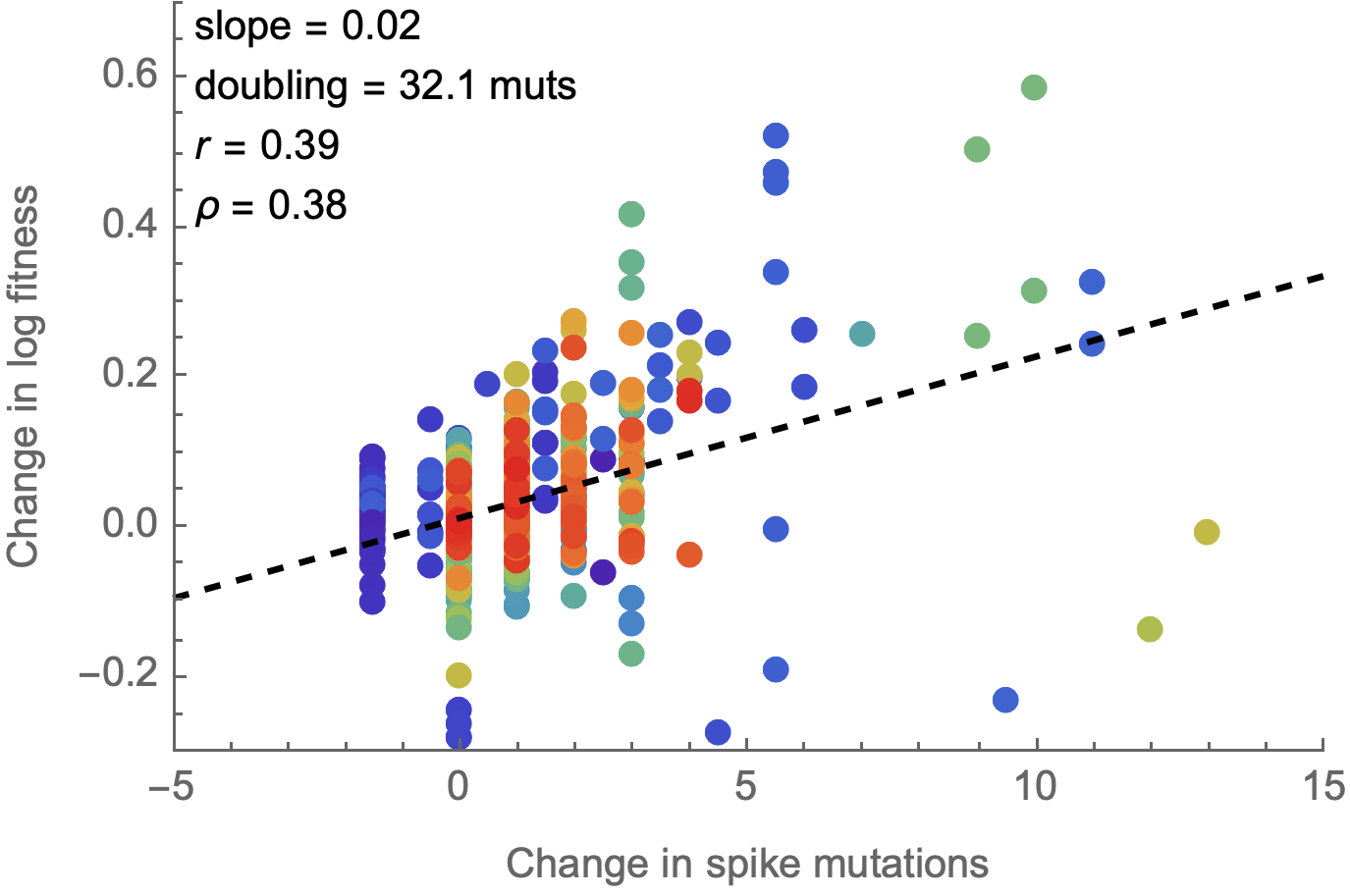
Spike mutations tend to increase fitness
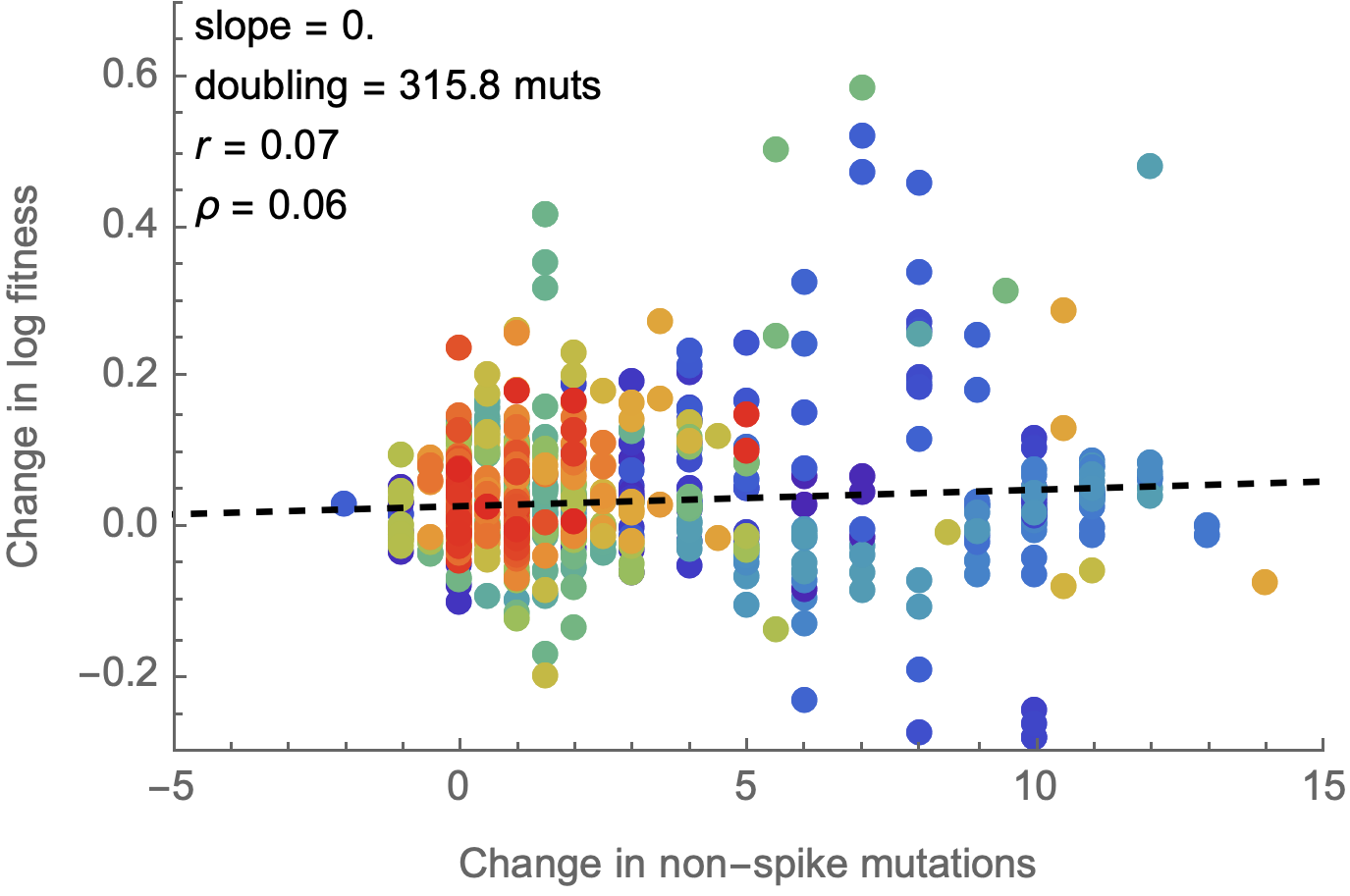
Non-spike mutations do not impact fitness on average
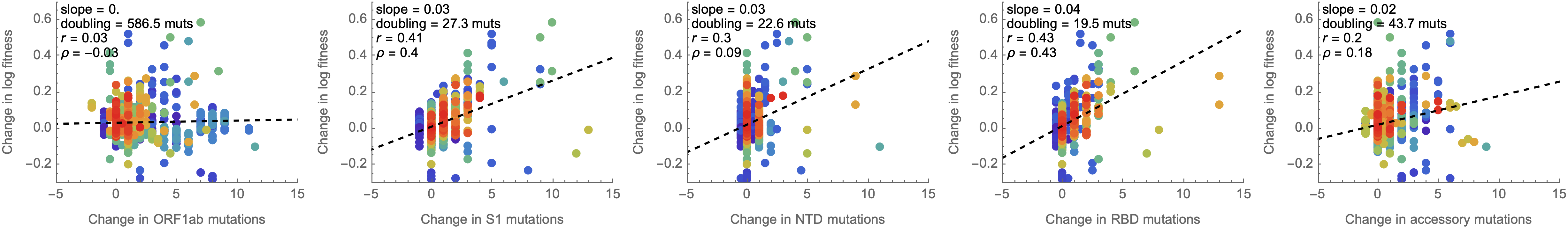
Looking across the genome shows that spike is the focus for positive selection, but accessory genes have some signal

Some attenuation of fitness effects over time
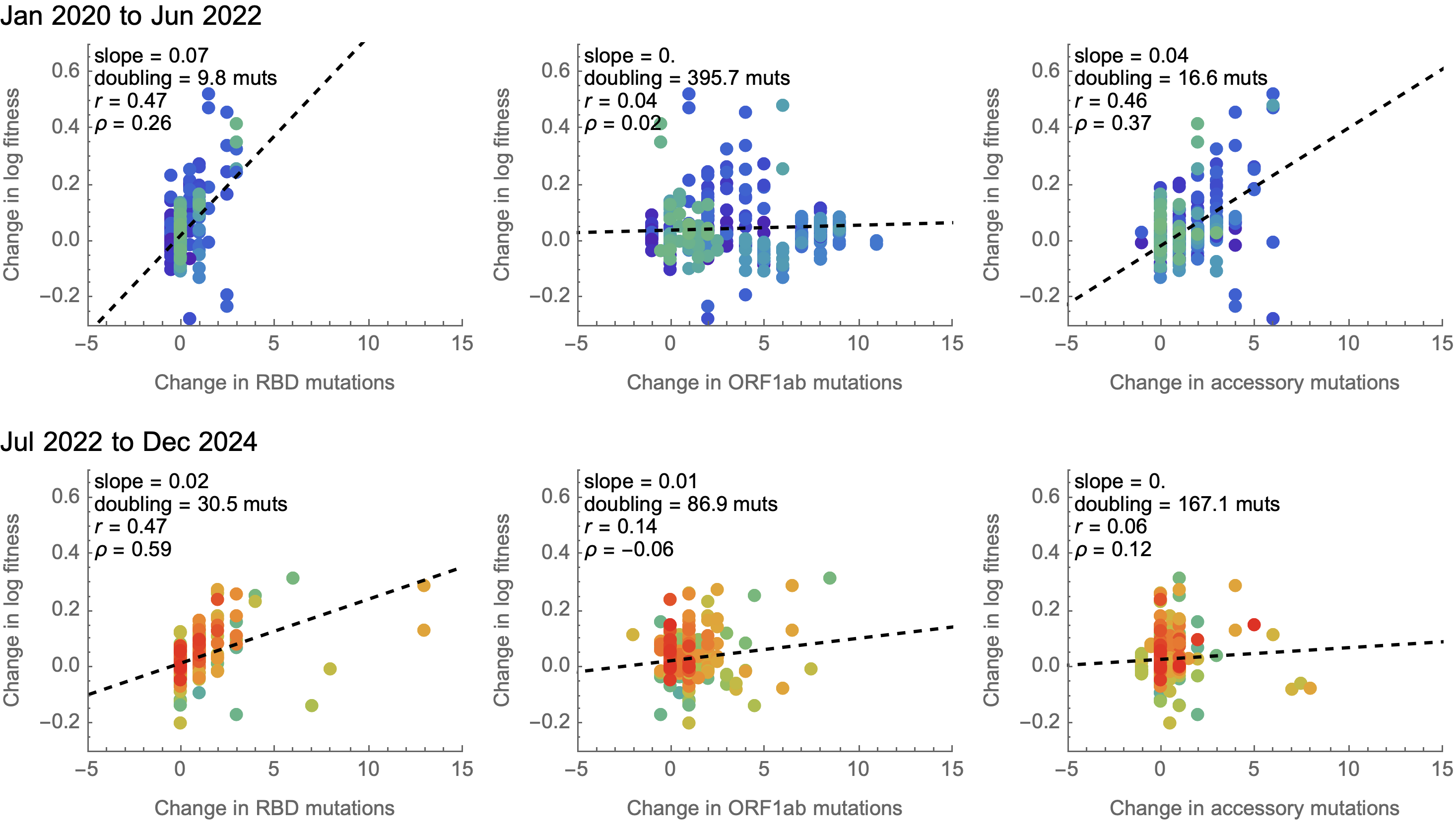
Predicting fitness effects
Simple linear model to combine predictors into a fitness estimate
In this case, a very simple model of mutation counts
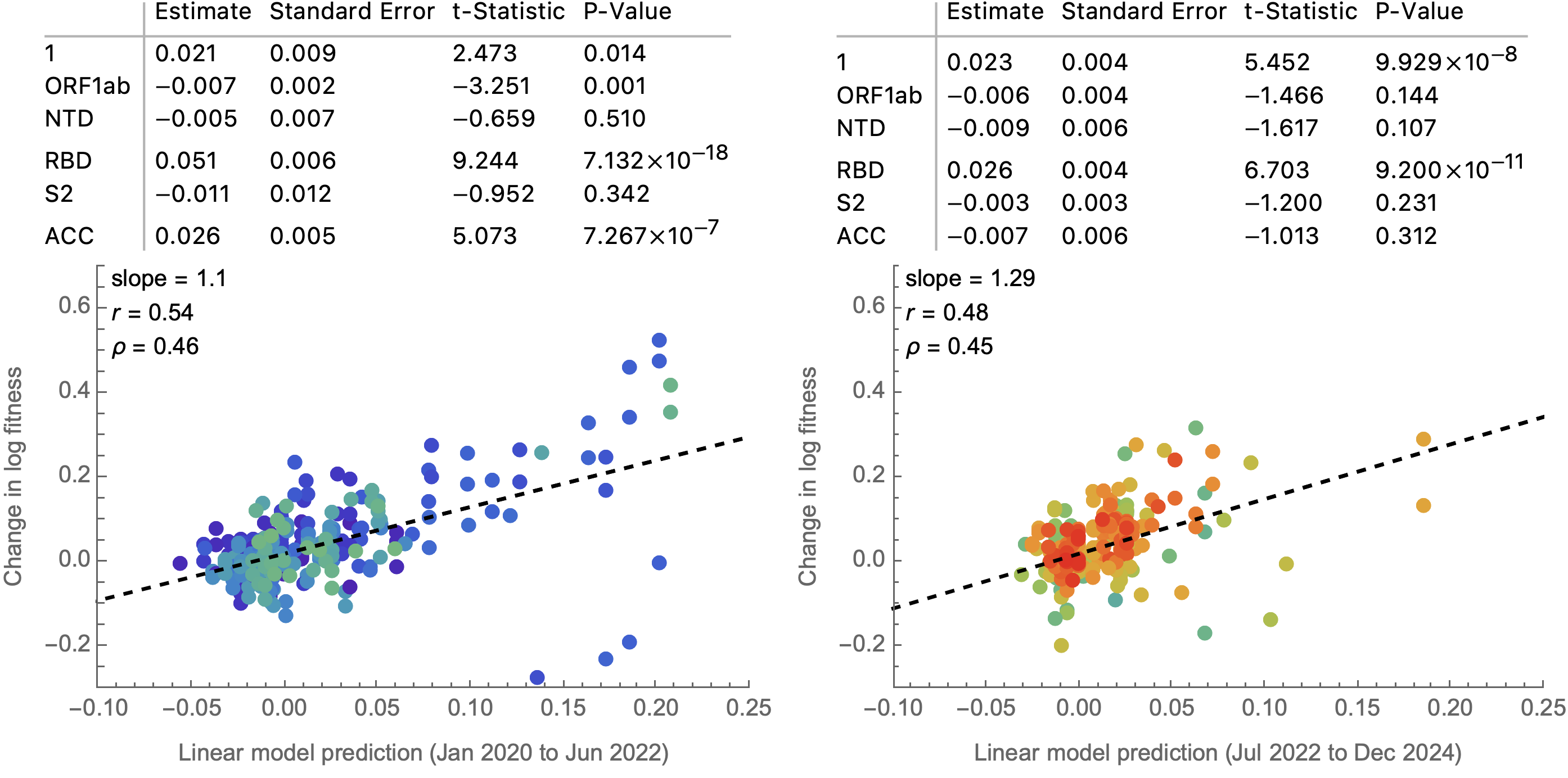
Framework to compare predictors of fitness
EvEscape combines a variational autoencoder for mutation effect + antibody accessibility + biochemical dissimilarity
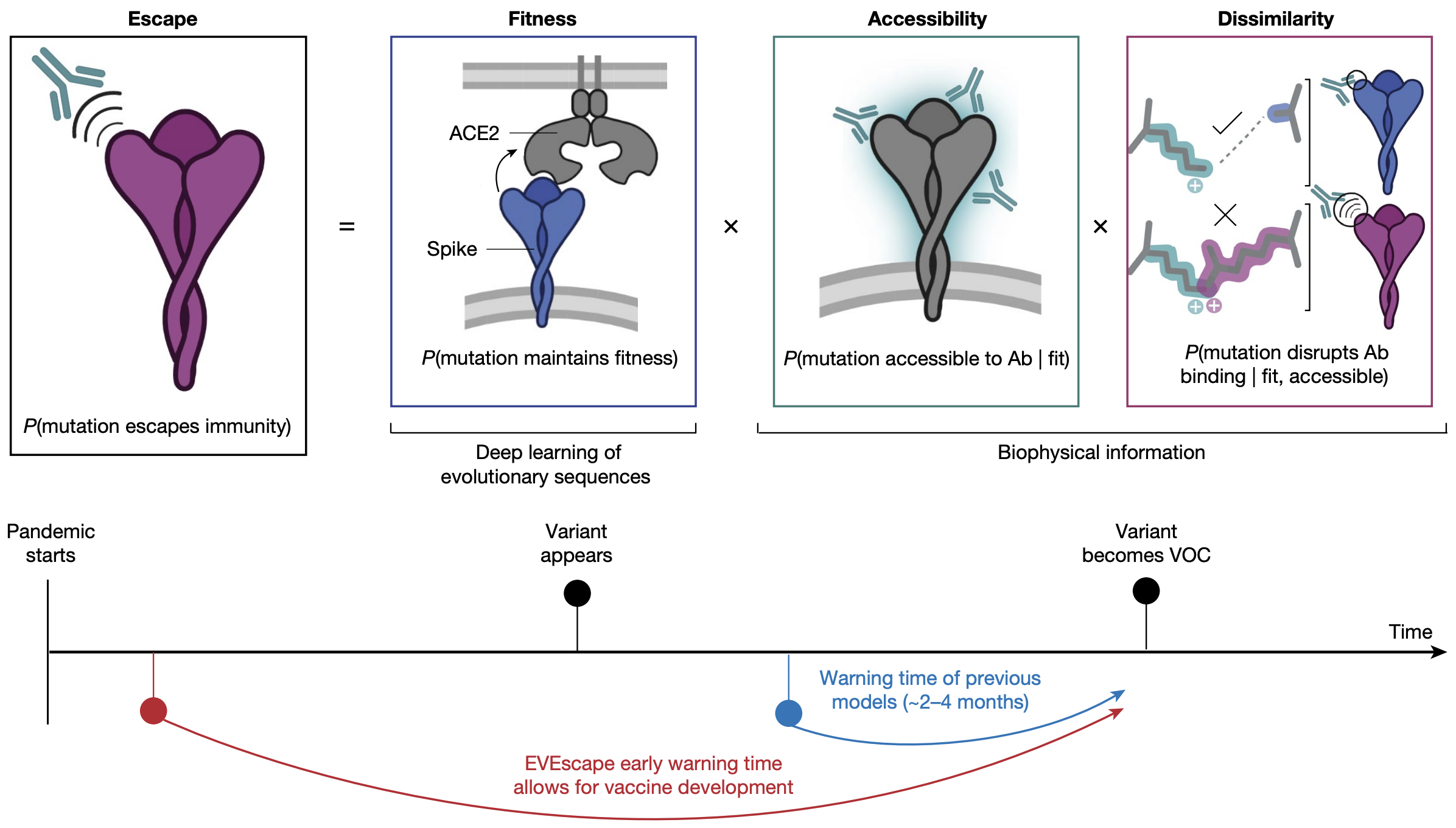 EvEscape from Thadani et al. 2023. Nature.
EvEscape from Thadani et al. 2023. Nature.
EvEscape does no better than counting spike mutations
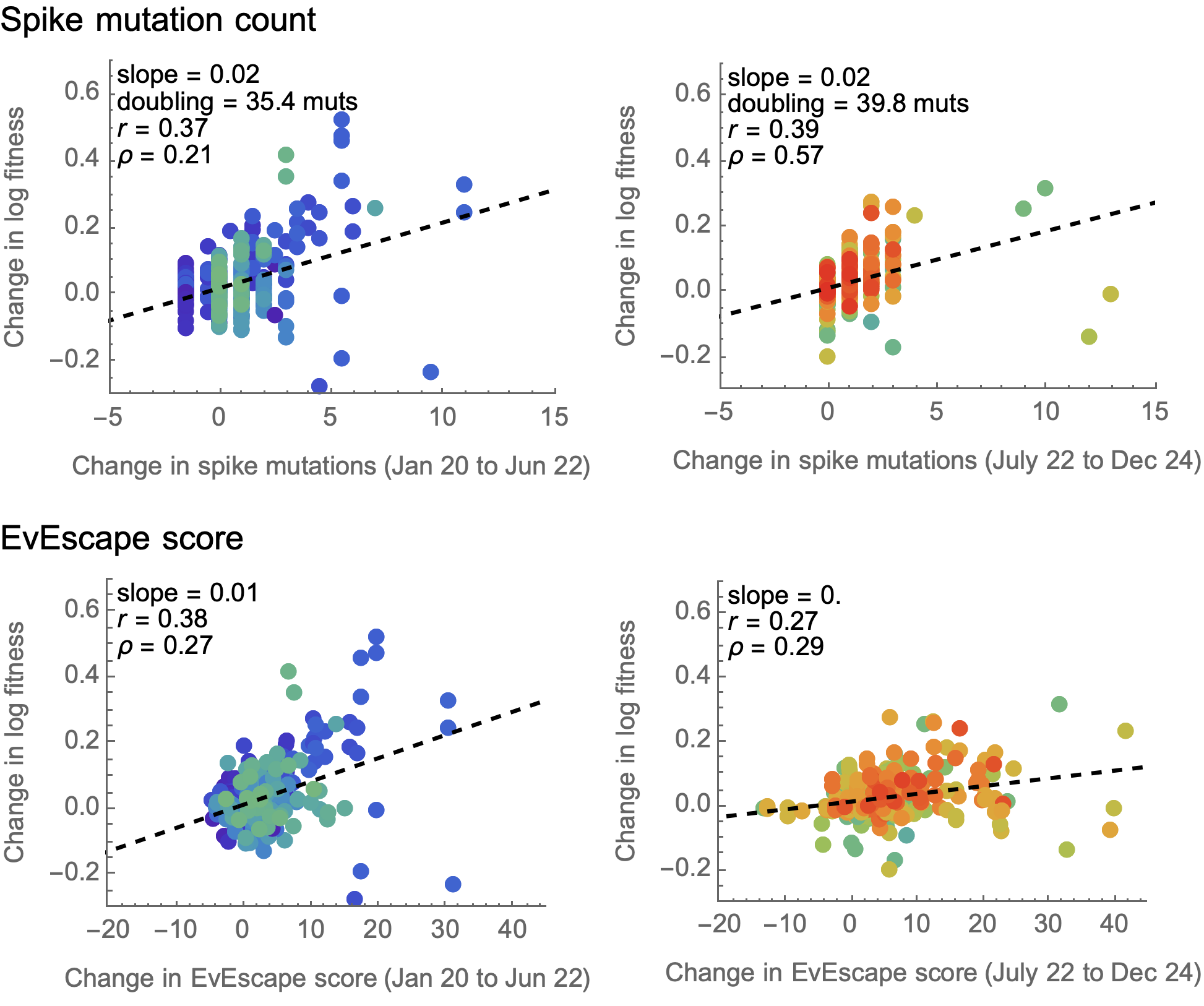 EvEscape from Thadani et al. 2023. Nature.
EvEscape from Thadani et al. 2023. Nature.
CoVFit uses spike protein embeddings from ESM-2 to predict MLR fitnesses
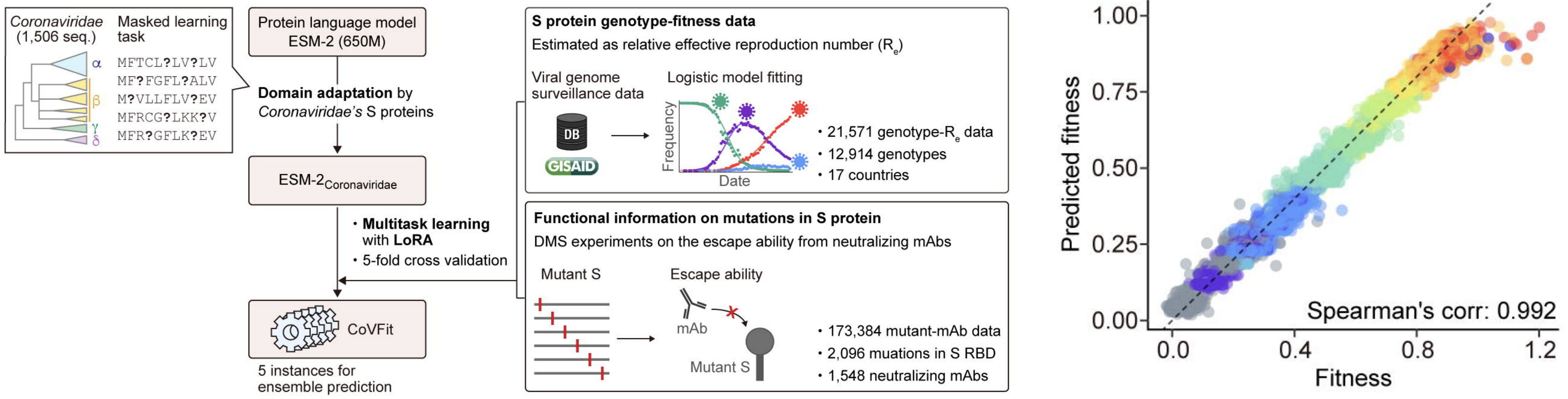 CoVFit from Ito et al. 2025. Nat Commun.
CoVFit from Ito et al. 2025. Nat Commun.
CoVFit performs poorly outside of training window
Note also the importance of properly assessing independent parent/child lineage deltas
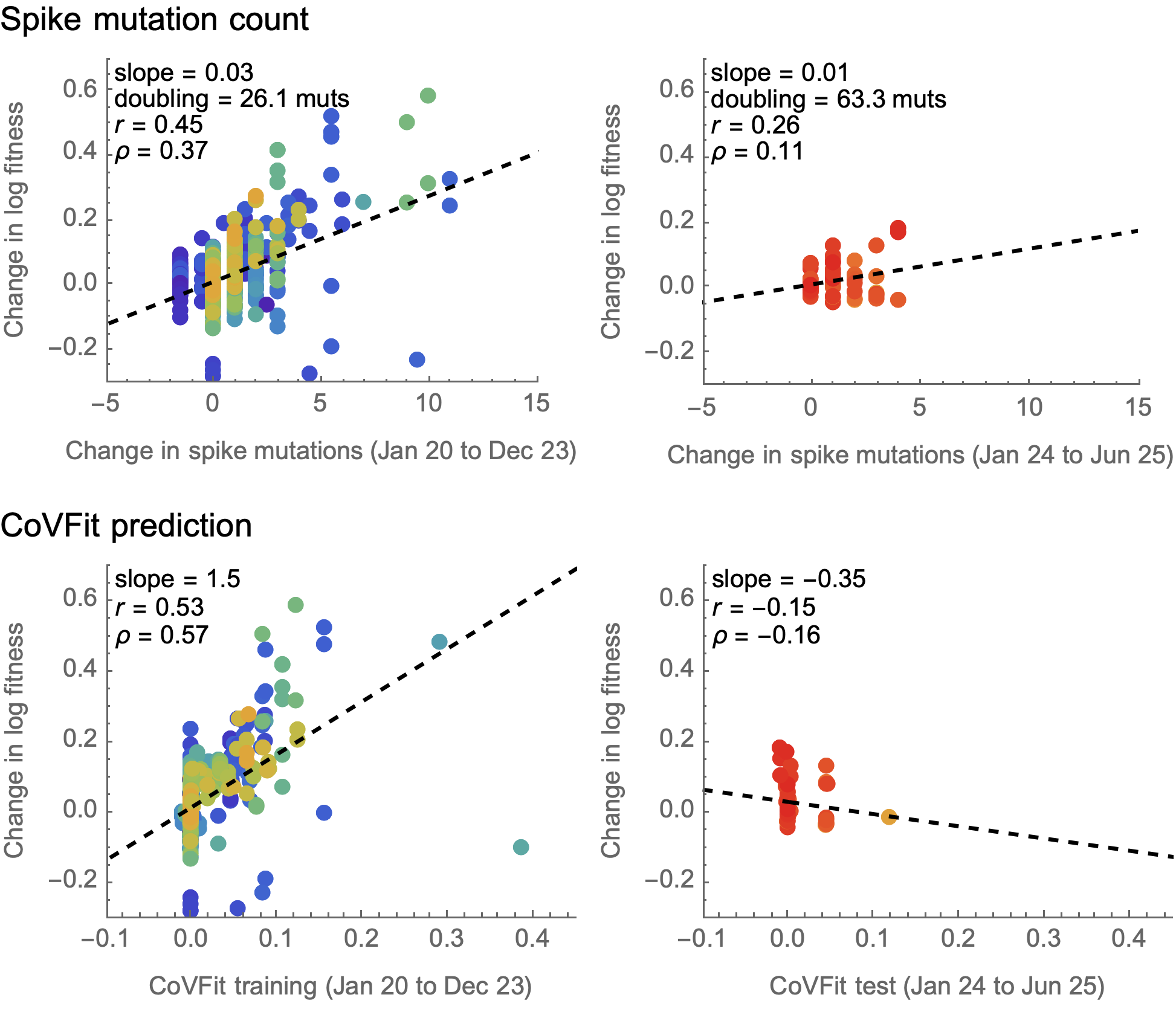 CoVFit from Ito et al. 2025. Nat Commun.
CoVFit from Ito et al. 2025. Nat Commun.
Semanticity to predict immune escape via dissimilarity of embeddings
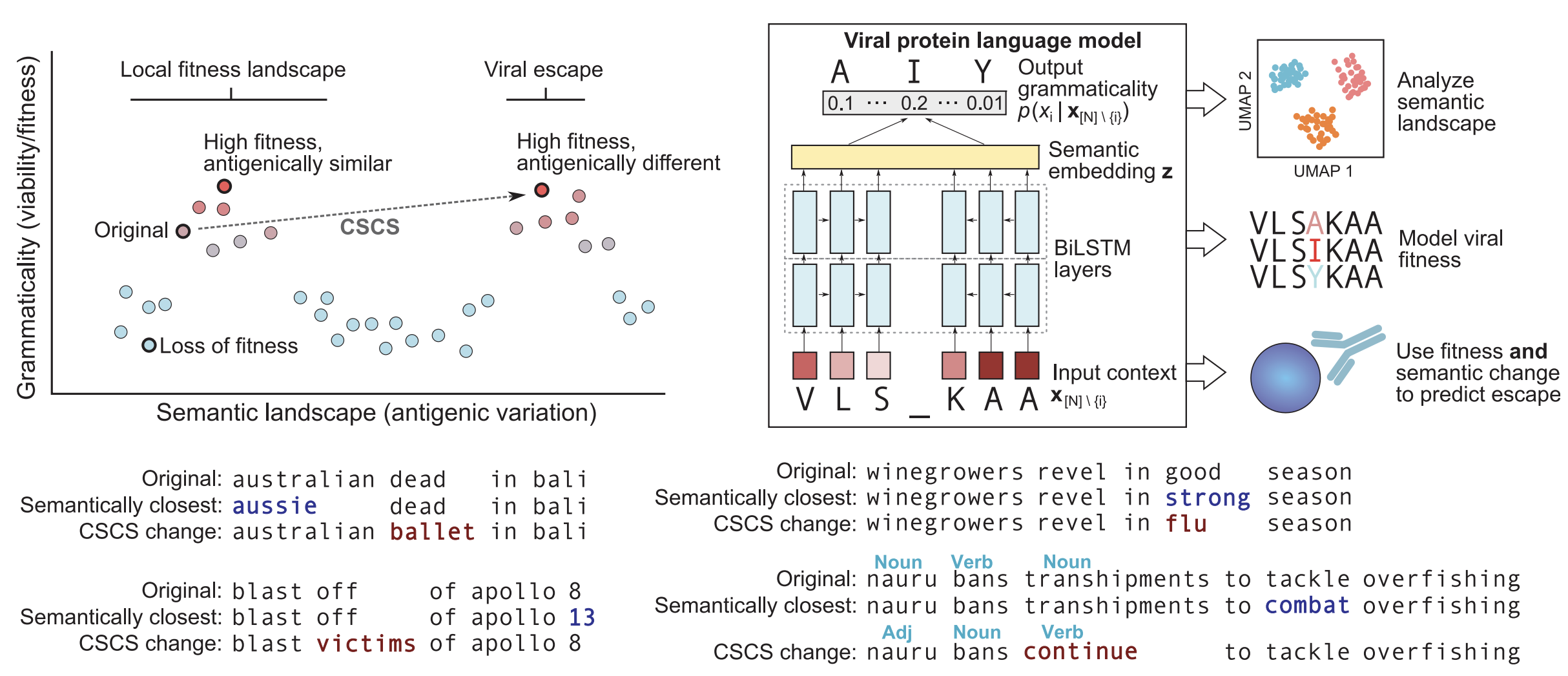 Semanticity following Hie et al. 2021. Science.
Semanticity following Hie et al. 2021. Science.
Re-implemented semanticity metric in ESM-2 via CLS token embedding
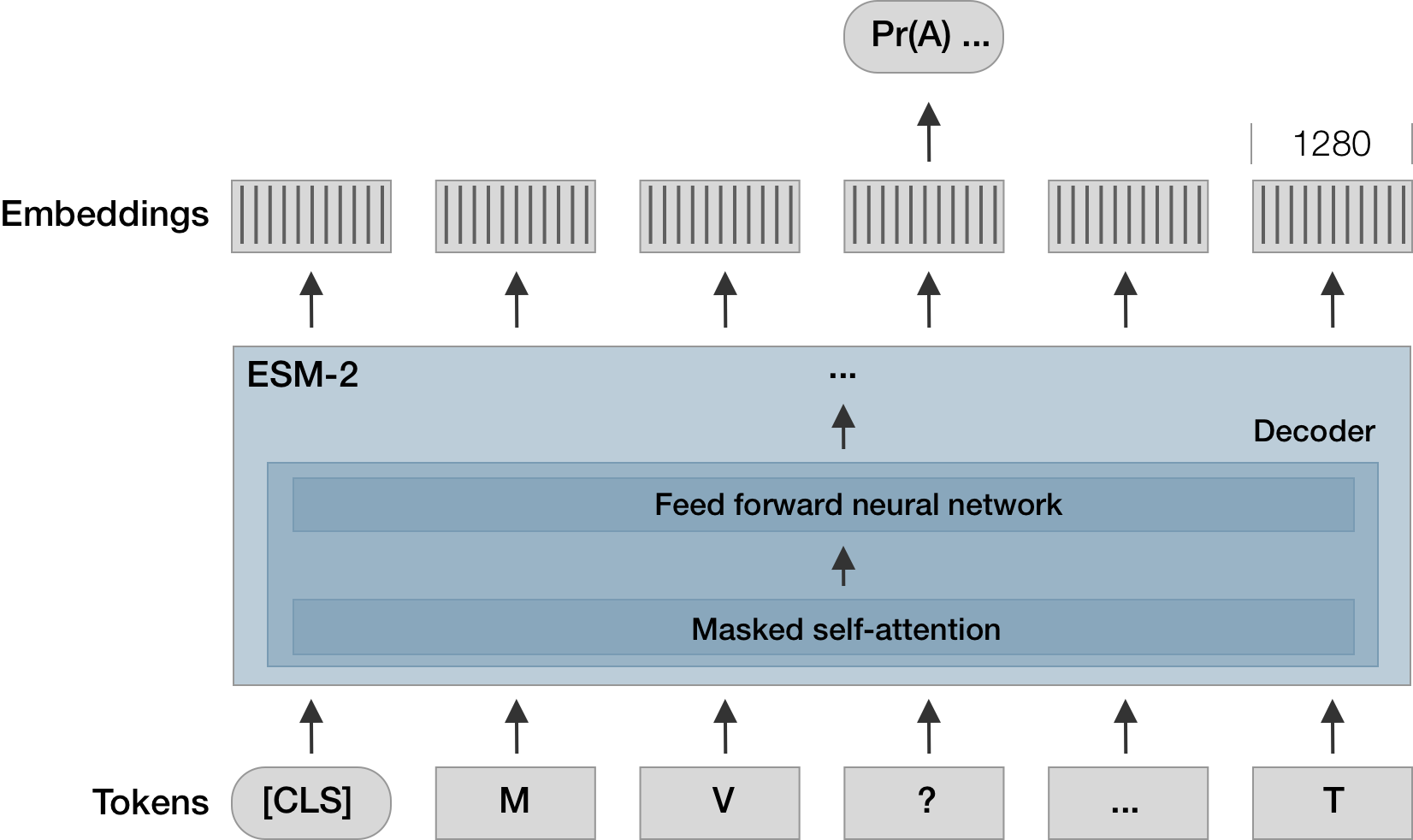 Semanticity following Hie et al. 2021. Science.
ESM-2 from Rives et al. 2021. PNAS.
Semanticity following Hie et al. 2021. Science.
ESM-2 from Rives et al. 2021. PNAS.
Semantic dissimilarity does no better than counting spike mutations
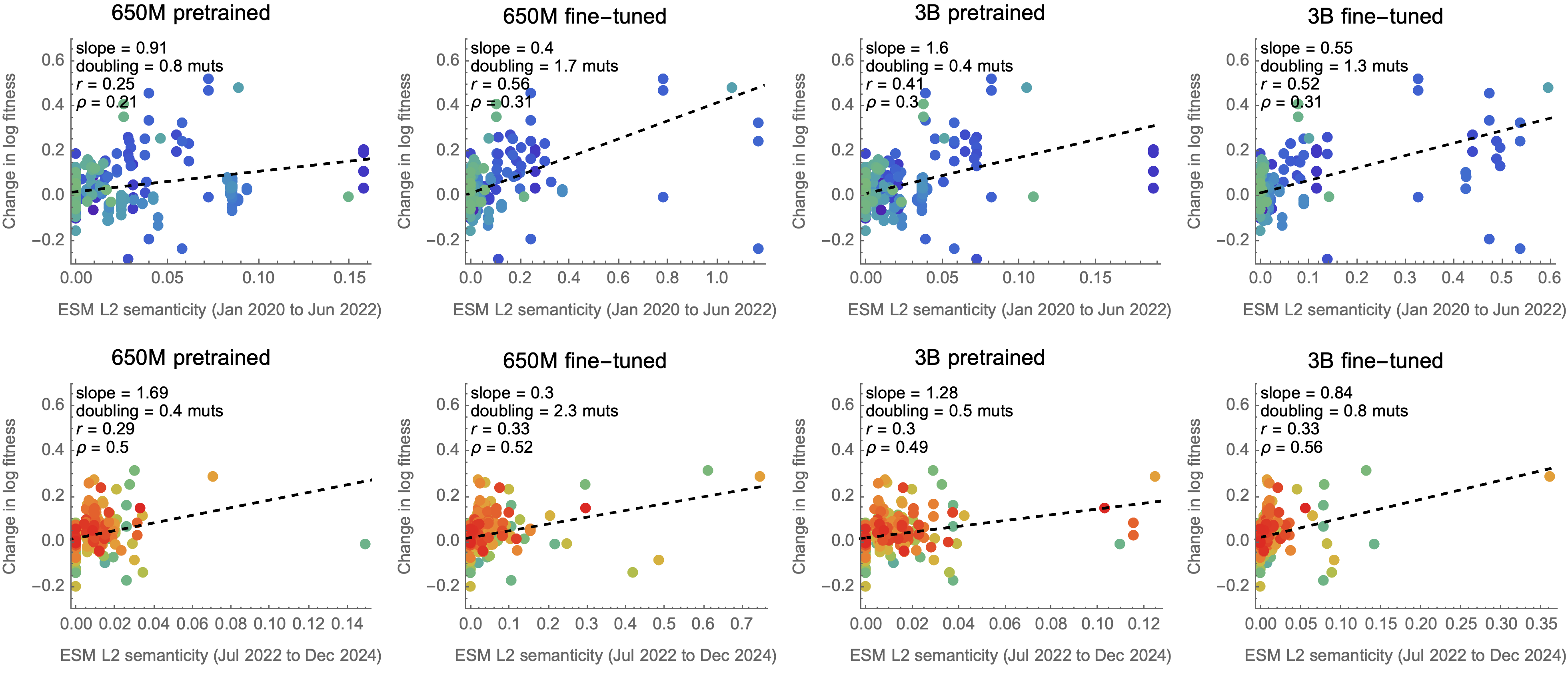 Semanticity following Hie et al. 2021. Science.
ESM-2 from Rives et al. 2021. PNAS.
Semanticity following Hie et al. 2021. Science.
ESM-2 from Rives et al. 2021. PNAS.
High-throughput experimental measurements of ACE2 binding and immune escape
But even these DMS measurements have short time horizons
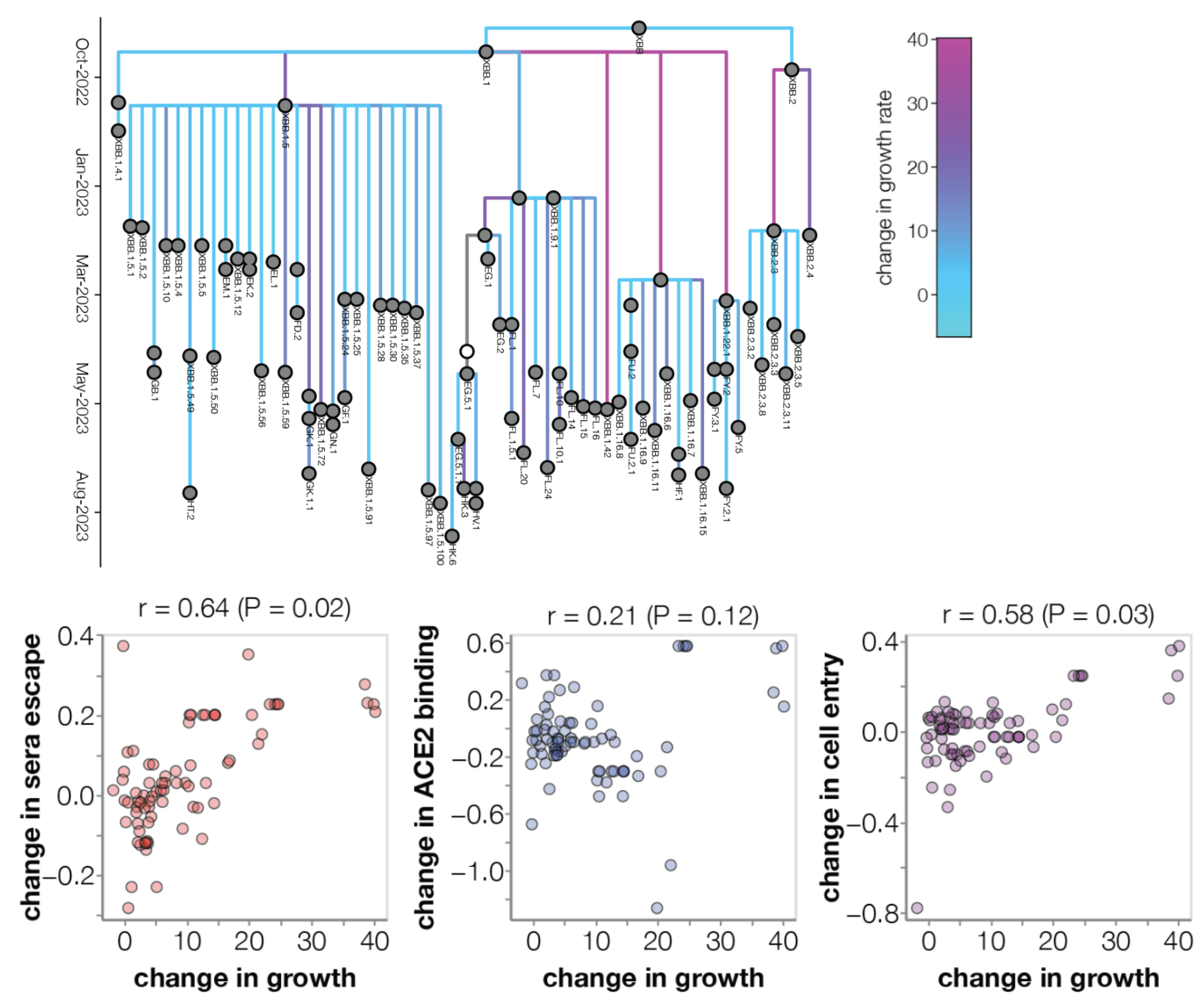
Even with excellent out-of-sample fitness prediction, we still need approaches that explicitly model mutations and emergence of novel lineages
Acknowledgements
Seasonal influenza and SARS-CoV-2 genomics: Data producers from all over the world, GISAID
Nextstrain: Richard Neher, Ivan Aksamentov, John SJ Anderson, Kim Andrews, Jennifer Chang, James Hadfield, Emma Hodcroft, John Huddleston, Jover Lee, Victor Lin, Cornelius Roemer, Thomas Sibley
MLR and fitness modeling: Marlin Figgins, Eslam Abousamra, Jover Lee, James Hadfield, John Huddleston, Philippa Steinberg, Jesse Bloom, Cornelius Roemer, Richard Neher
Bedford Lab:
![]() John Huddleston,
John Huddleston,
![]() James Hadfield,
James Hadfield,
![]() Katie Kistler,
Katie Kistler,
![]() Jover Lee,
Jover Lee,
![]() Marlin Figgins,
Marlin Figgins,
![]() Victor Lin,
Victor Lin,
![]() Nashwa Ahmed,
Nashwa Ahmed,
![]() Cécile Tran Kiem,
Cécile Tran Kiem,
![]() Philippa Steinberg,
Philippa Steinberg,
![]() Jacob Dodds,
Jacob Dodds,
![]() Amin Bemanian,
Amin Bemanian,
![]() Carlos Avendano,
Carlos Avendano,
![]() Aayush Verma
Aayush Verma




