Evolutionary forecasting for SARS-CoV-2
Trevor Bedford
Fred Hutchinson Cancer Center / Howard Hughes Medical Institute
11 Jul 2023
Monthly Meeting
Northwest PGCoE
Slides at: bedford.io/talks
SARS-CoV-2 continues to show remarkable capacity for evolution
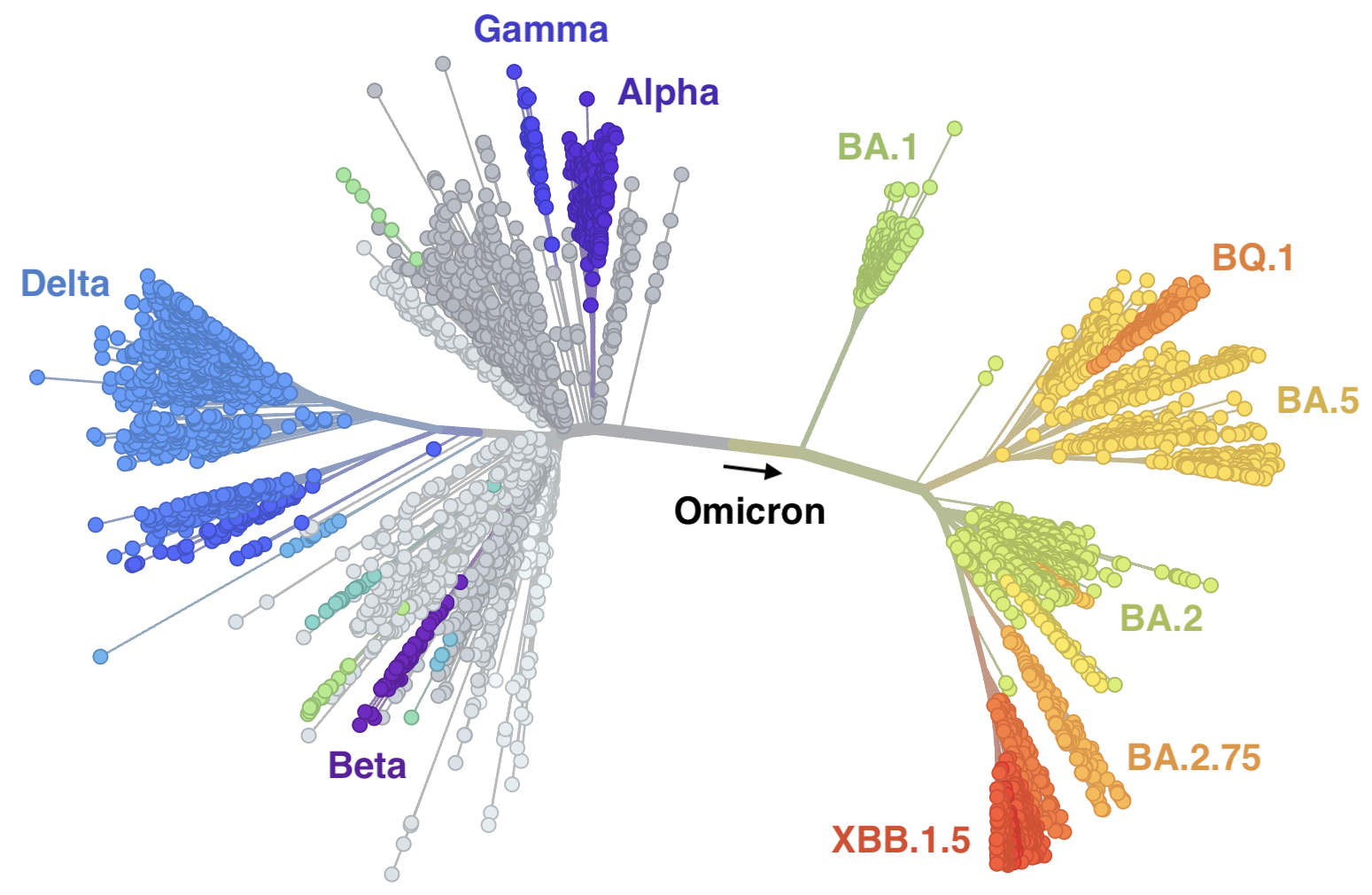
Mutations at spike S1 propel escape from population immunity
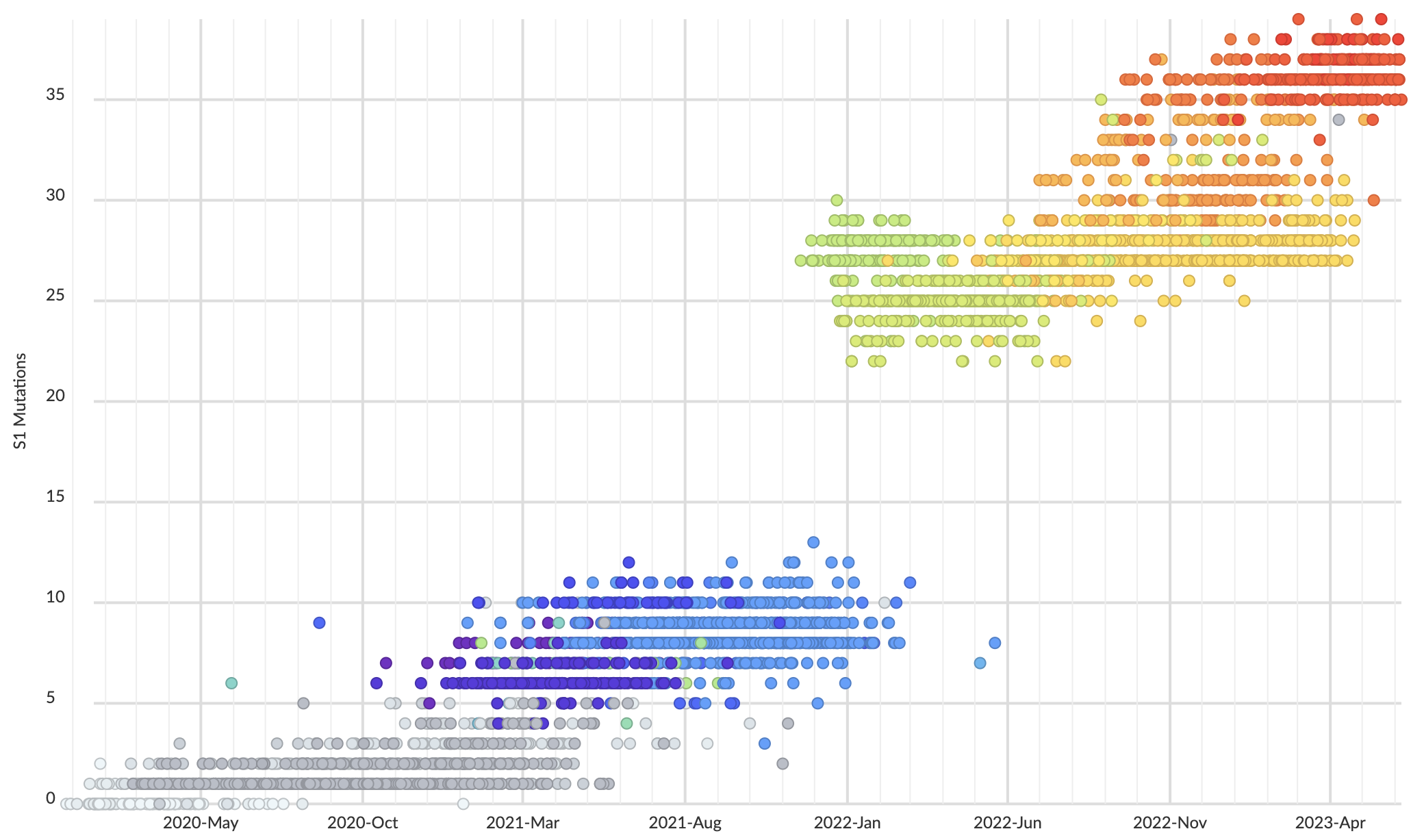
These mutations are accruing much more rapidly than other endemic viruses
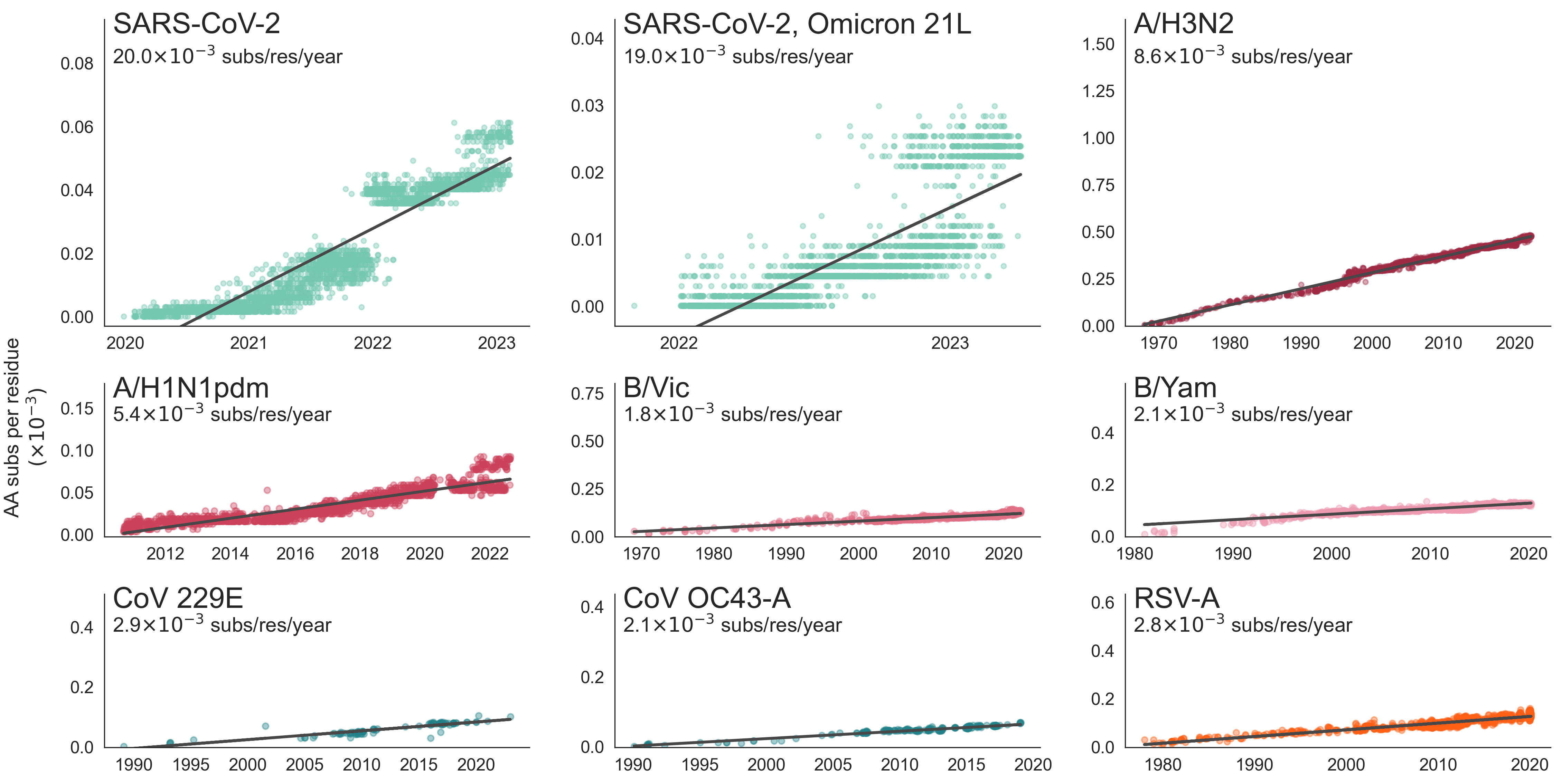
New variants emerge that escape from existing population immunity and spread rapidly
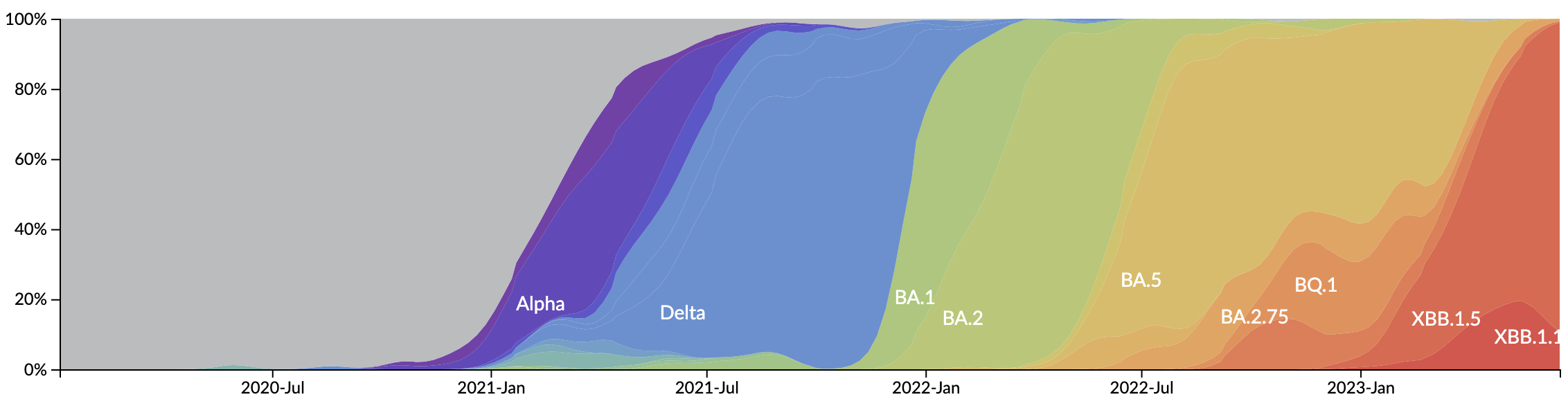
Variant frequency dynamics
Population genetic expectation of variant frequency under selection
$x' = \frac{x \, (1+s)}{x \, (1+s) + (1-x)}$ for frequency $x$ over one generation with selective advantage $s$
$x(t) = \frac{x_0 \, (1+s)^t}{x_0 \, (1+s)^t + (1-x_0)}$ for initial frequency $x_0$ over $t$ generations
Trajectories are linear once logit transformed via $\mathrm{log}(\frac{x}{1 - x})$
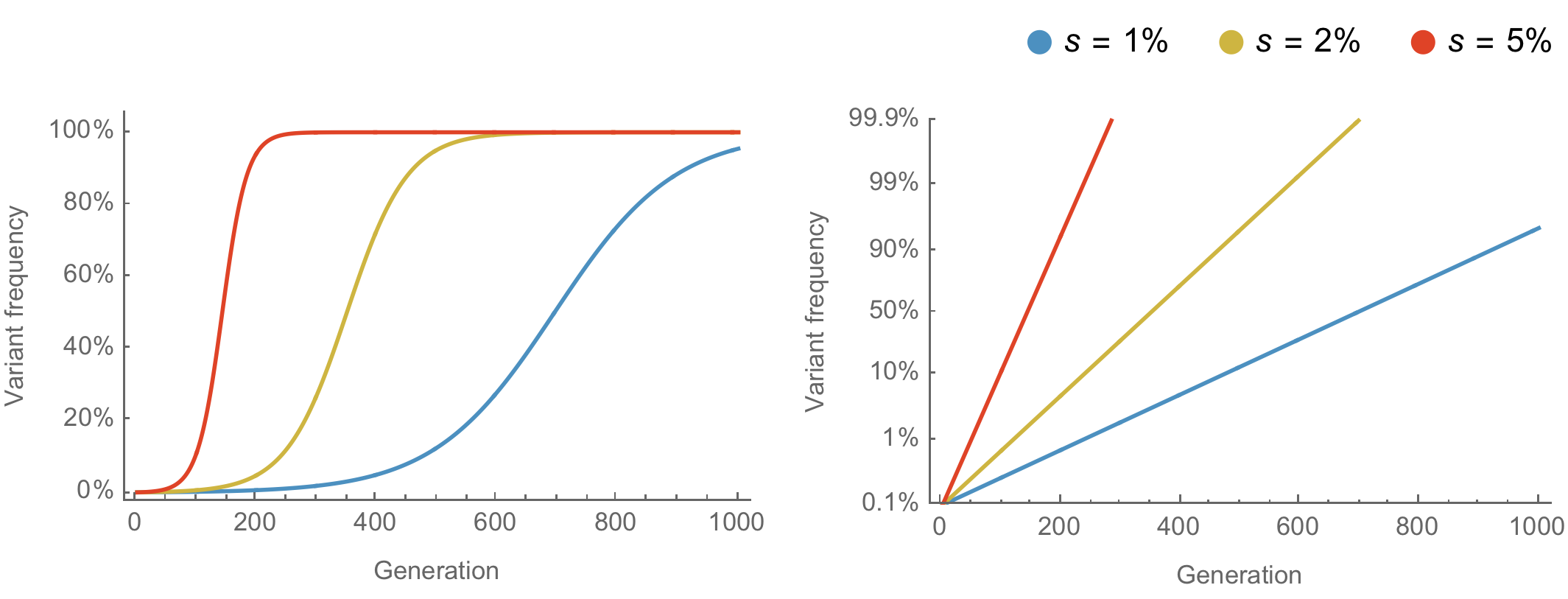
Variants show consistent frequency dynamics in logit space
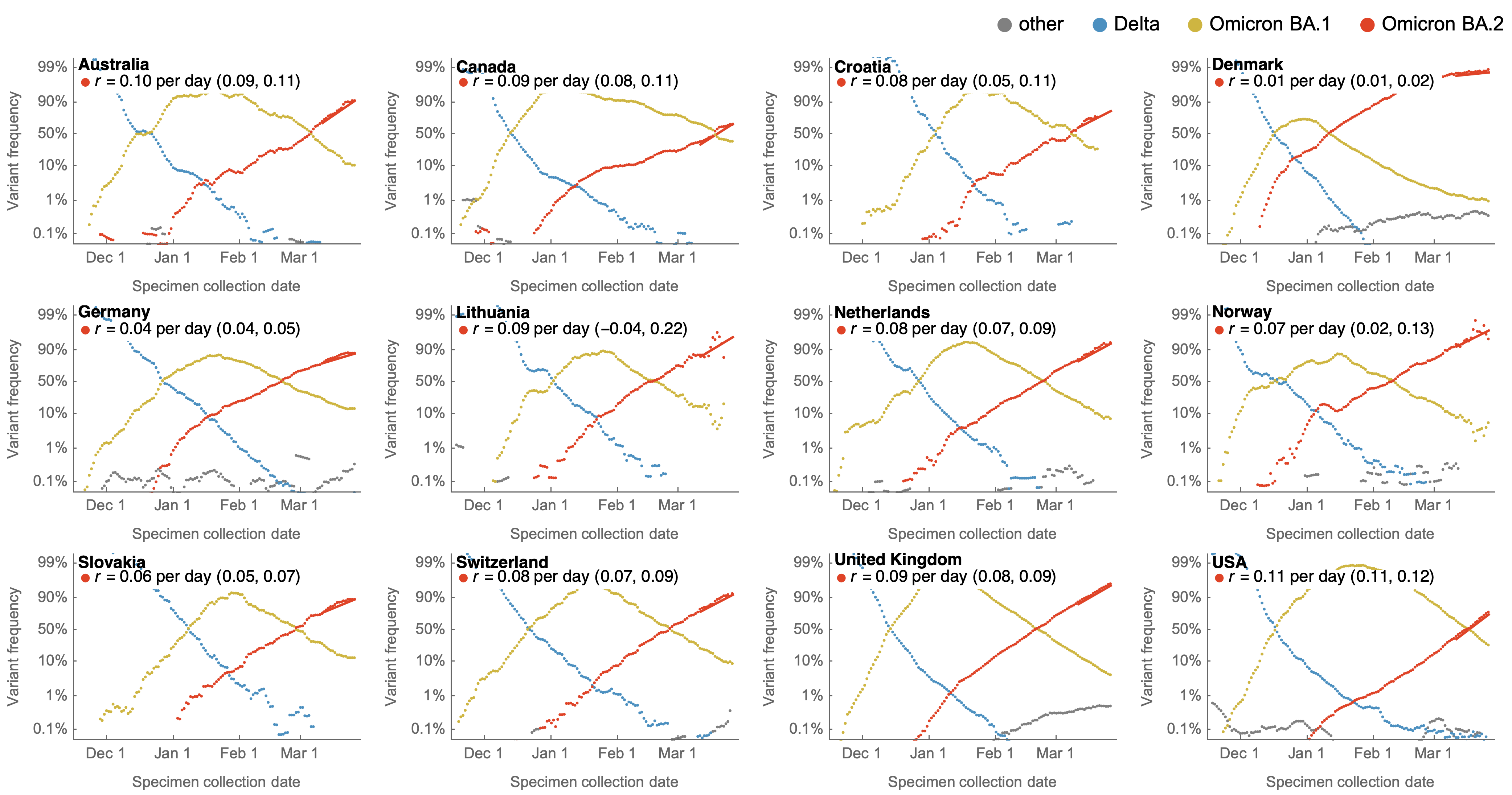
Variants show consistent frequency dynamics in logit space
Multinomial logistic regression
Multinomial logistic regression across $n$ variants models the probability of a virus sampled at time $t$ belonging to variant $i$ as
$$\mathrm{Pr}(X = i) = x_i(t) = \frac{p_i \, \mathrm{exp}(f_i \, t)}{\sum_{1 \le j \le n} p_j \, \mathrm{exp}(f_j \, t) }$$
with $2n$ parameters consisting of $p_i$ the frequency of variant $i$ at initial timepoint and $f_i$ the growth rate or fitness of variant $i$.
The model is fit to minimize "log loss" of predicted variant vs observed variant across observations in dataset.
MLR implemented in evofr package
location variant date sequences Japan 22B 2023-02-10 242 Japan 22B 2023-02-11 56 Japan 22B 2023-02-12 70 Japan 22E 2023-02-10 80 Japan 22E 2023-02-11 21 Japan 22E 2023-02-12 27 USA 22B 2023-02-10 41 USA 22B 2023-02-11 23 USA 22B 2023-02-12 23 USA 22E 2023-02-10 368 USA 22E 2023-02-11 236 USA 22E 2023-02-12 246 ...
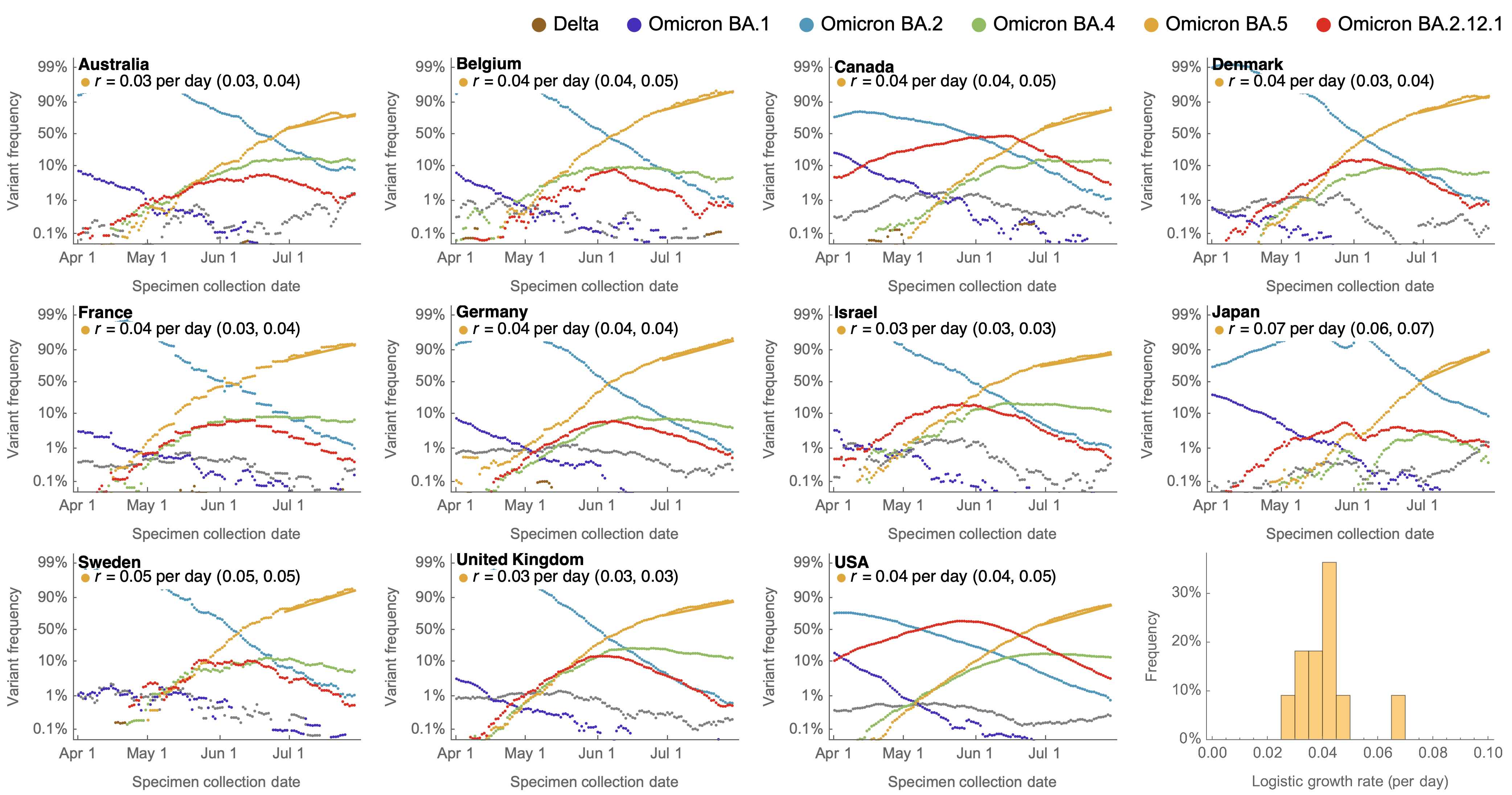
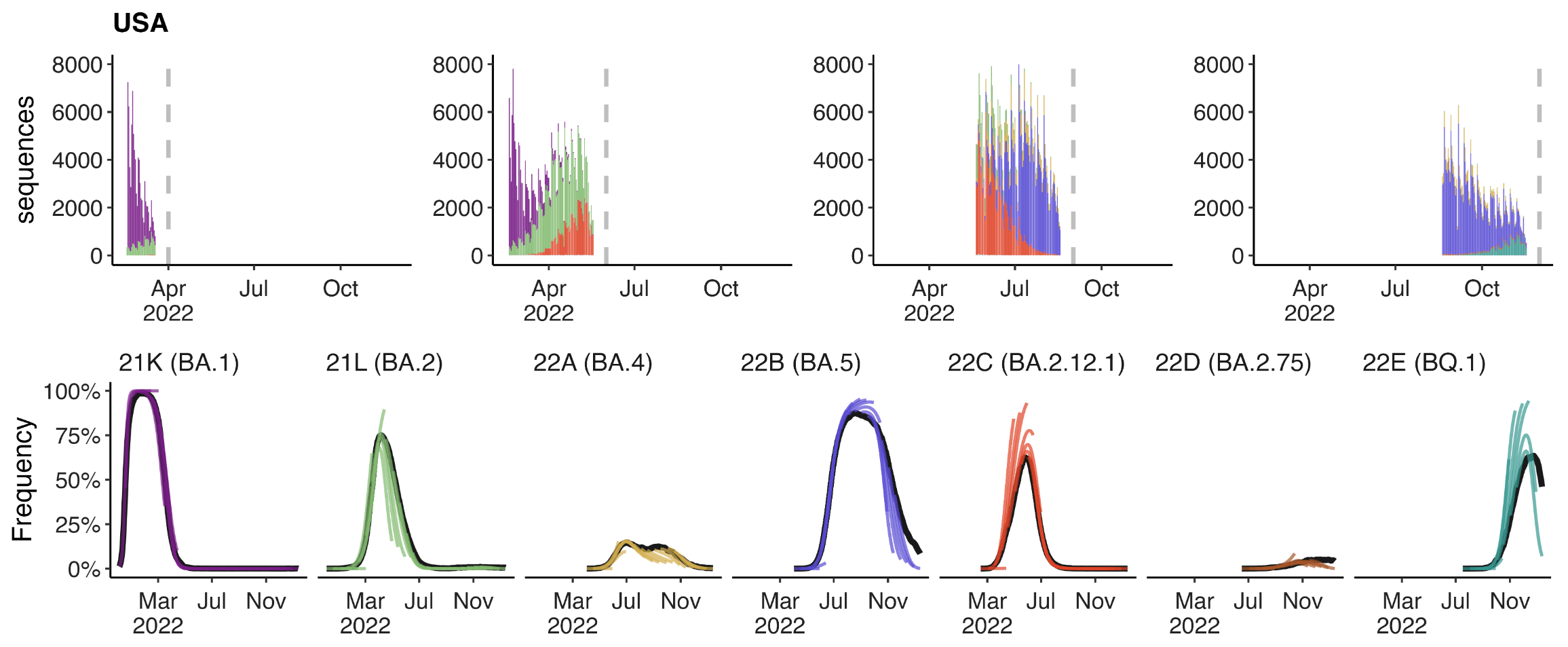
Multinomial logistic regression fits variant frequencies well
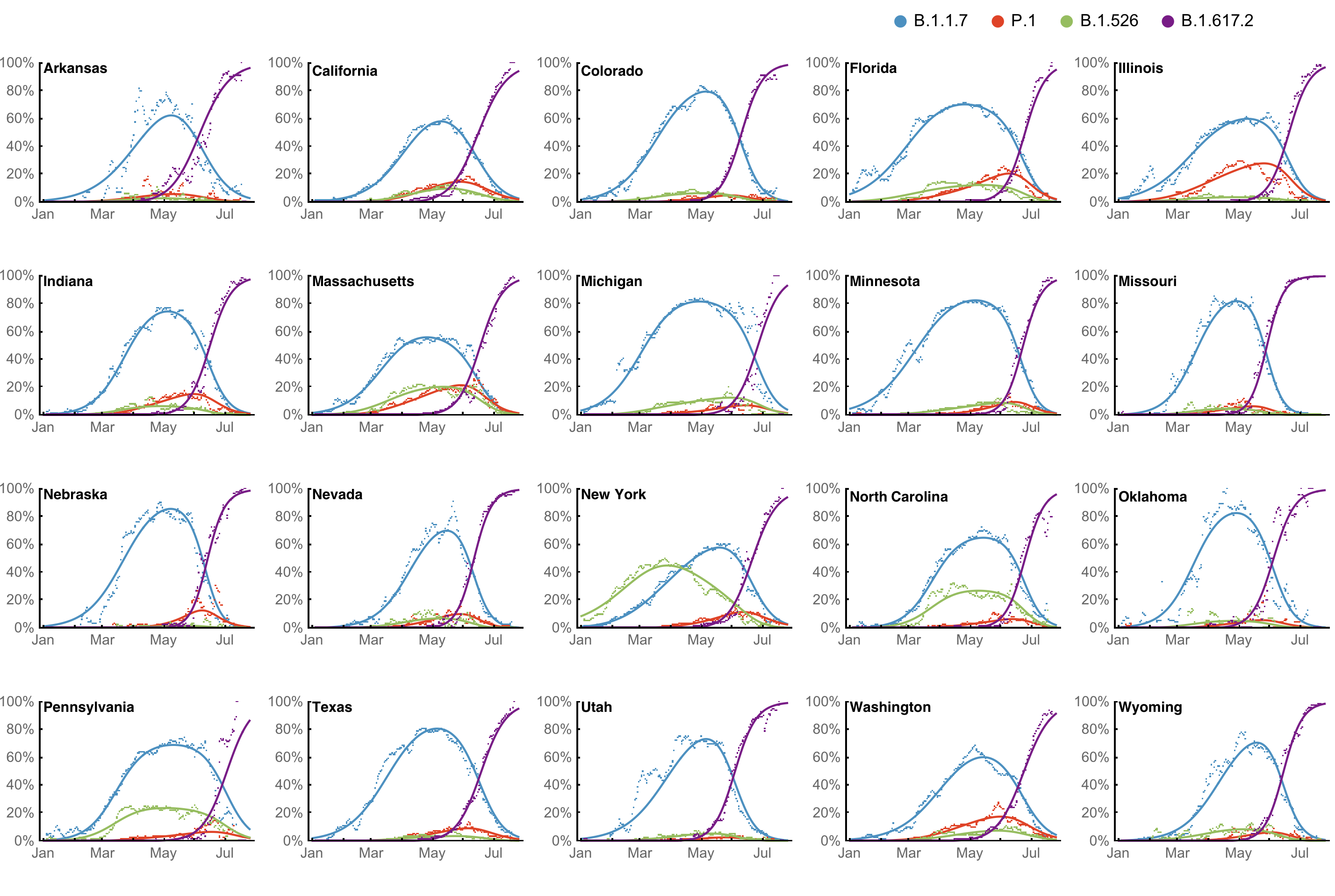
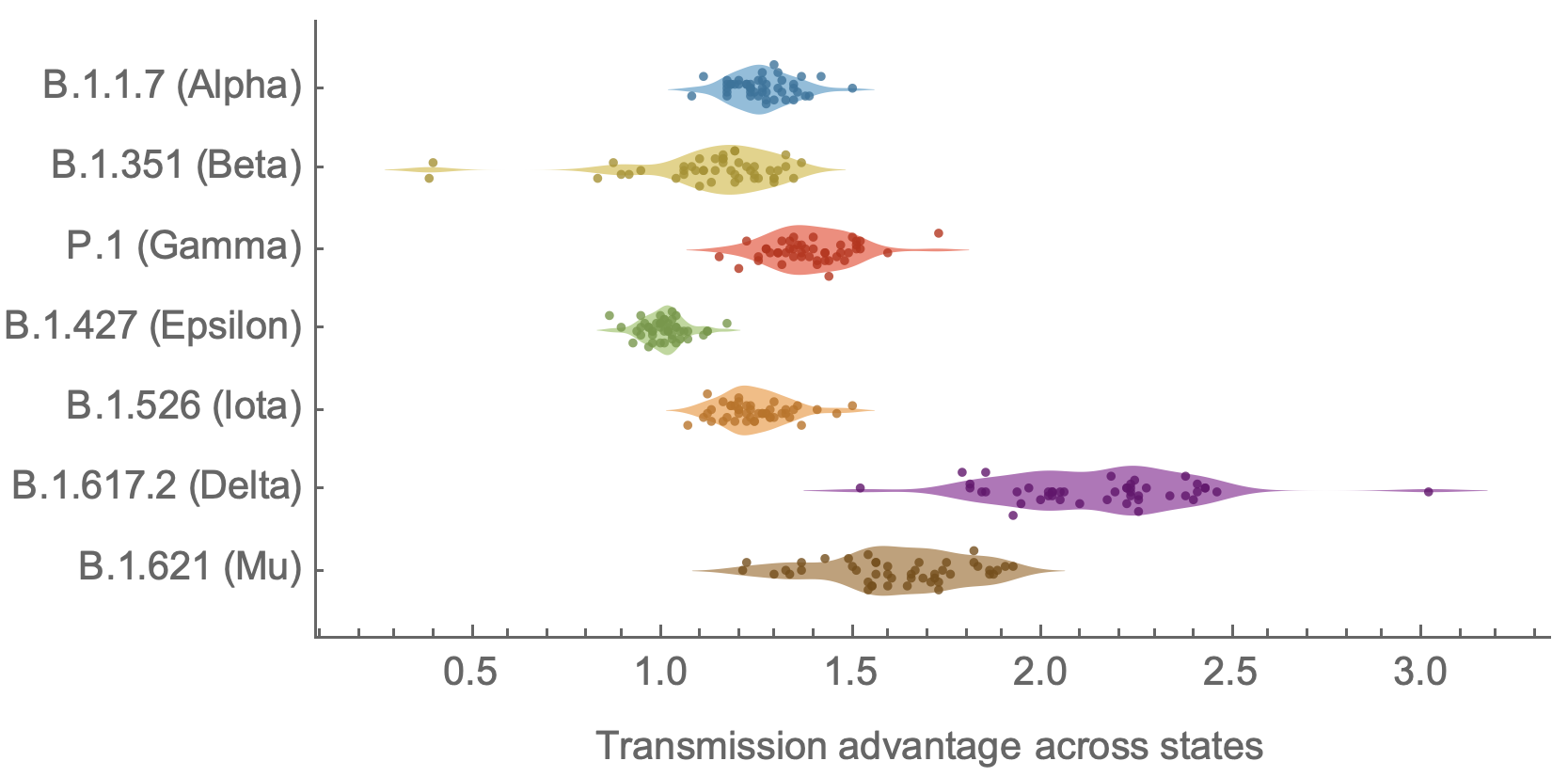
Original VOC viruses had substantially increased transmissibility
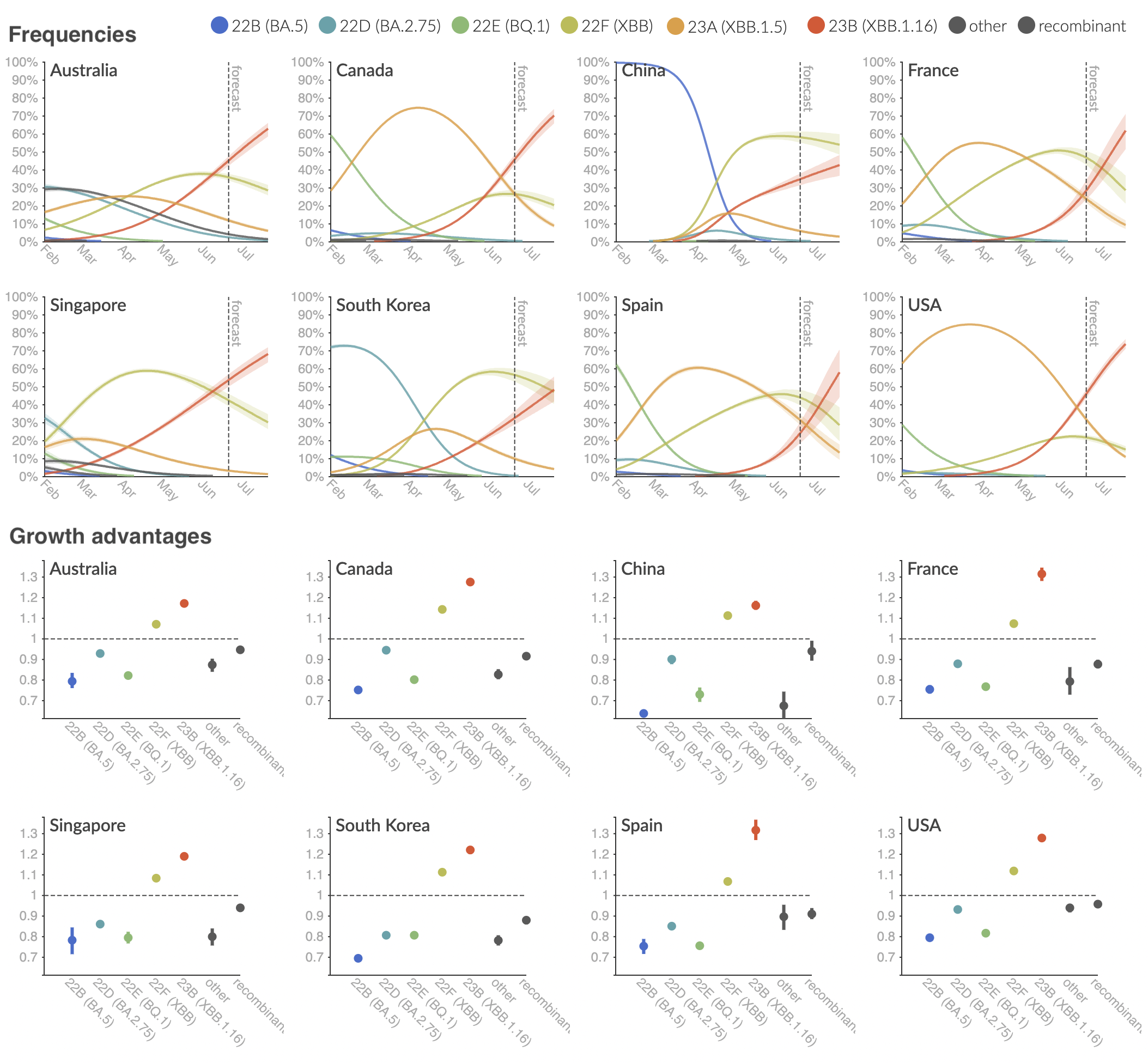
Variant frequencies across countries from Feb 2022 to present
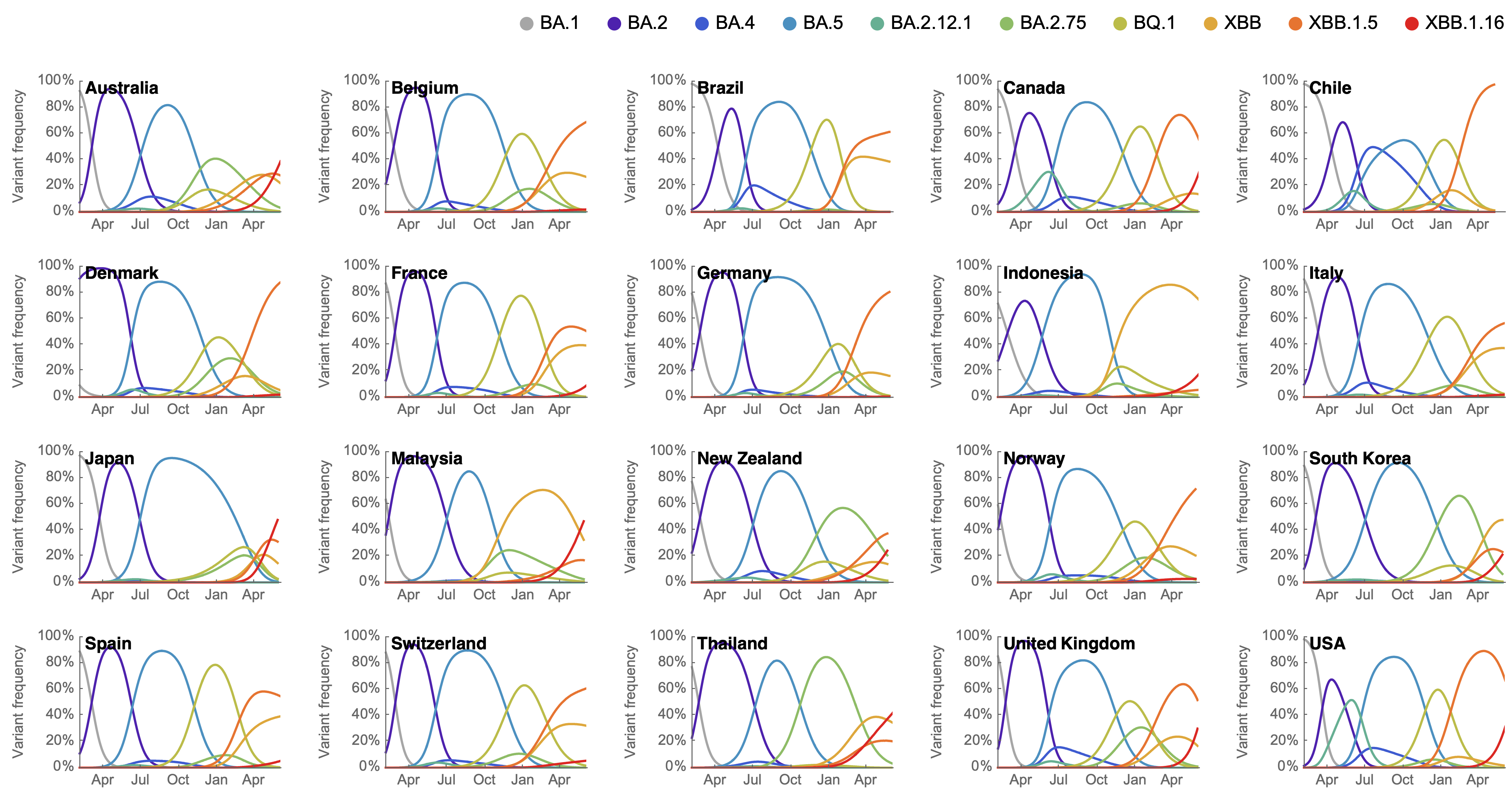
We find that recent variants like XBB.1.5 are ~300% fitter than original Omicron BA.1
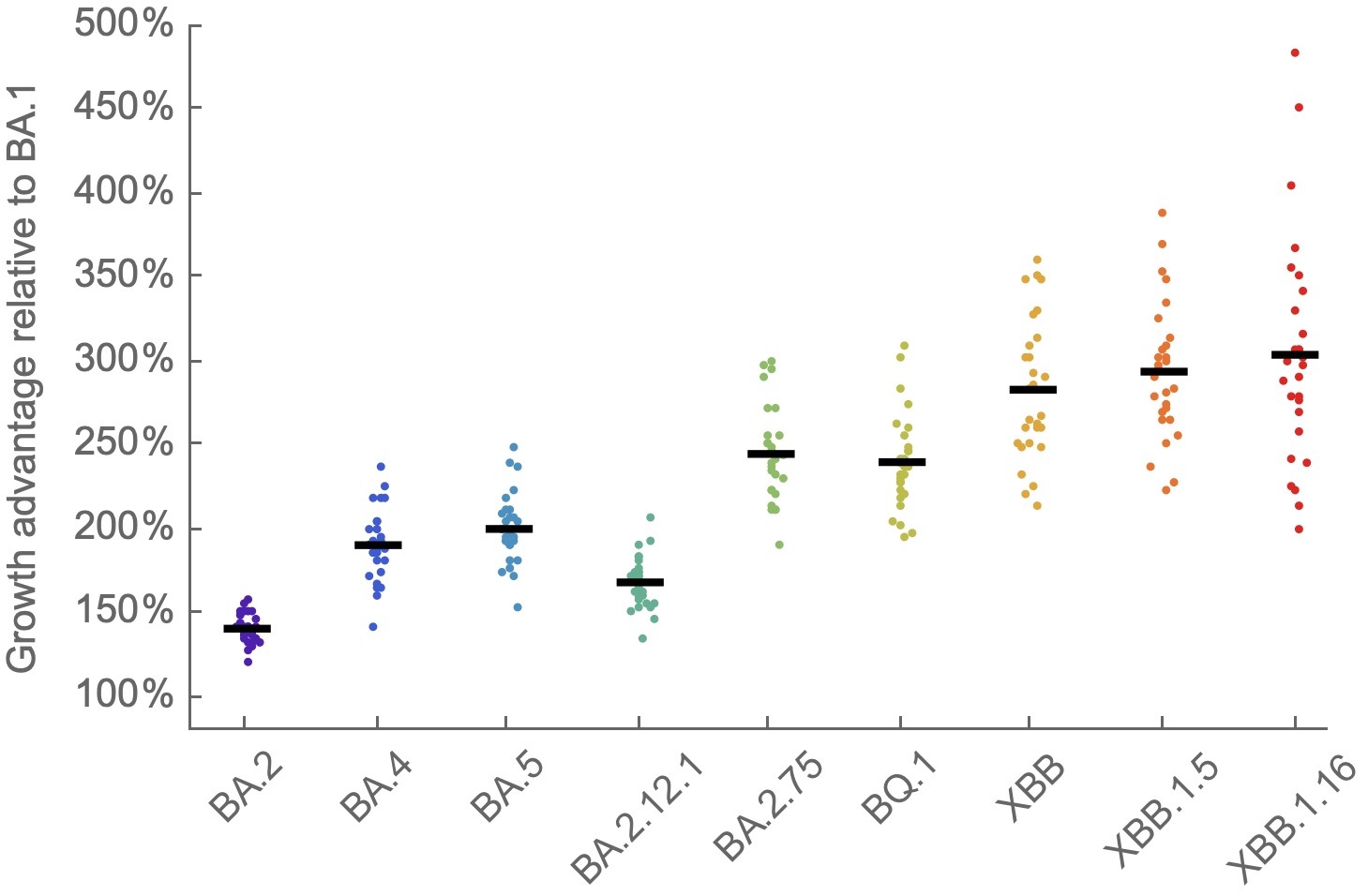
Evolution driving epidemics
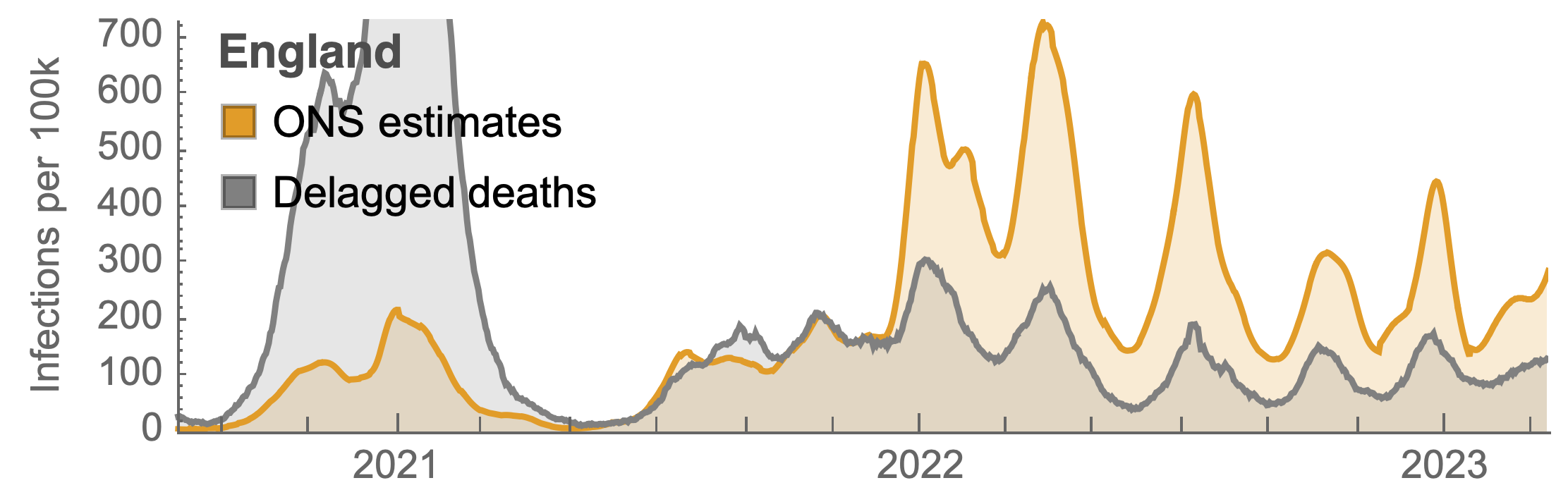
Many fewer reported cases in England post-Omicron
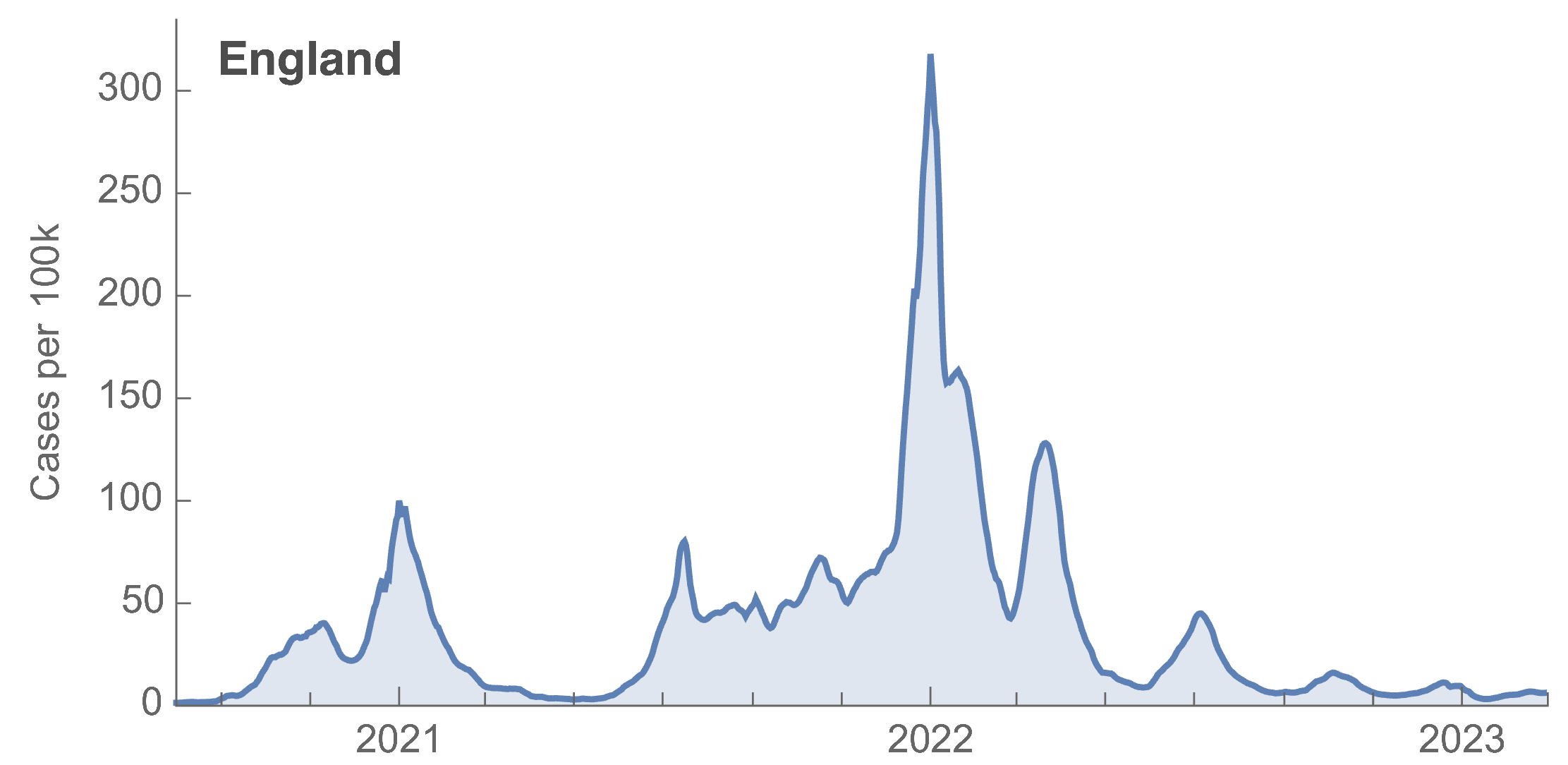
ONS Infection Survey provides rare source of ground truth
Roughly 1 in 3 infections detected in 2021, while 1 in 40 in 2023
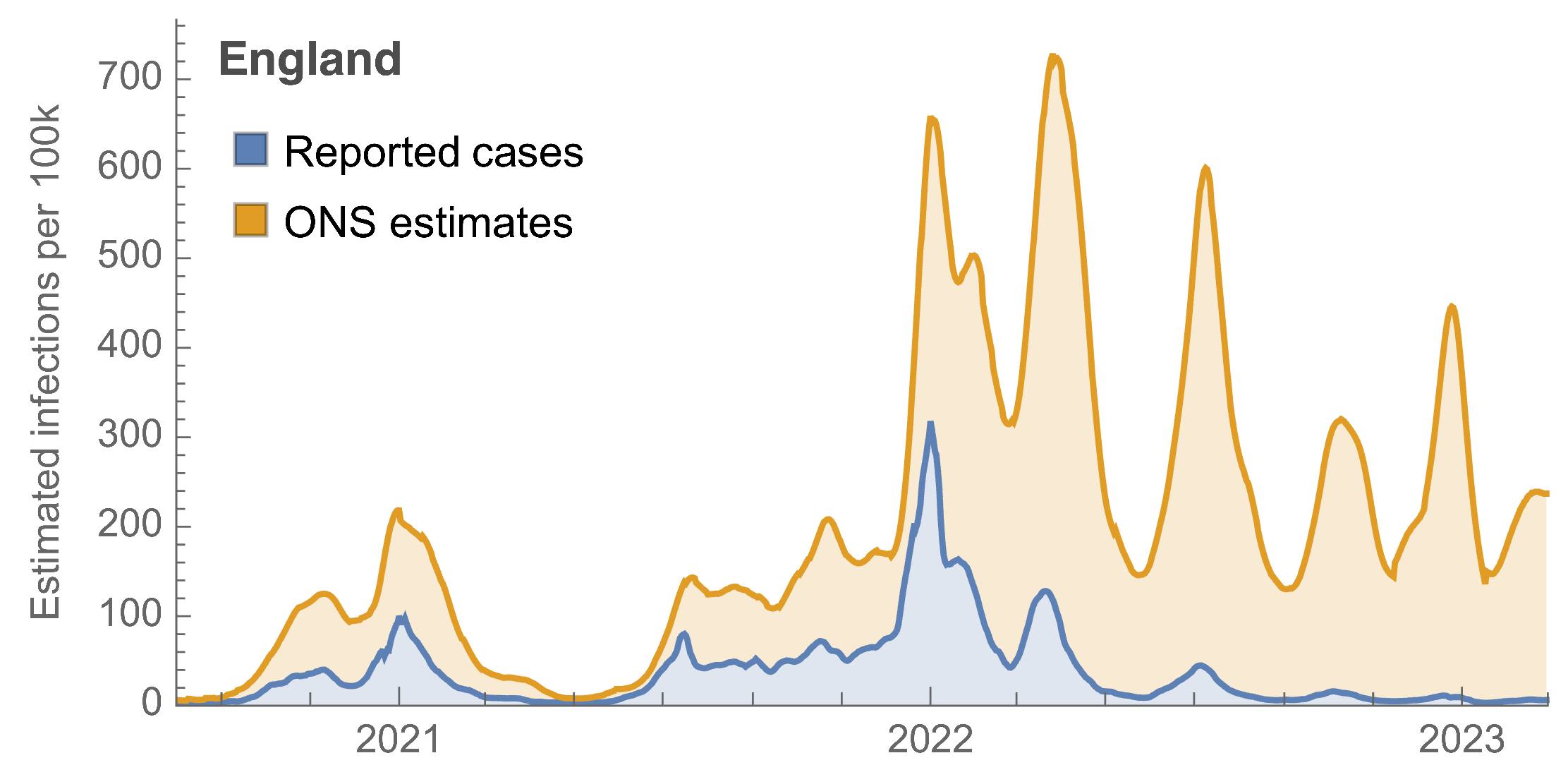
Partitioning ONS incidence based on sequencing data shows variant-driven epidemics
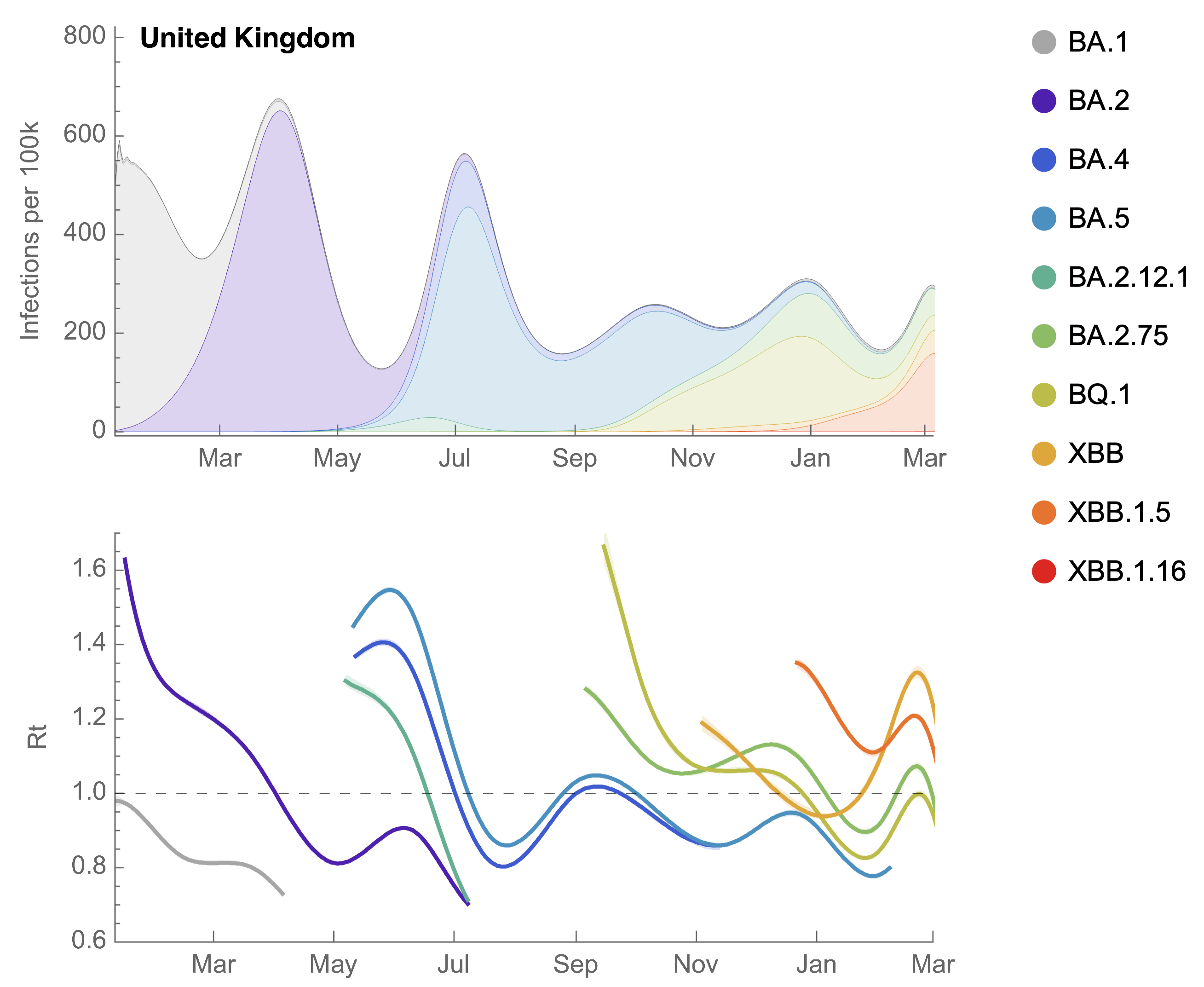
~110% population attack rate from March 2022 to March 2023
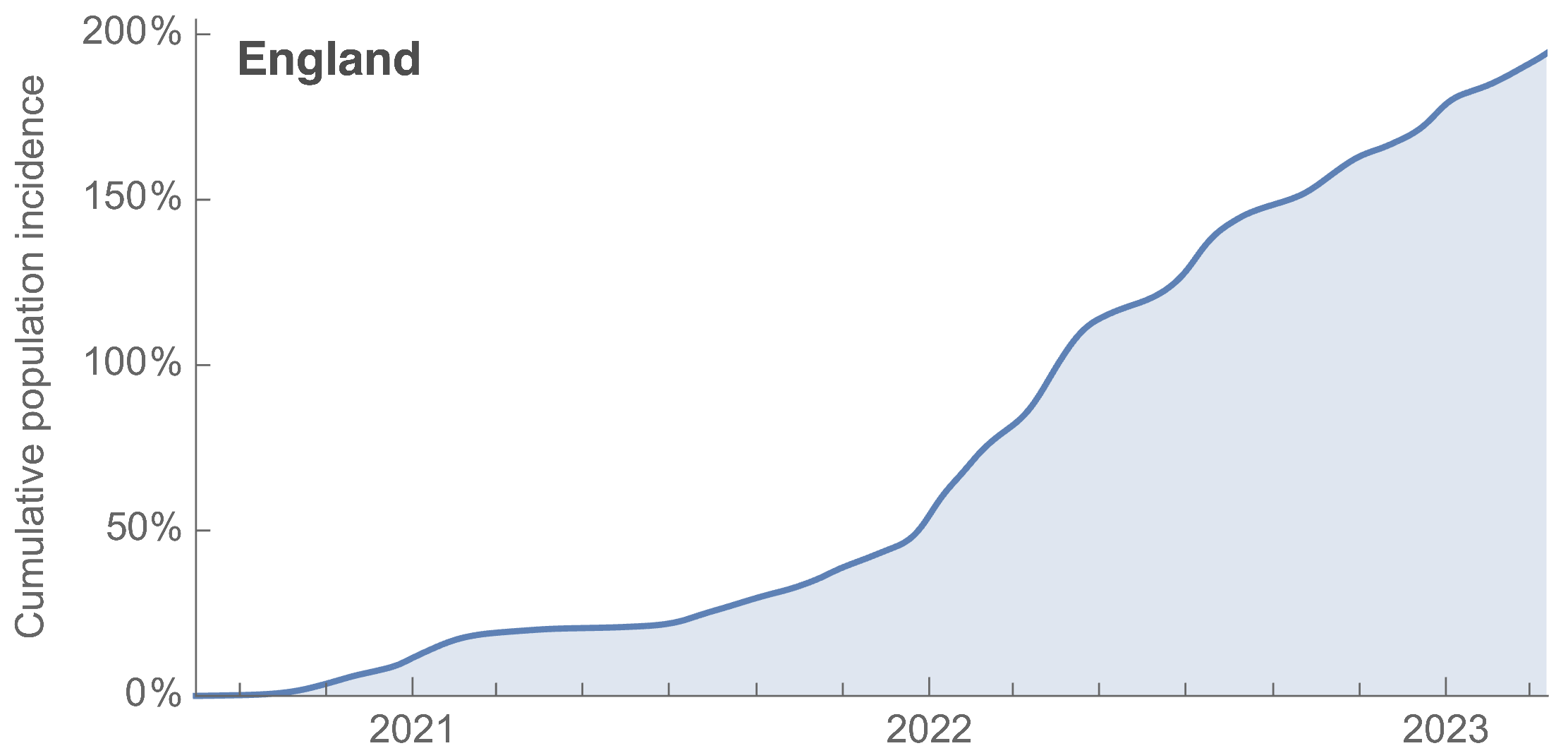
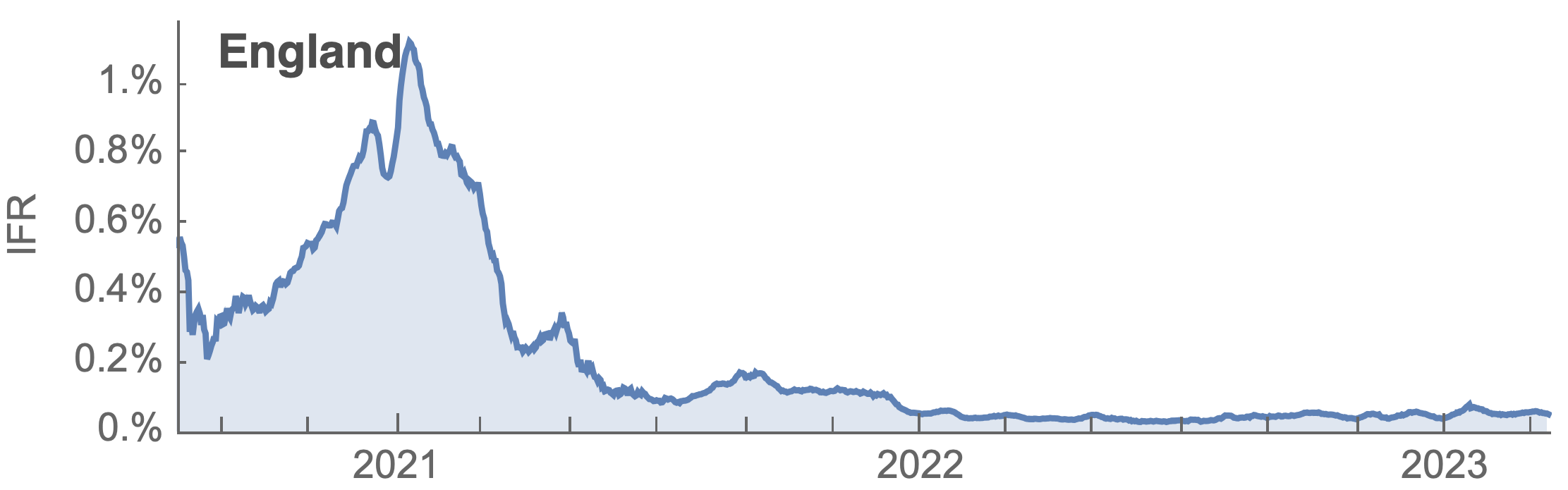
Post-Omicron period shows consistent IFR of 0.04%
Forecasting
Assessing MLR models for short-term frequency forecasting
MLR models generate accurate short-term forecasts
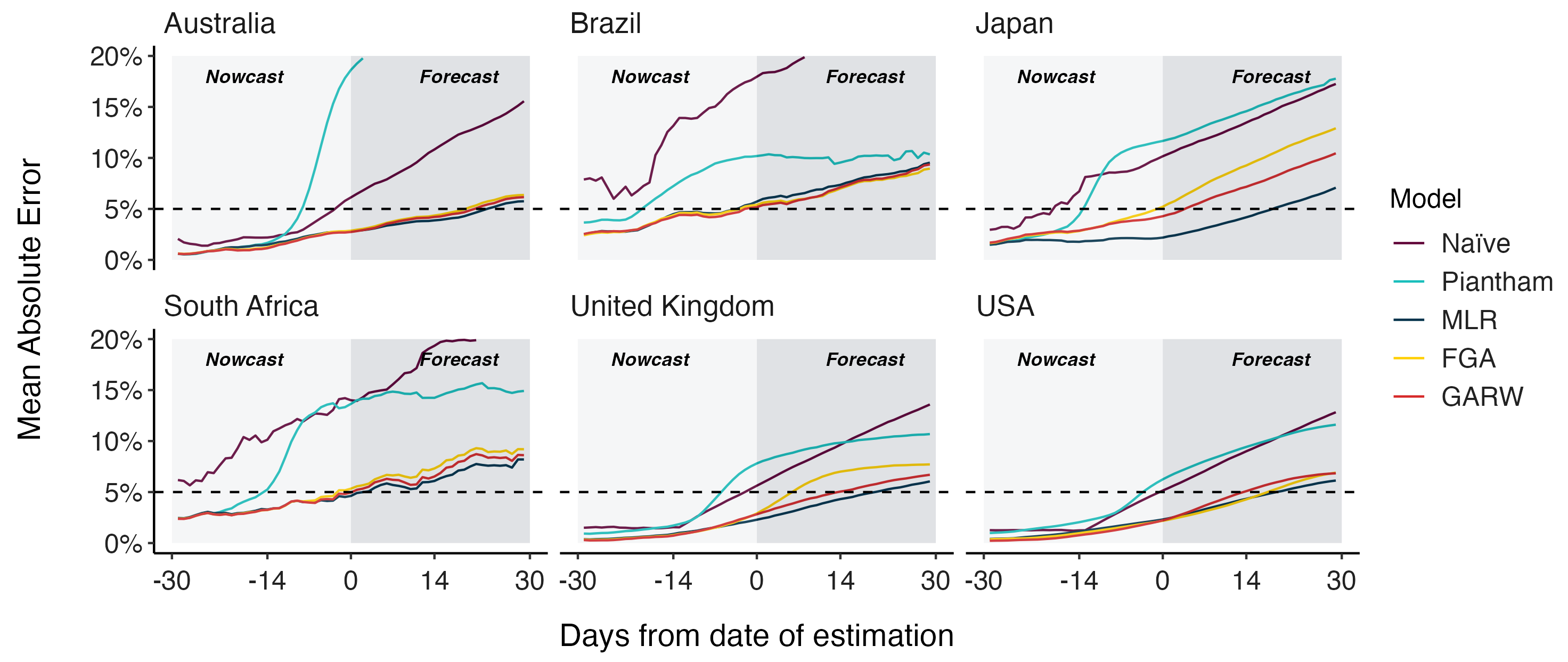
Now have clade and lineage forecasts continuously updated
Hierarchical MLR model
The hierarchical model allows pooling of growth advantages across locations. This allows us to include locations with fewer sequences and to better estimate growth advantage of rare lineages.
| Initial frequency | Growth advantage | |||
|---|---|---|---|---|
| Japan | $p_{23A}$ | $p_{23B}$ | $f_{23A}$ | $f_{23B}$ |
| USA | $p_{23A}$ | $p_{23B}$ | $f_{23A}$ | $f_{23B}$ |
| hierarchical | $f_{23A}$ | $f_{23B}$ | ||
Multinomial logistic regression should work well for SARS-CoV-2 prediction, except new variants have been emerging fast enough that the prediction horizon is really quite short
Could we predict the spread of new mutations using DMS data?
Escape from antibodies that potently neutralize BA.2
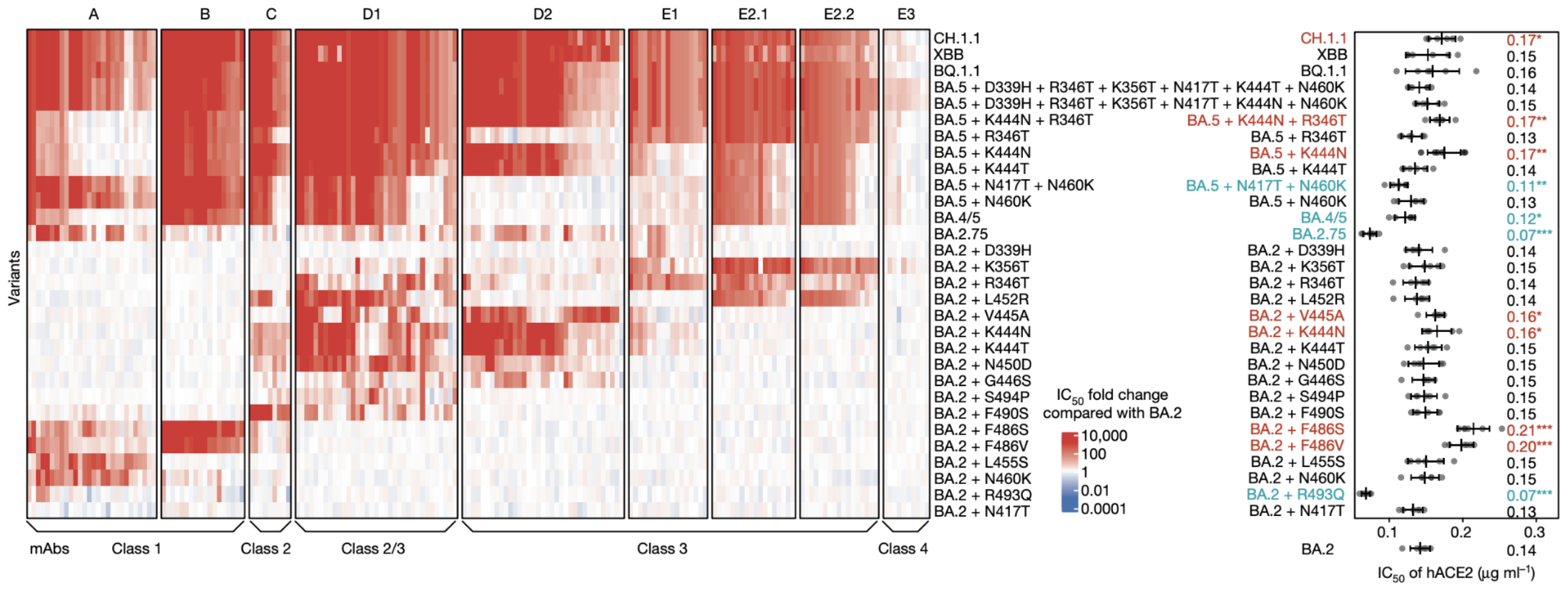
Can calculate escape of arbitrary RBD against antibodies known to neutralize BA.2
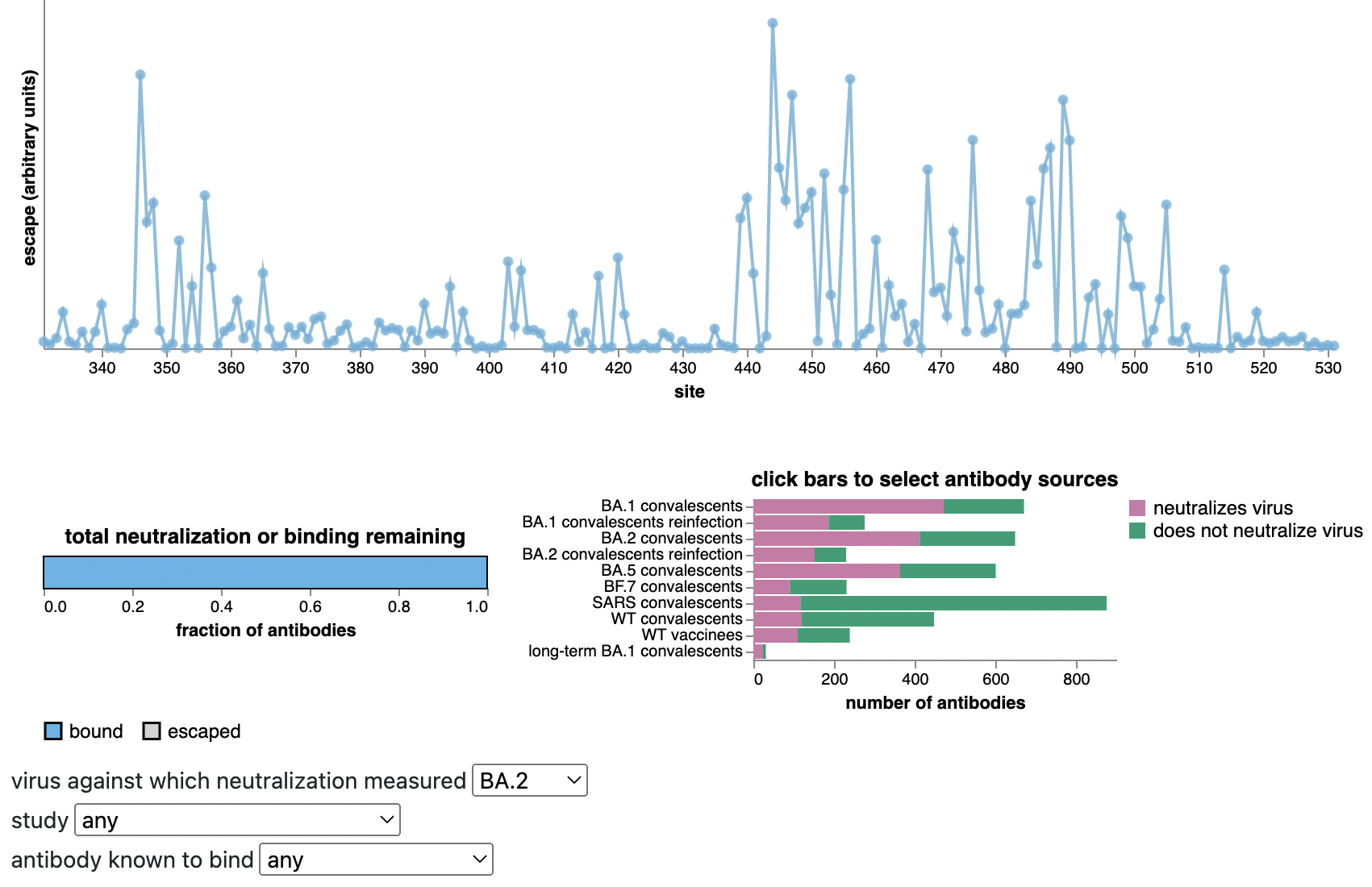
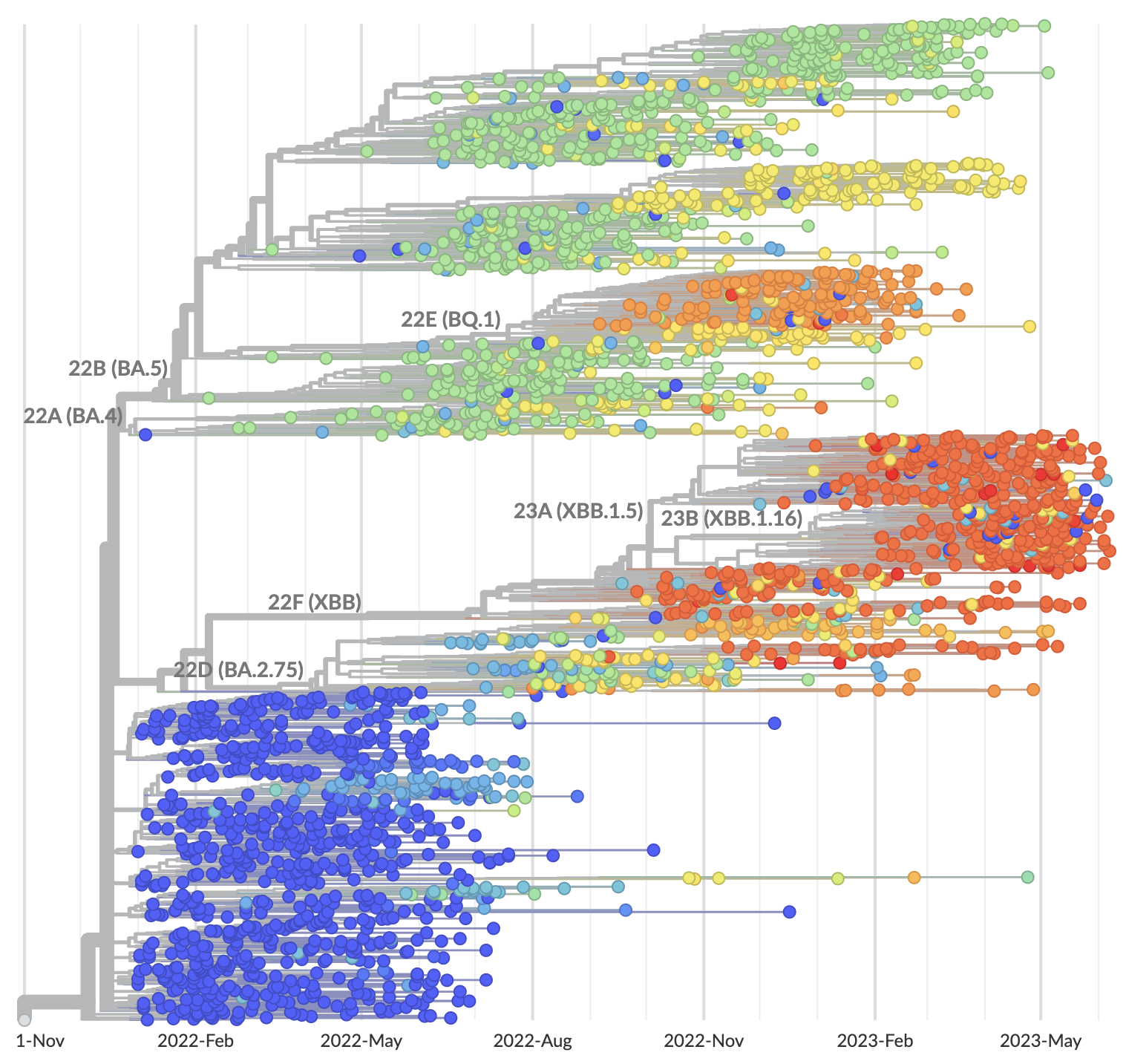
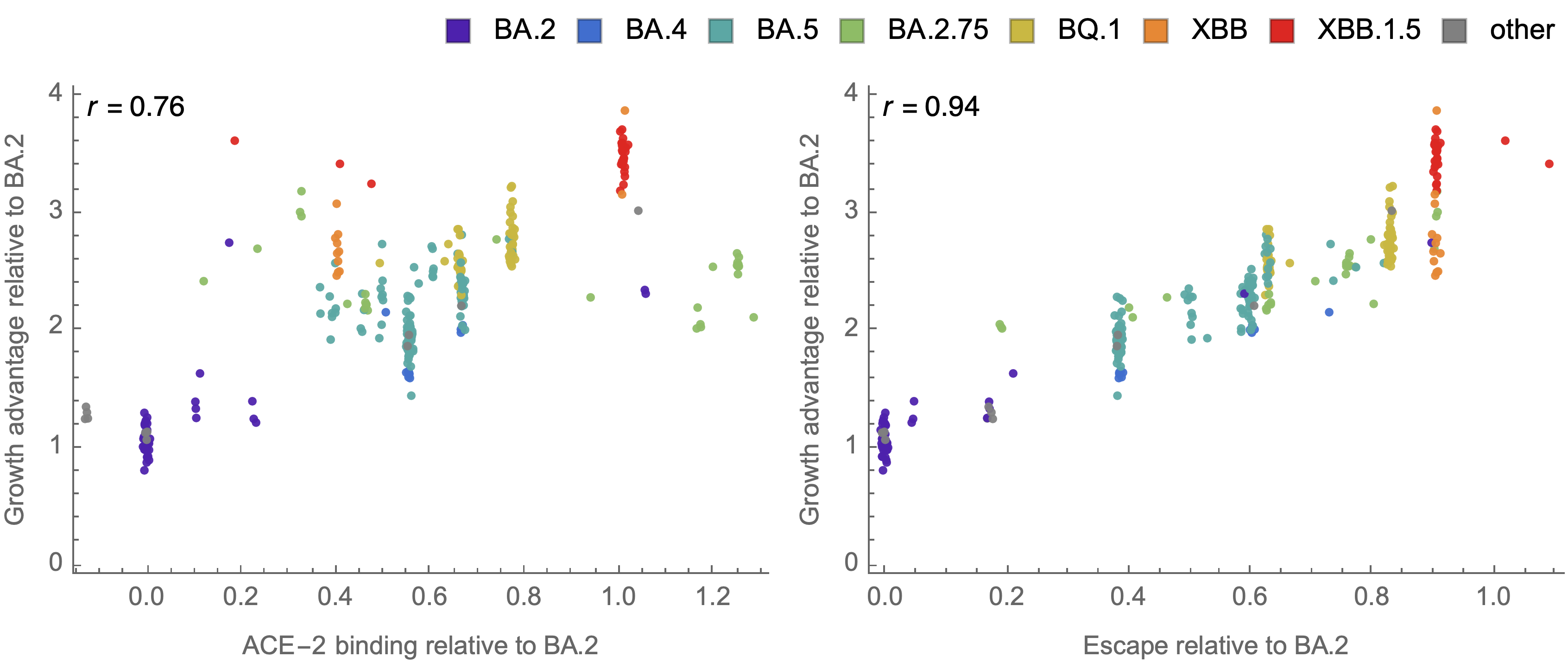
Strong correlation between DMS immune escape and lineage-level MLR growth advantage
Continued research
- Application of MLR models to other pathogens, such as seasonal influenza
- Assessing and improving accuracy of "live" models at nextstrain.org/sars-cov-2/forecasts/
- Implementing DMS priors to predict fitness of emerging and yet-to-emerge lineages
Acknowledgements
SARS-CoV-2 genomic epi: Data producers from all over the world, GISAID and the Nextstrain team
Bedford Lab:
![]() John Huddleston,
John Huddleston,
![]() James Hadfield,
James Hadfield,
![]() Katie Kistler,
Katie Kistler,
![]() Thomas Sibley,
Thomas Sibley,
![]() Jover Lee,
Jover Lee,
![]() Cassia Wagner,
Cassia Wagner,
![]() Miguel Paredes,
Miguel Paredes,
![]() Nicola Müller,
Nicola Müller,
![]() Marlin Figgins,
Marlin Figgins,
![]() Victor Lin,
Victor Lin,
![]() Jennifer Chang,
Jennifer Chang,
![]() Allison Li,
Allison Li,
![]() Eslam Abousamra,
Eslam Abousamra,
![]() Donna Modrell,
Donna Modrell,
![]() Nashwa Ahmed,
Nashwa Ahmed,
![]() Cécile Tran Kiem
Cécile Tran Kiem






