Phylodynamics and molecular evolution of SARS-CoV-2
Trevor Bedford (@trvrb)
Associate Professor, Fred Hutchinson Cancer Research Center
2 Dec 2021
Epidemics8
Slides at: bedford.io/talks
1. Real-time tracking of SARS-CoV-2 evolution
2. Emergence of variants of concern
3. Current circulation patterns
4. Assessing adaptive evolution
5. Variant transmission dynamics
6. Omicron variant
Real-time tracking of SARS-CoV-2 evolution
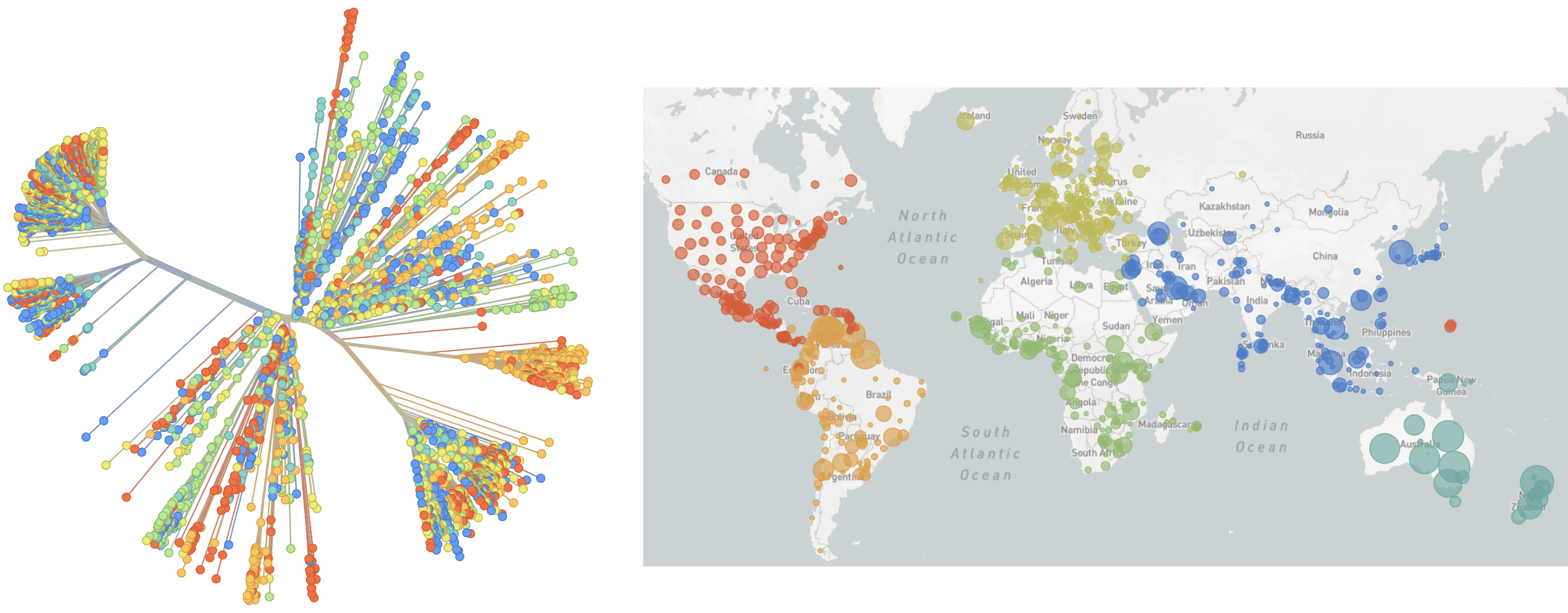
Over 5.5M SARS-CoV-2 genomes shared to GISAID and evolution tracked in real-time at nextstrain.org
![]() Richard Neher,
Richard Neher,
![]() Emma Hodcroft,
Emma Hodcroft,
![]() James Hadfield,
James Hadfield,
![]() Thomas Sibley,
Thomas Sibley,
![]() John Huddleston,
John Huddleston,
![]() Ivan Aksamentov,
Ivan Aksamentov,
![]() Moira Zuber,
Moira Zuber,
![]() Jover Lee,
Jover Lee,
![]() Cassia Wagner,
Cassia Wagner,
![]() Denisse Sequeira,
Denisse Sequeira,
![]() Cornelius Roemer,
Cornelius Roemer,
![]() Victor Lin,
Victor Lin,
![]() Jennifer Chang
Jennifer Chang
SARS-CoV-2 lineages establish globally in February and March
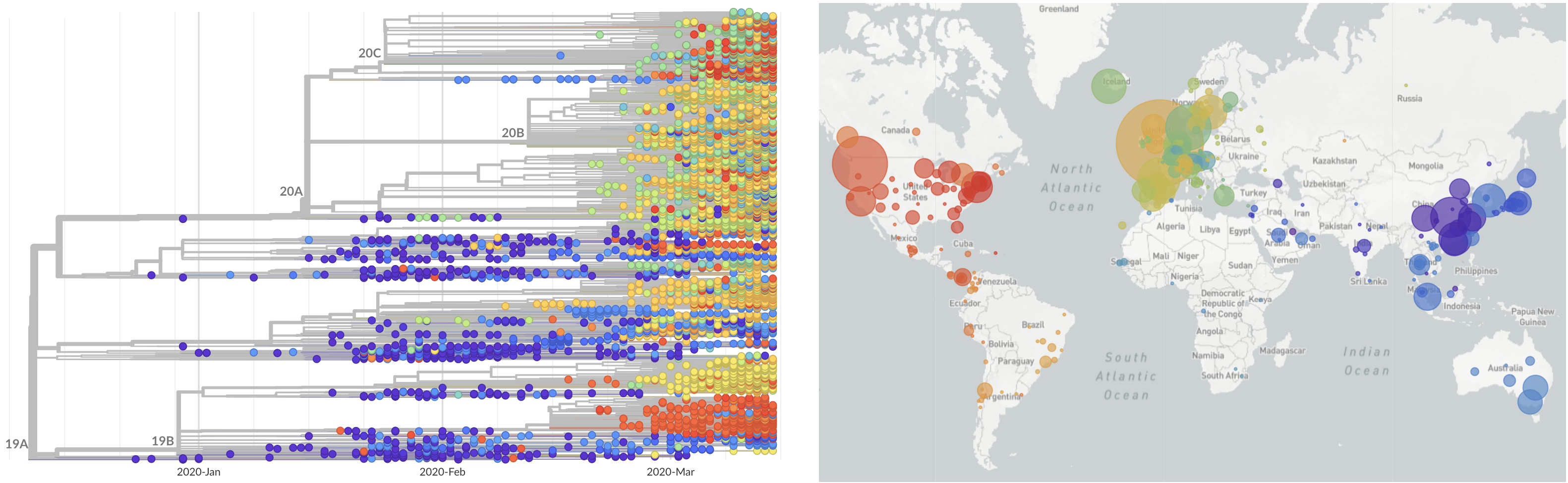
Limited early mutations like spike D614G spread globally during initial wave
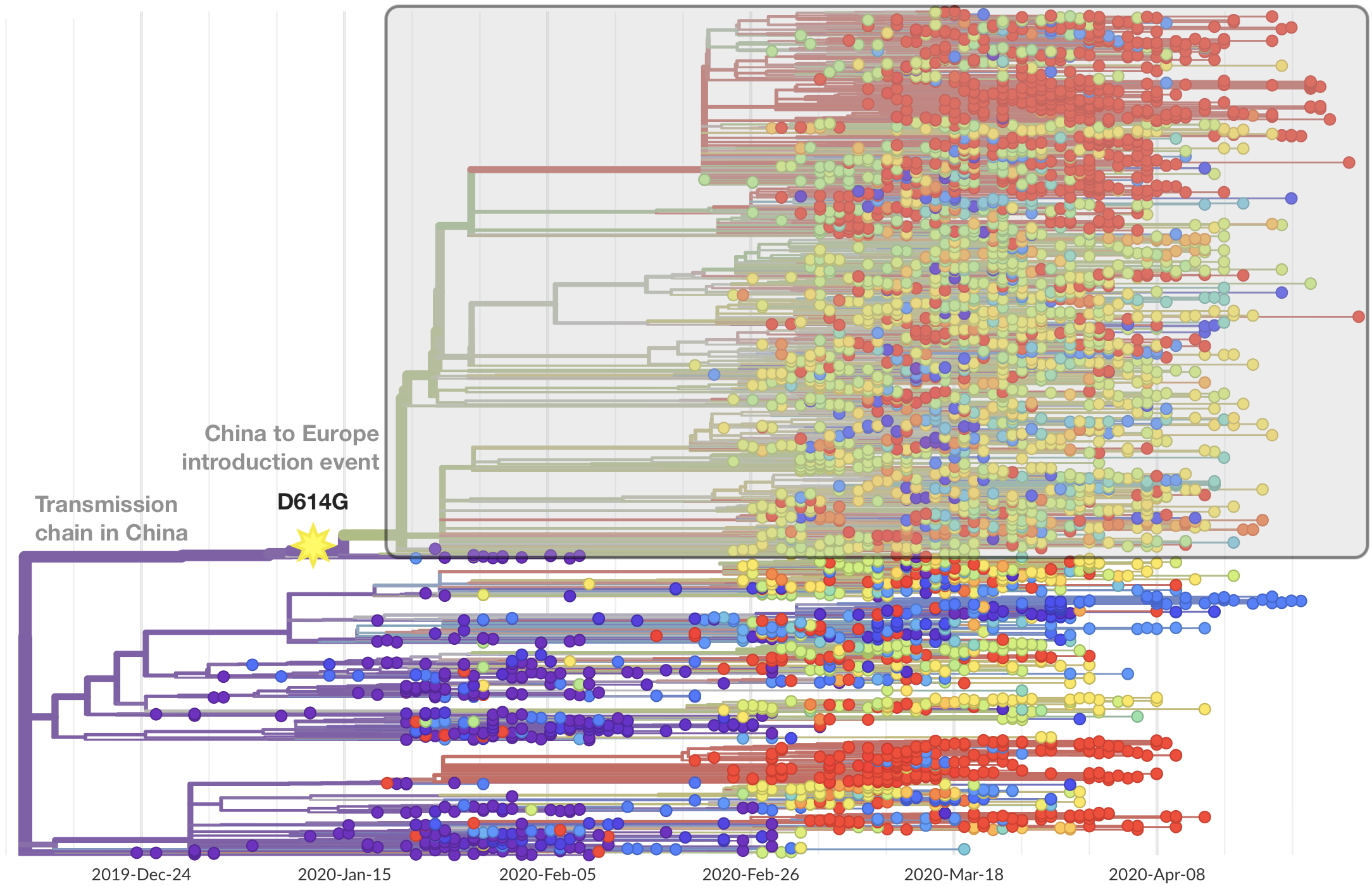
Mutations in summer and fall 2020 were confined to regional dominance
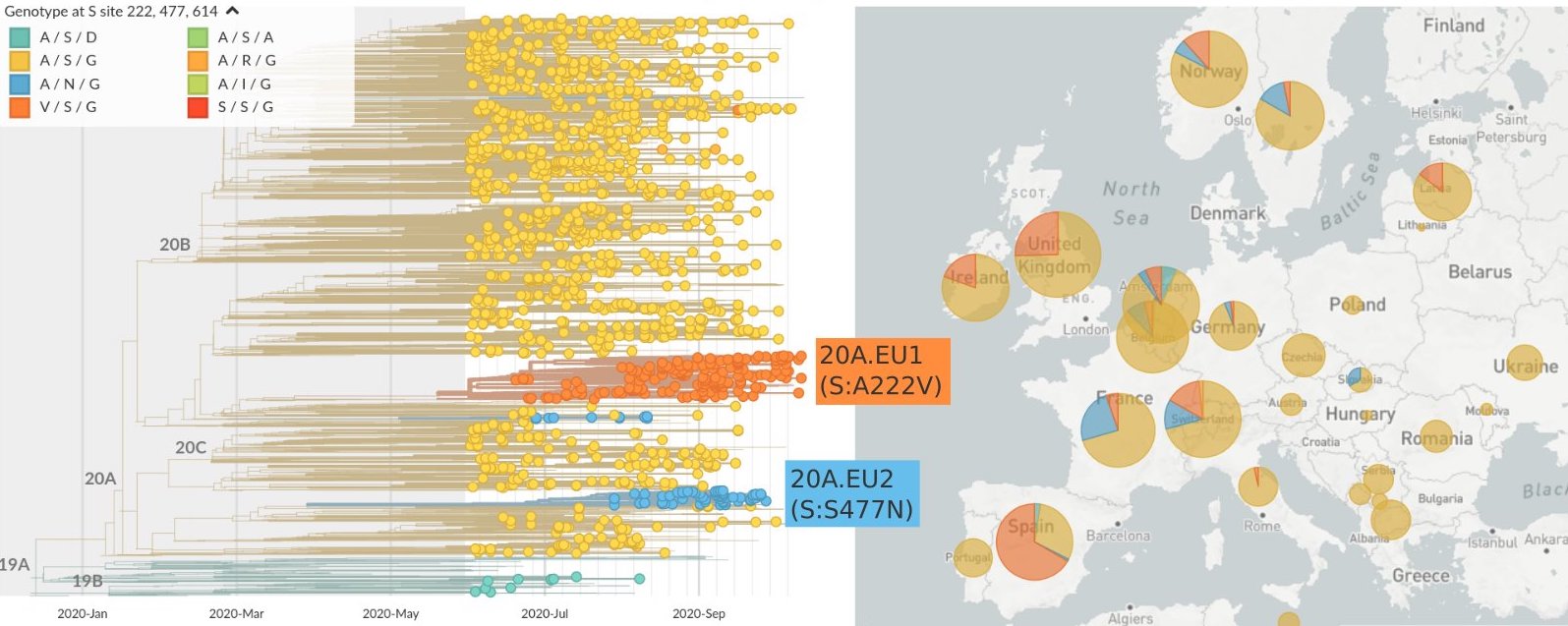
Emergence of variants of concern
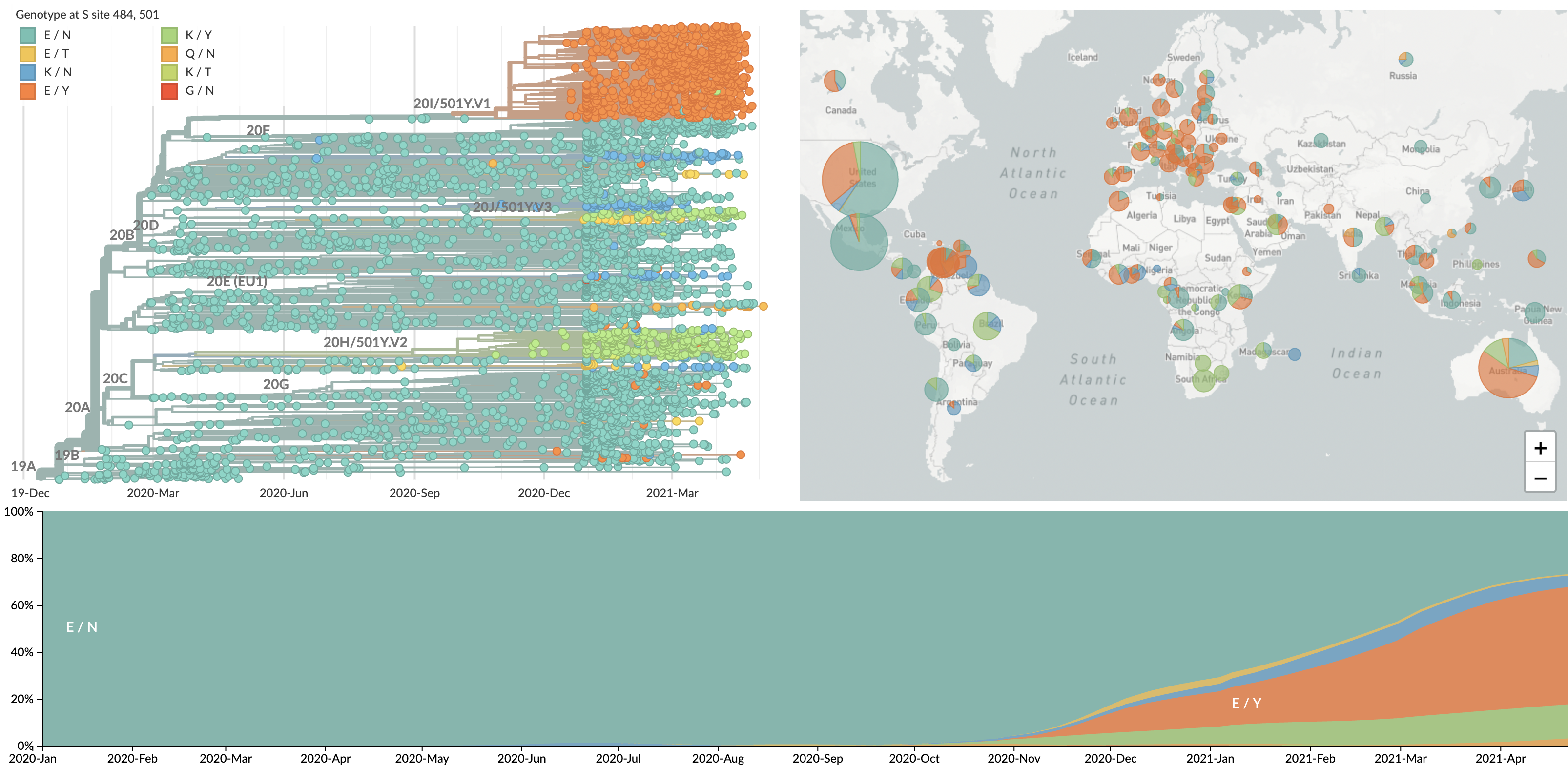
Repeated emergence of 484K and 501Y across the world
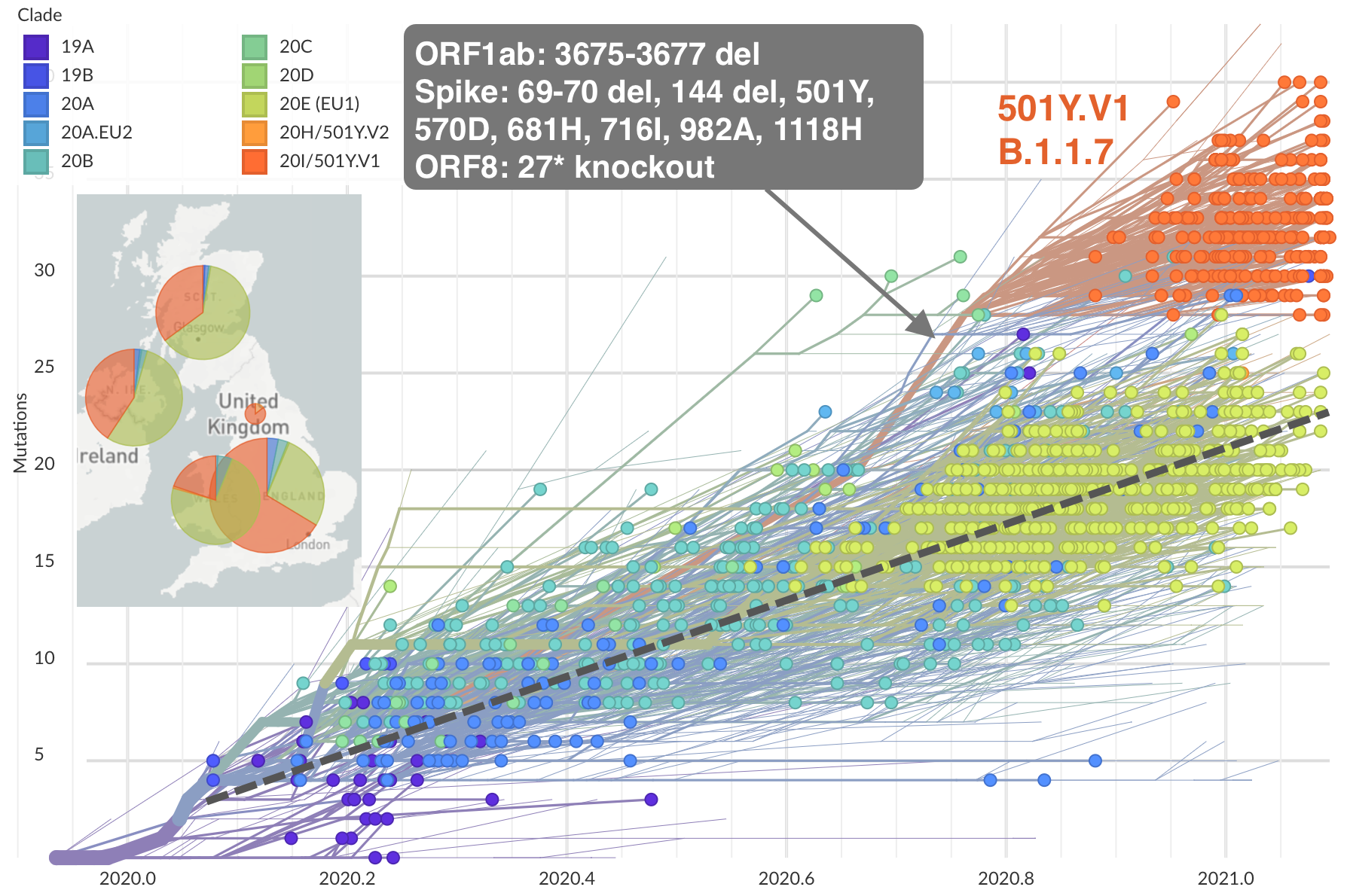
Emergence of Alpha (B.1.1.7) in the UK
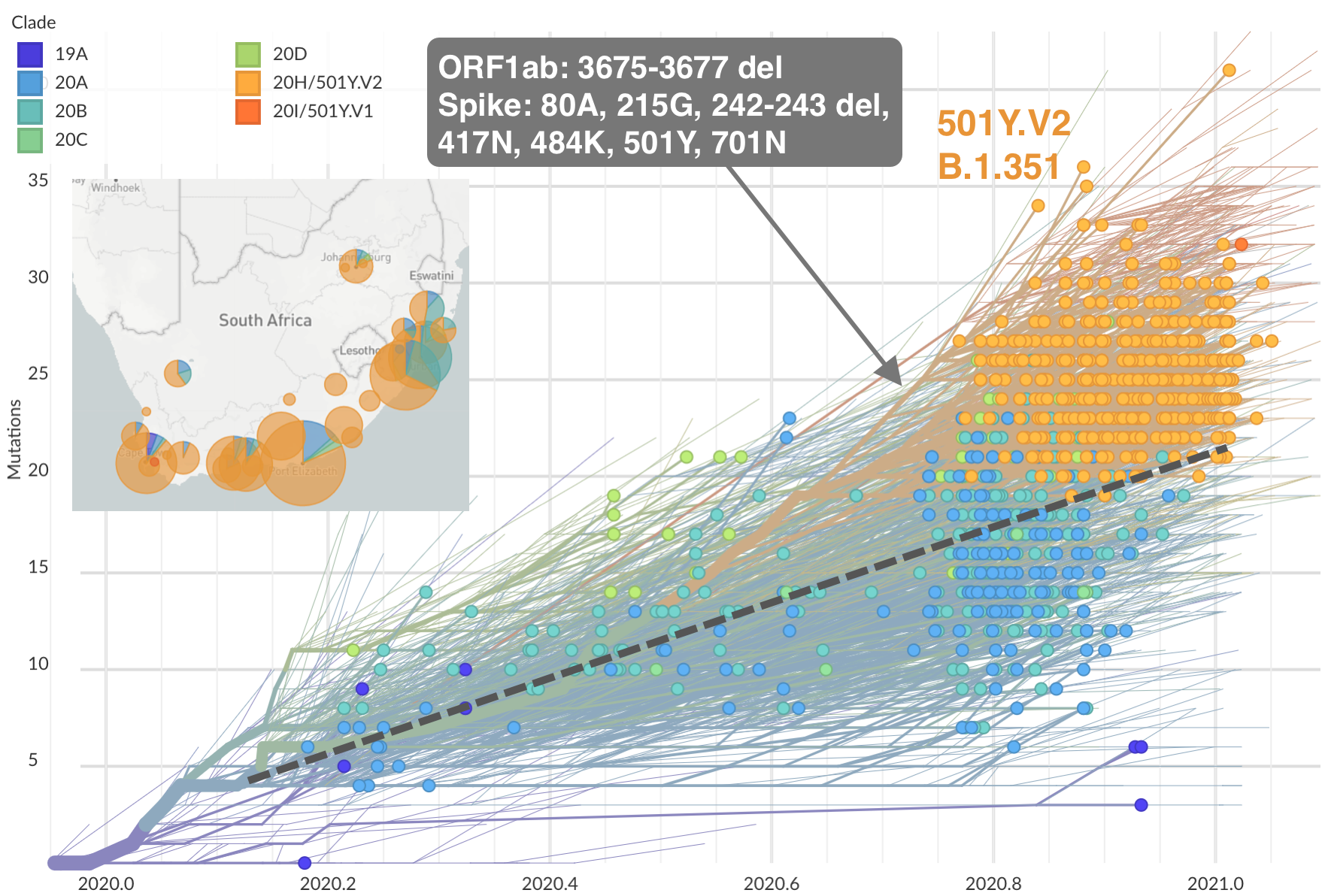
Emergence of Beta (B.1.351) in the South Africa
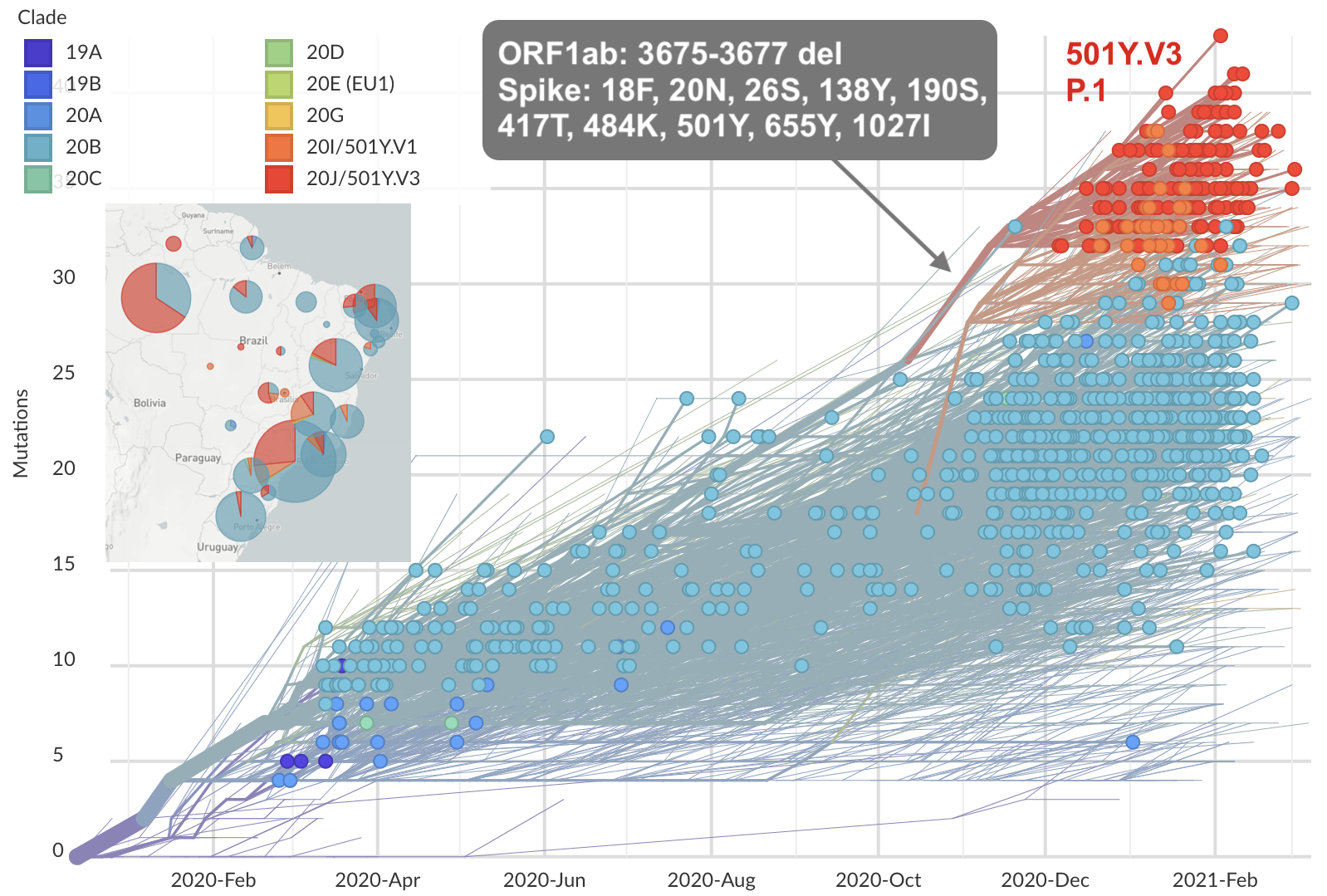
Emergence of Gamma (P.1) in the Brazil
Working hypothesis of within-host evolution occurring during infection, driven by natural selection against host immunity
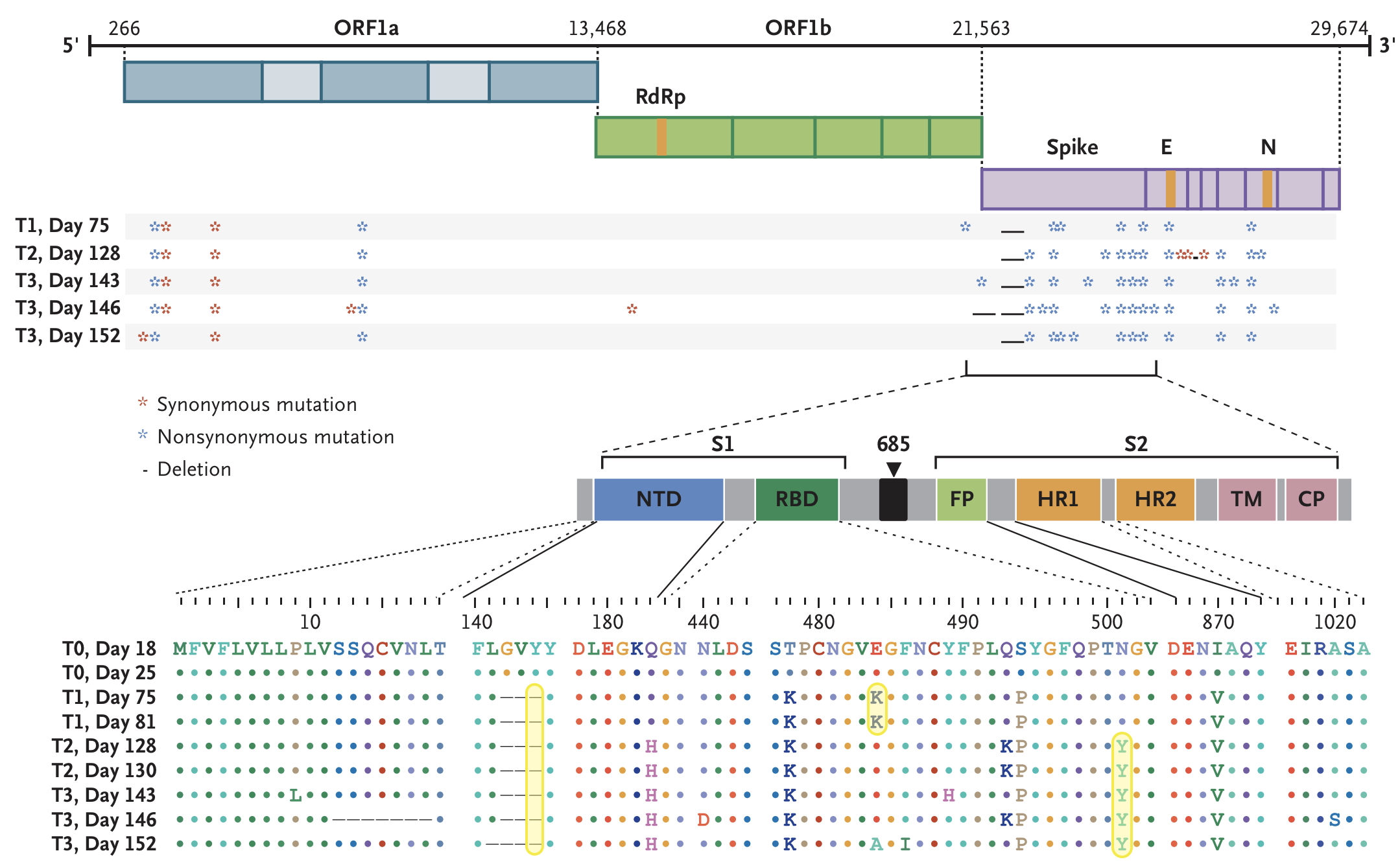
Rapid within-host evolution during persistent infection
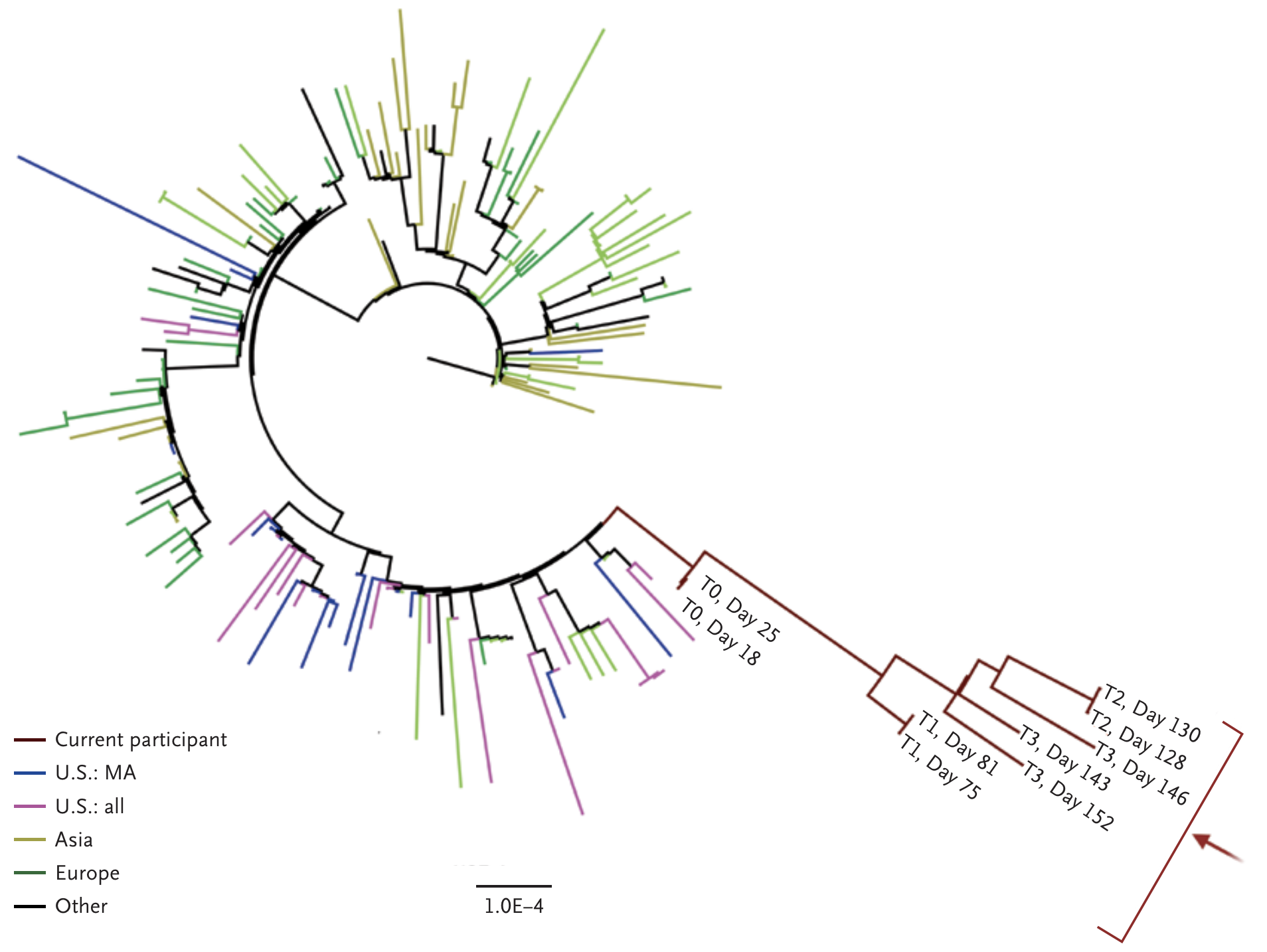
484K and 501Y observed during this evolution
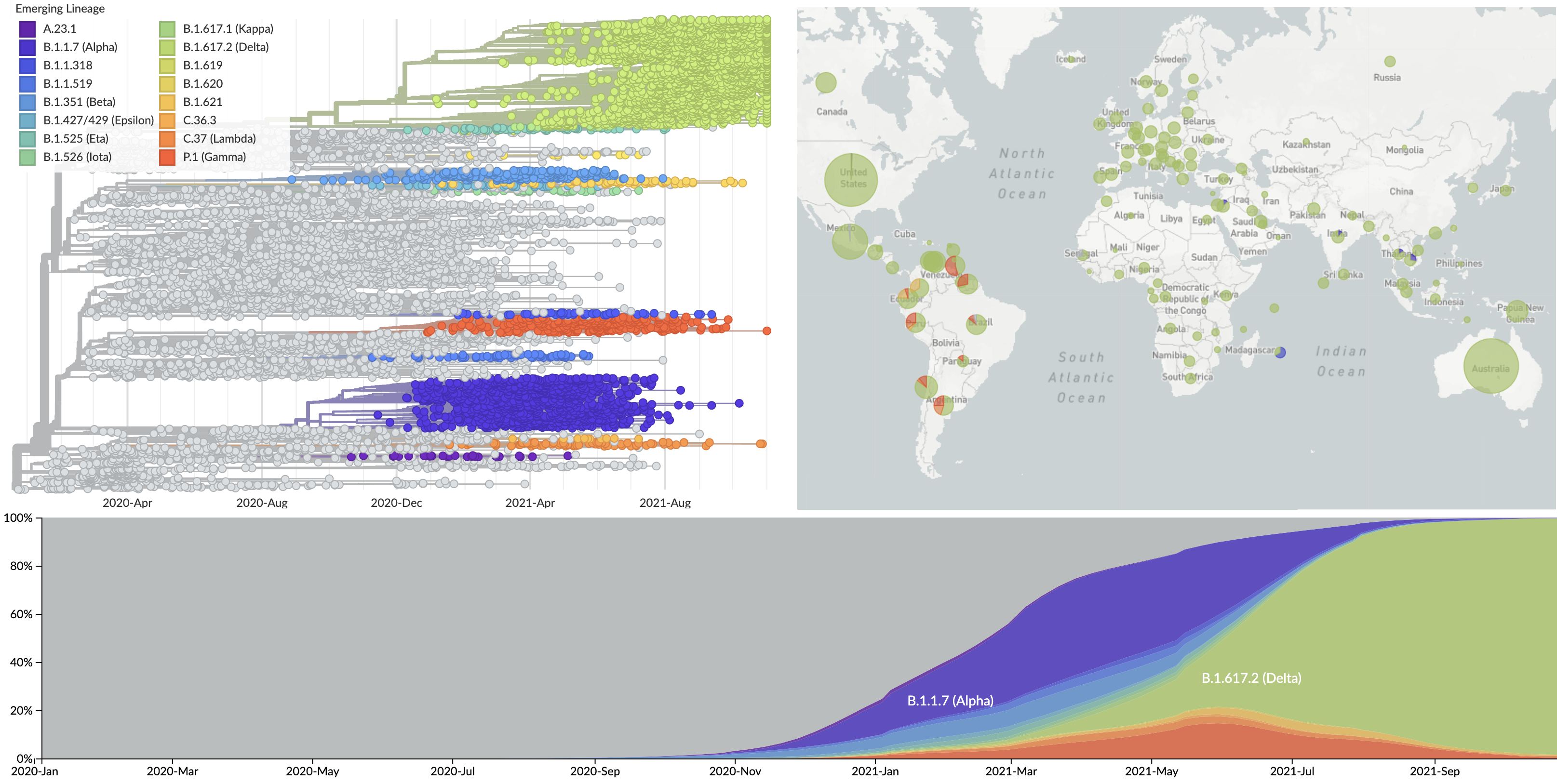
Current circulation patterns
Spread of VOC / VOI lineages across the world
Delta outcompeting other variants and seemed poised to sweep
Variants show excess mutations across the genome
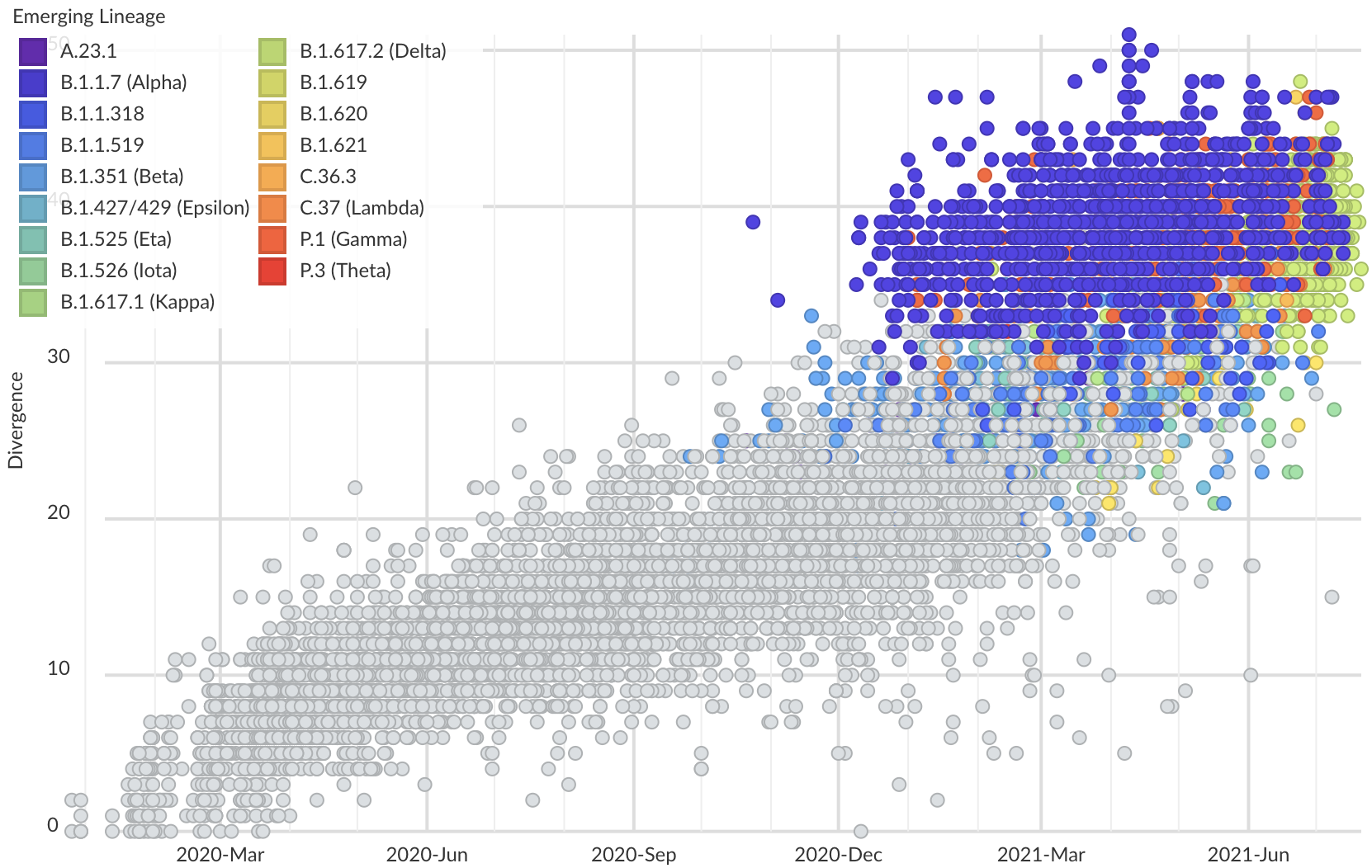
But show most substantial excess in the S1 domain of spike
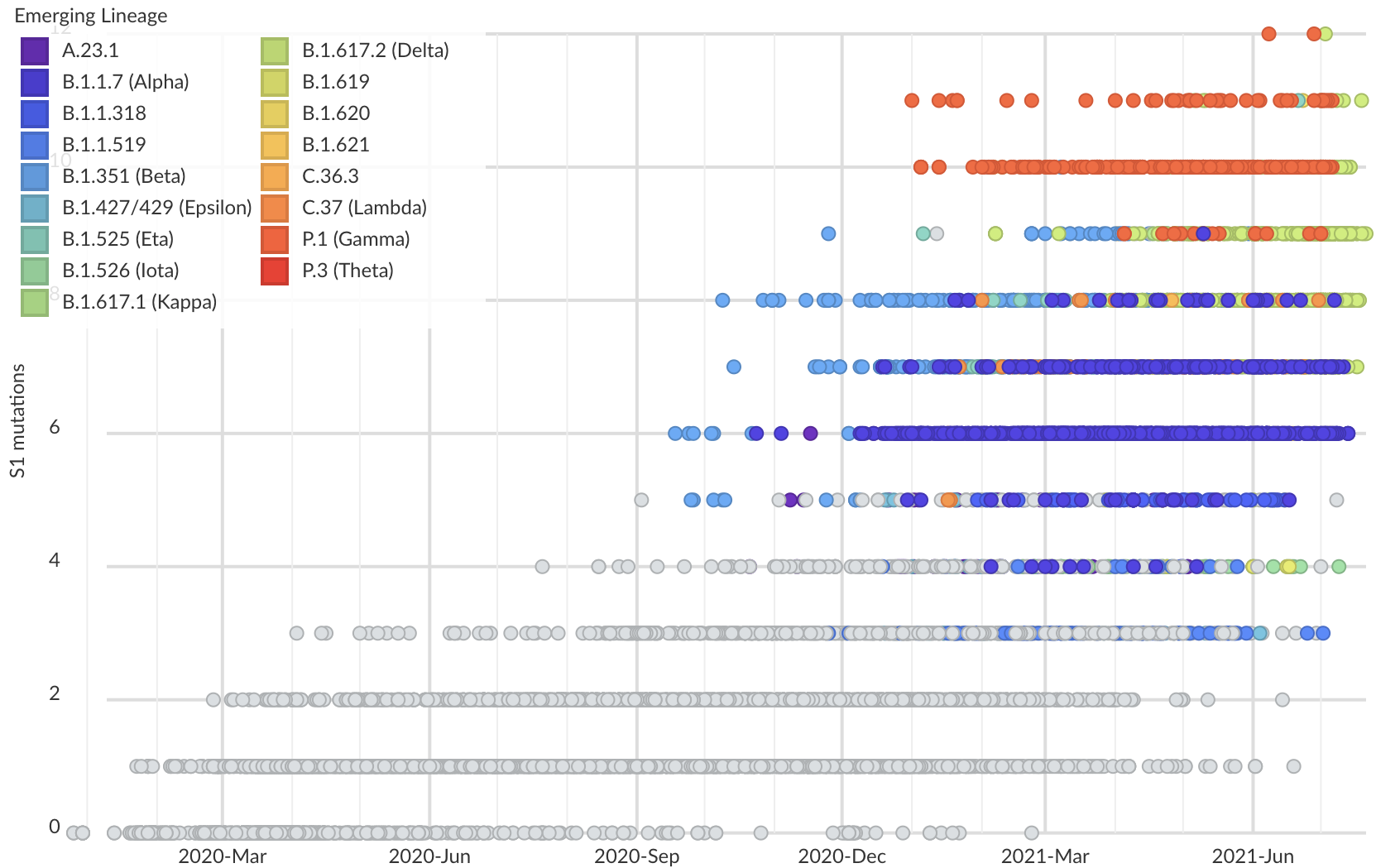
Assessing adaptive evolution
Rapid and parallel adaptive mutations in spike S1 drive clade success in SARS-CoV-2
 Katie Kistler,
Katie Kistler,
 John Huddleston
John Huddleston
Phylogeny of 10k genomes equitably sampled in space and time
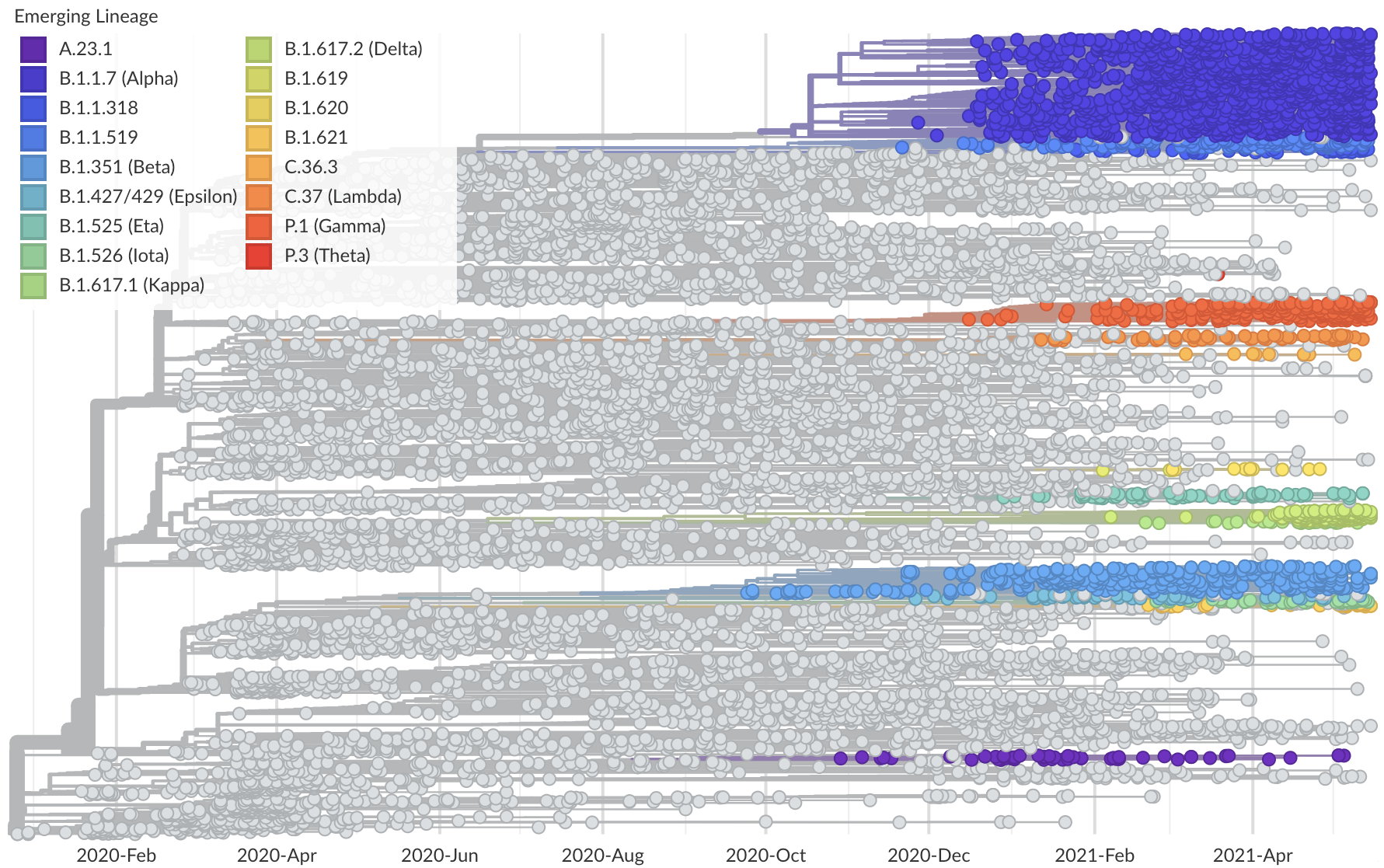
Measure clade growth as a proxy for viral fitness
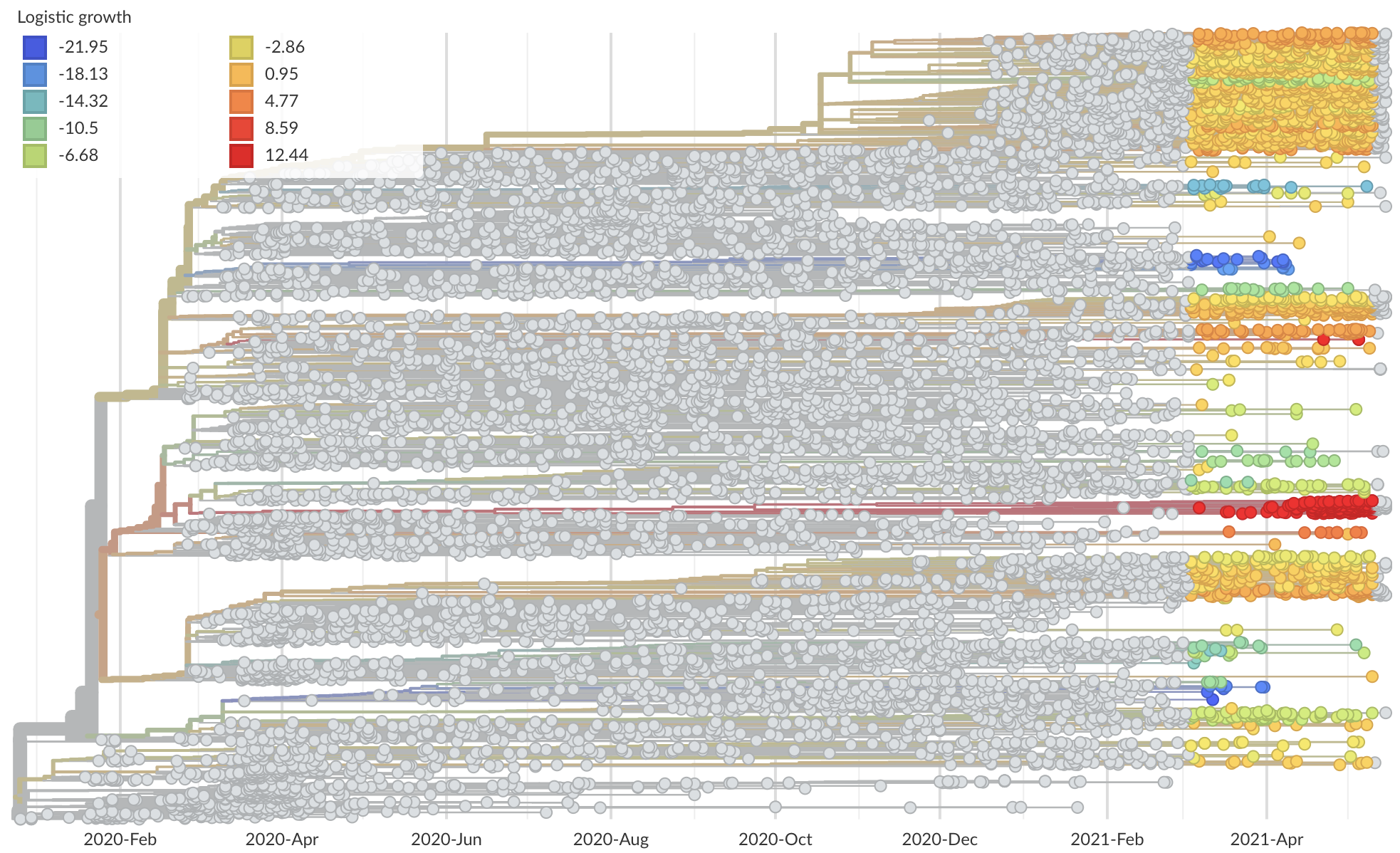
Clades with more S1 nonsynonymous mutations grow faster
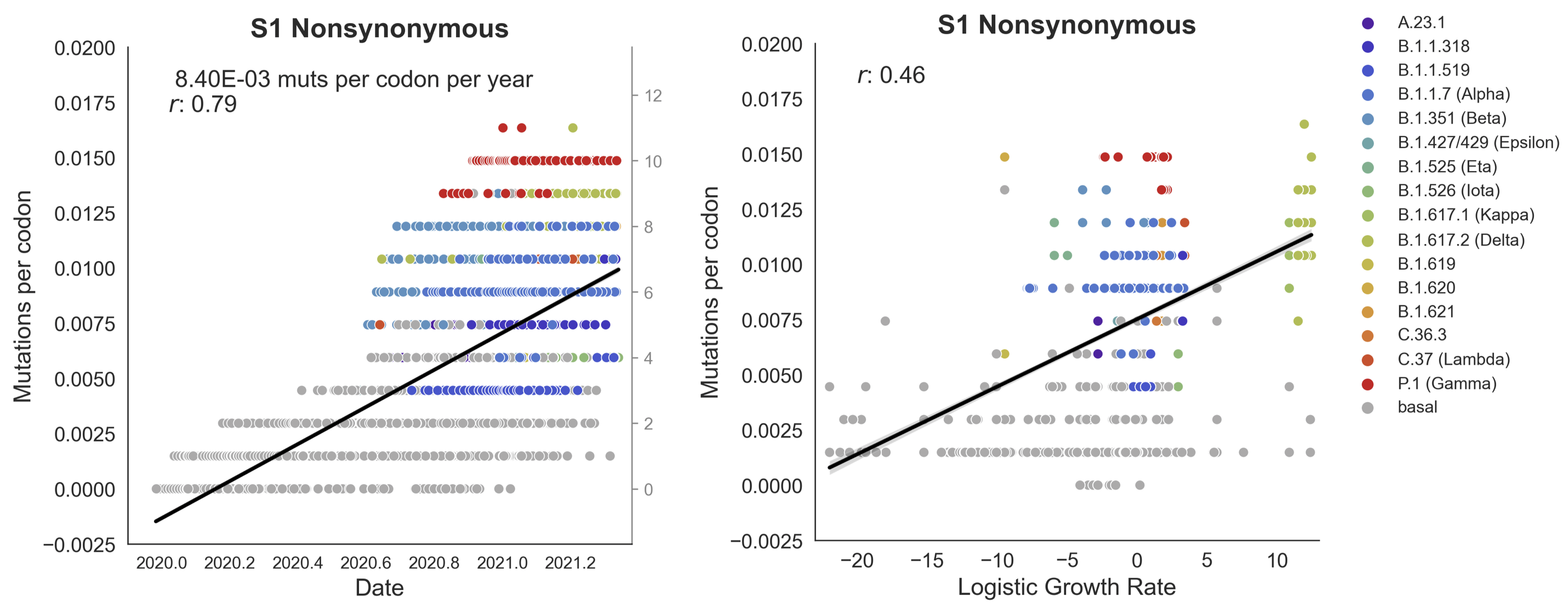
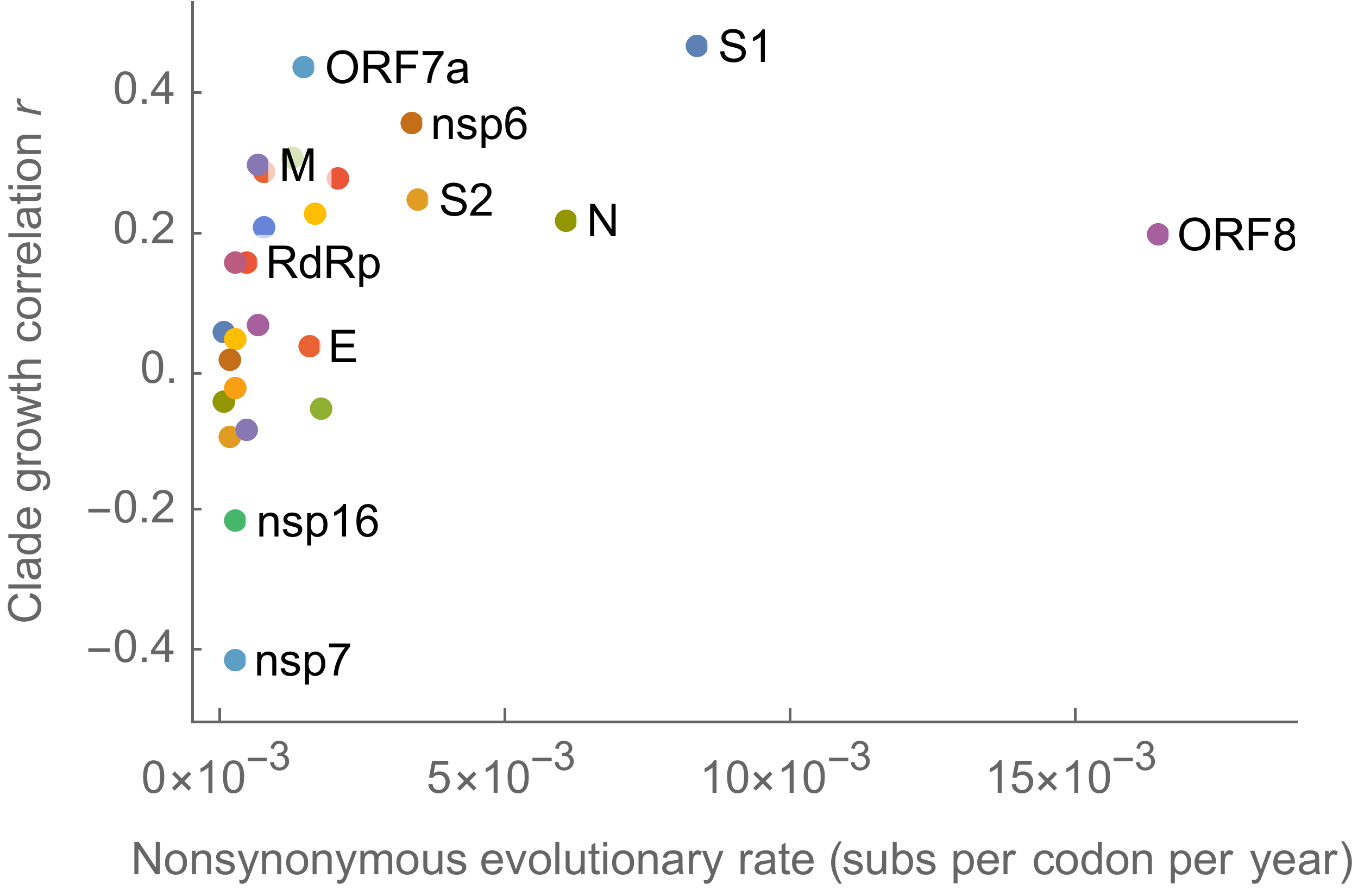
S1 is quickly evolving and highly correlated with clade growth
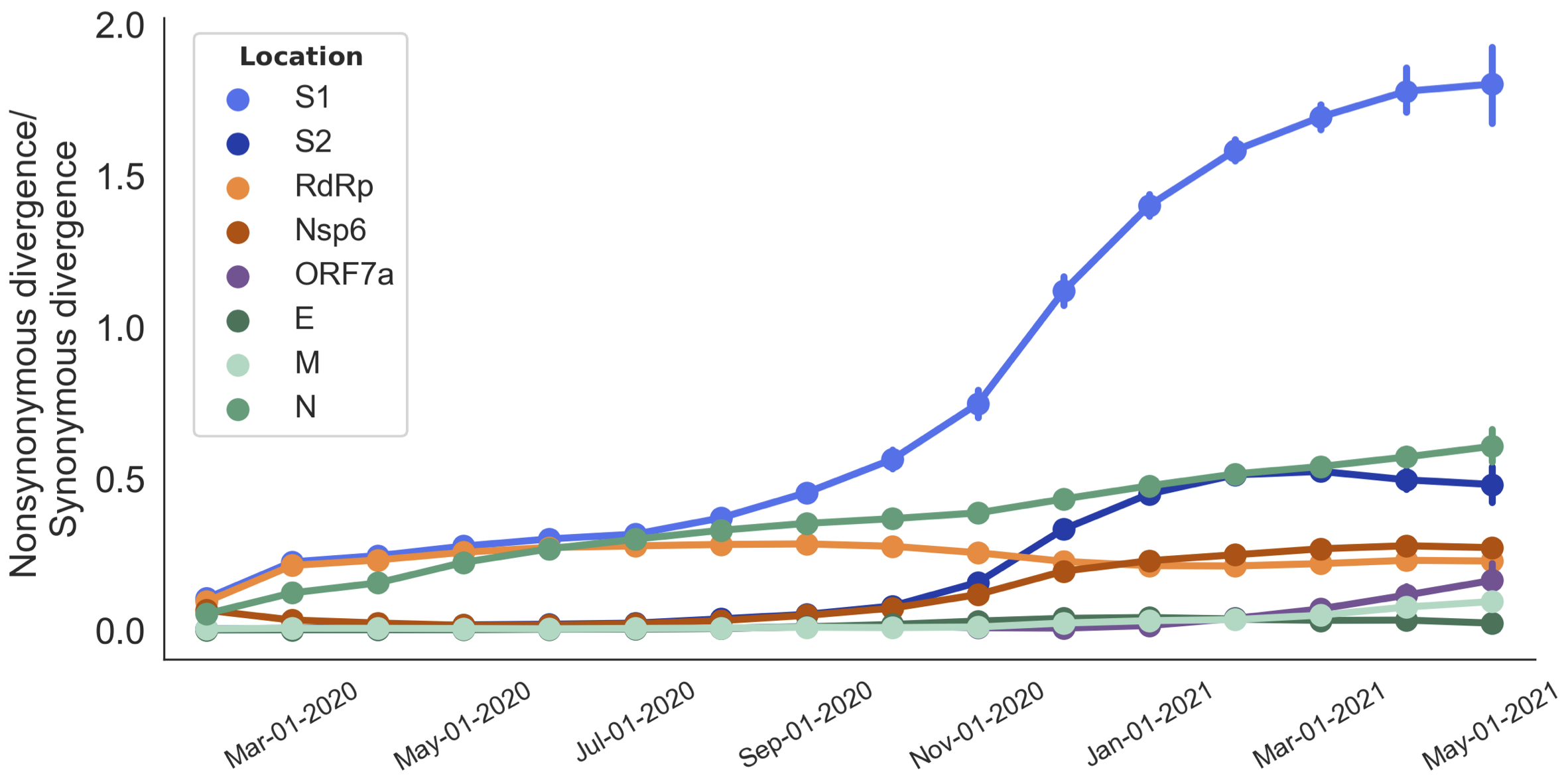
dN/dS to root further highlights adaptive evolution
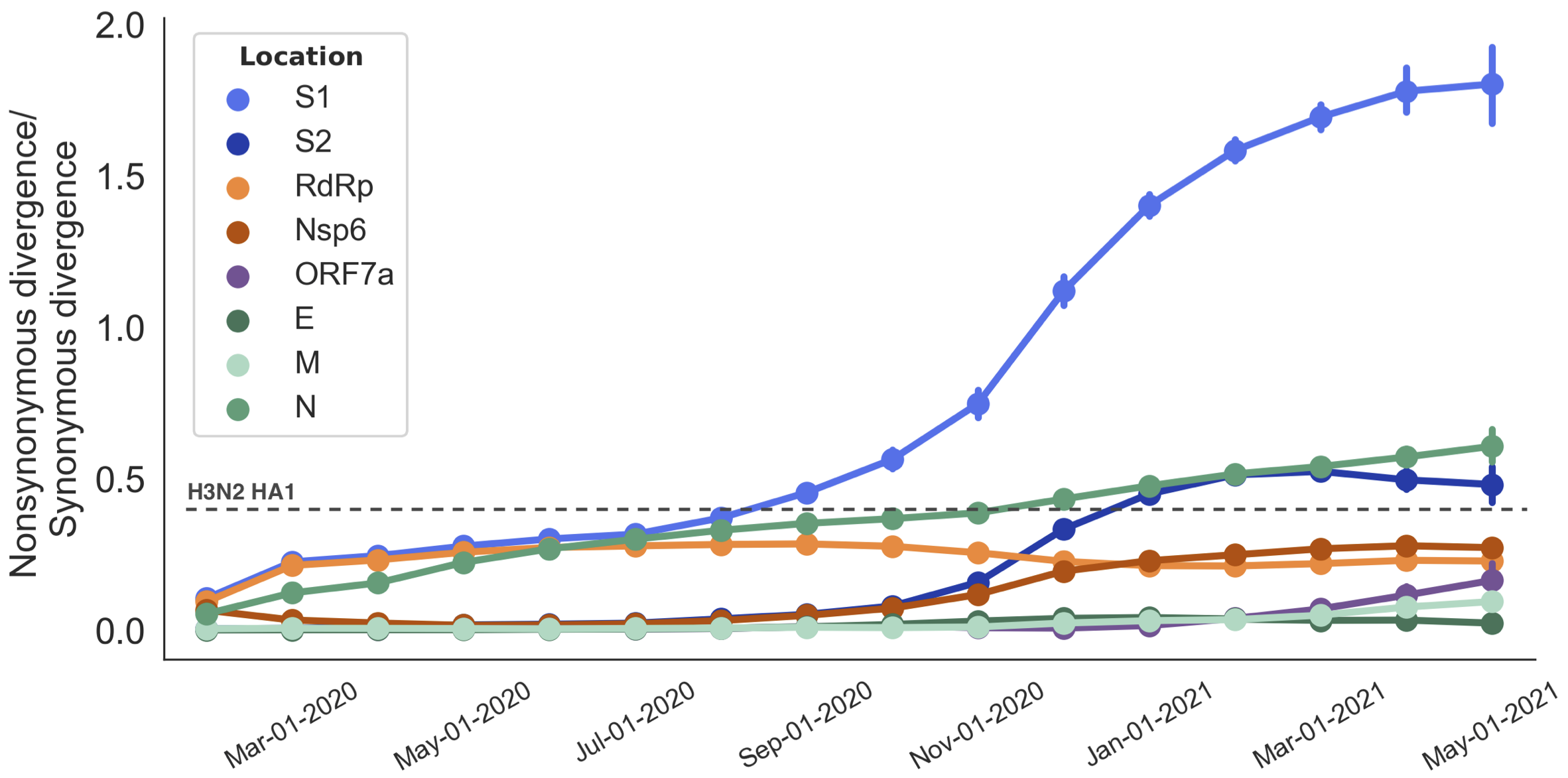
Rapid pace of adaptive evolution relative to H3N2 influenza
Mutations in S1 arising via within-host pressures result increase viral fitness and are enriched in the viral population by natural selection
Although selection has not been primarily for antigenic drift, observed level of adaptability suggests its potential
Variant transmission dynamics
Multiple approaches to modeling fitness differences between circulating variants
Multinomial logistic regression models work well on frequencies
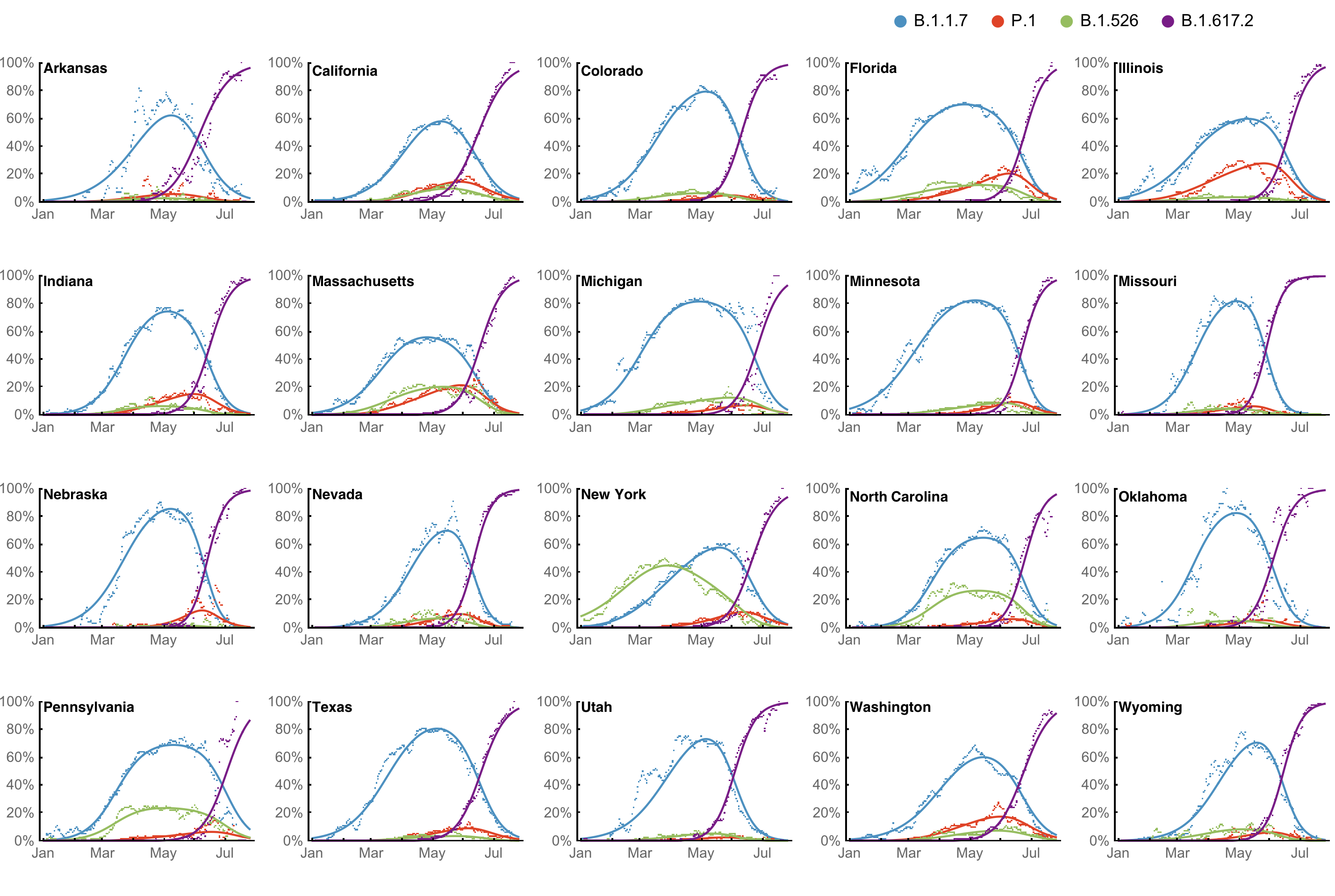
However, frequency of a variant may rise while cases fall
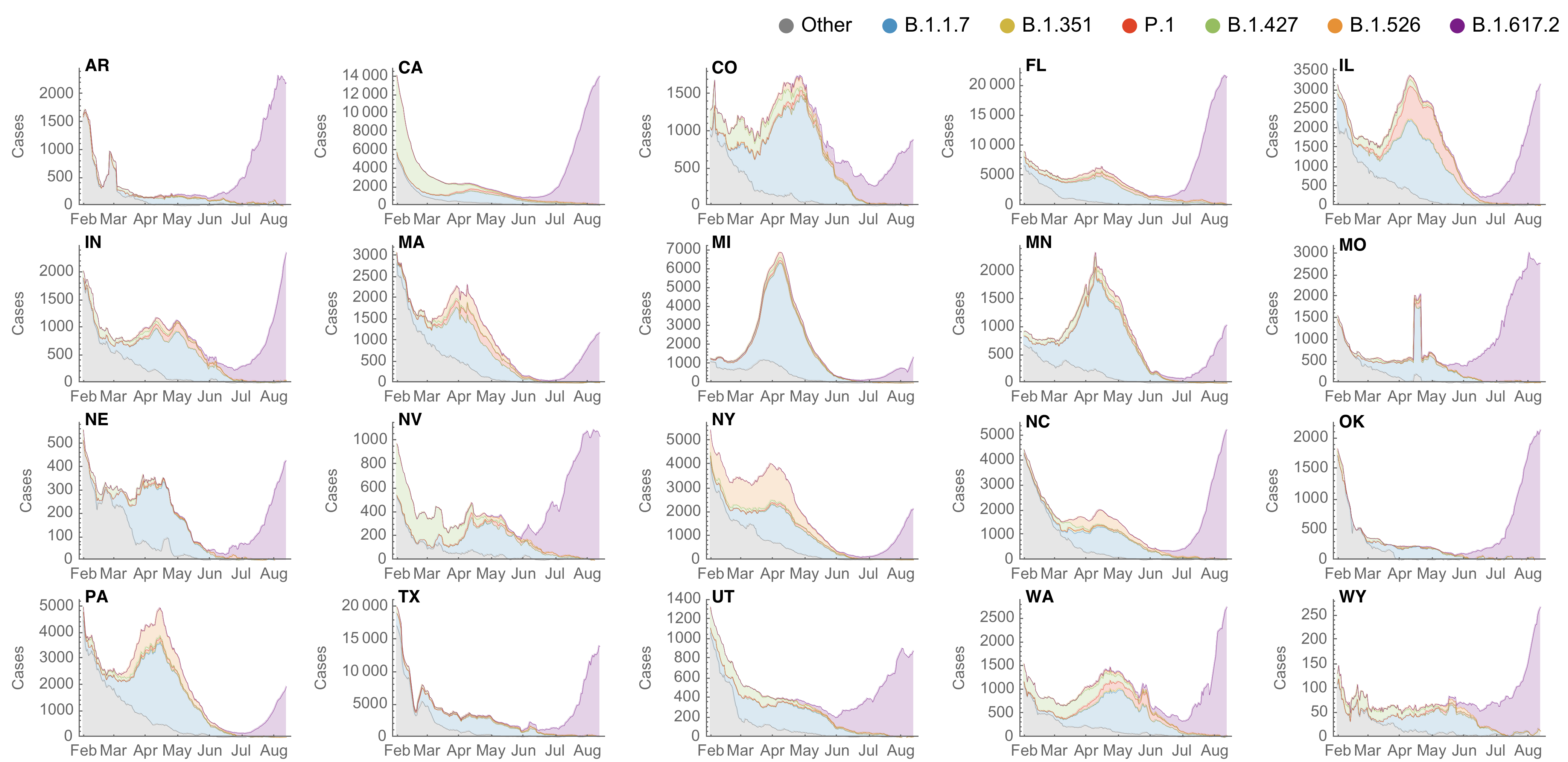
SARS-CoV-2 variant dynamics across US states show consistent differences in transmission rates
 Marlin Figgins
Marlin Figgins
Estimation of variant-specific Rt through time using state-level data
State-level case counts are partitioned based on frequencies of sequenced cases
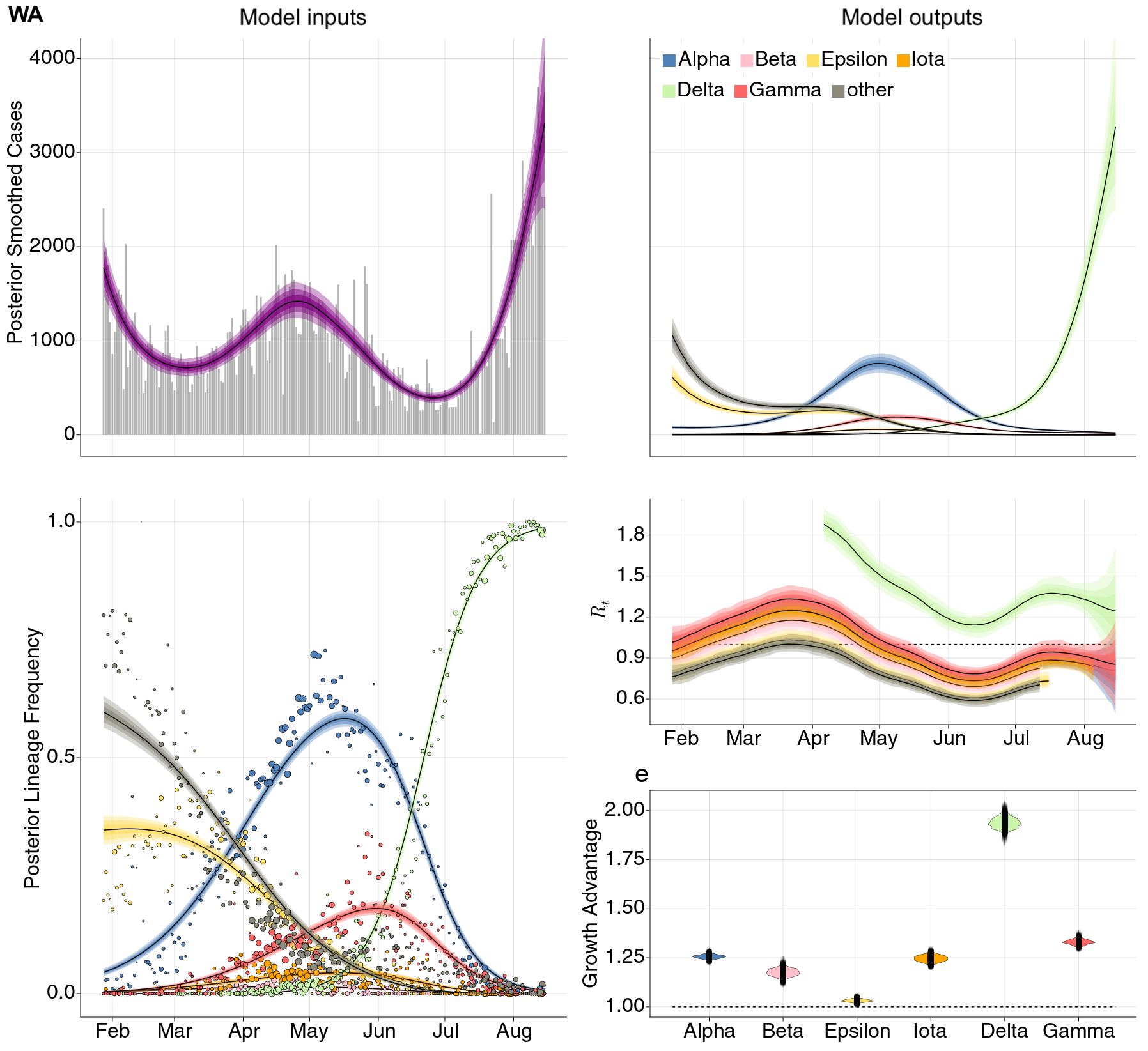
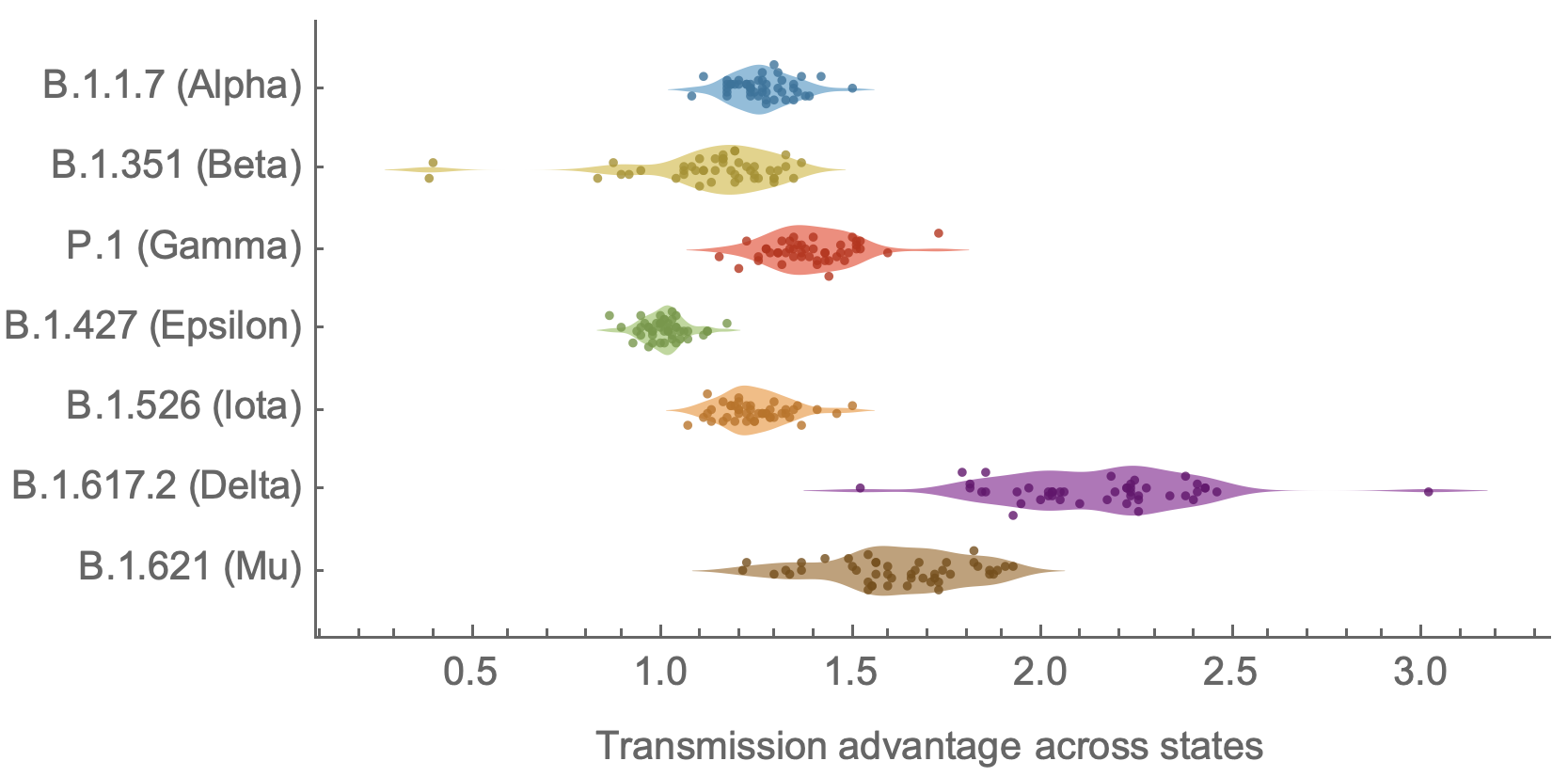
Differences in intrinsic Rt across variants, but all trending downwards
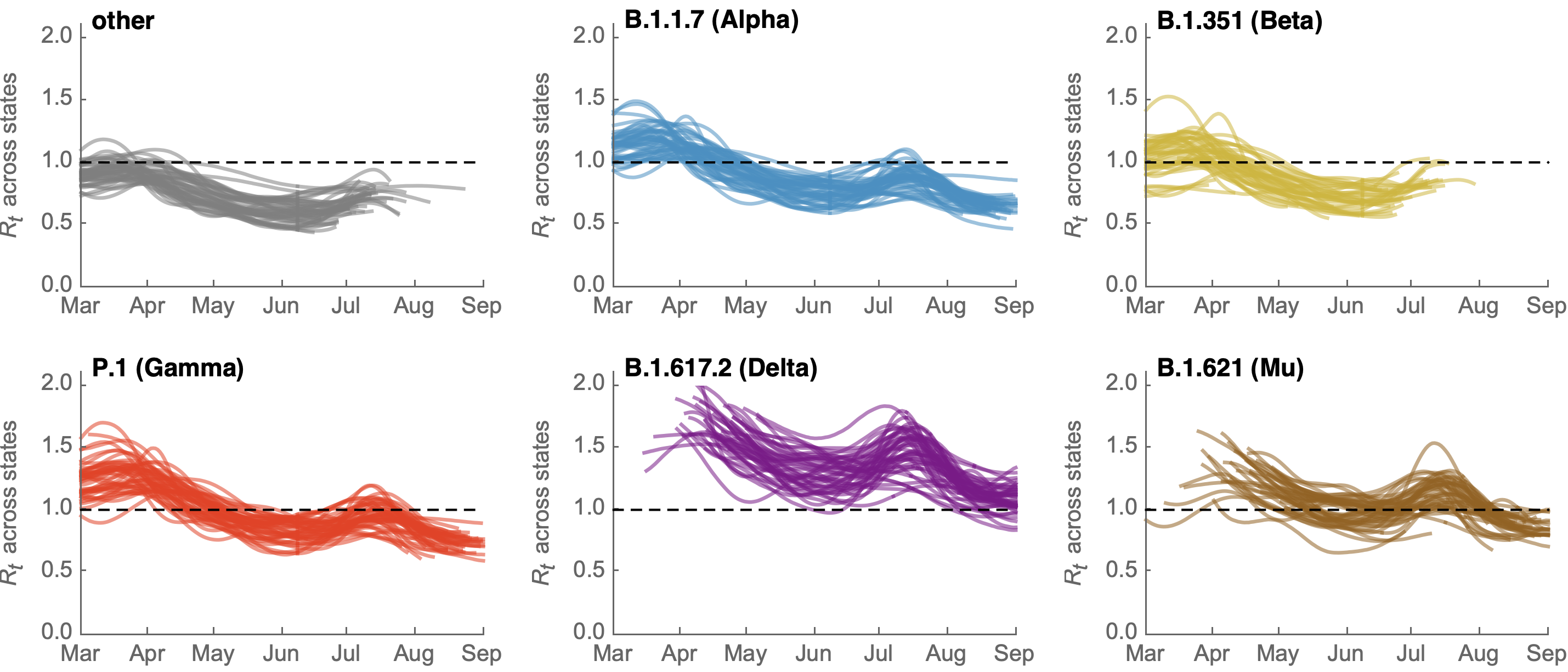
Consistent differences in variant-specific transmission rate across states
Consistent reductions in variant-specific Rt from vaccination
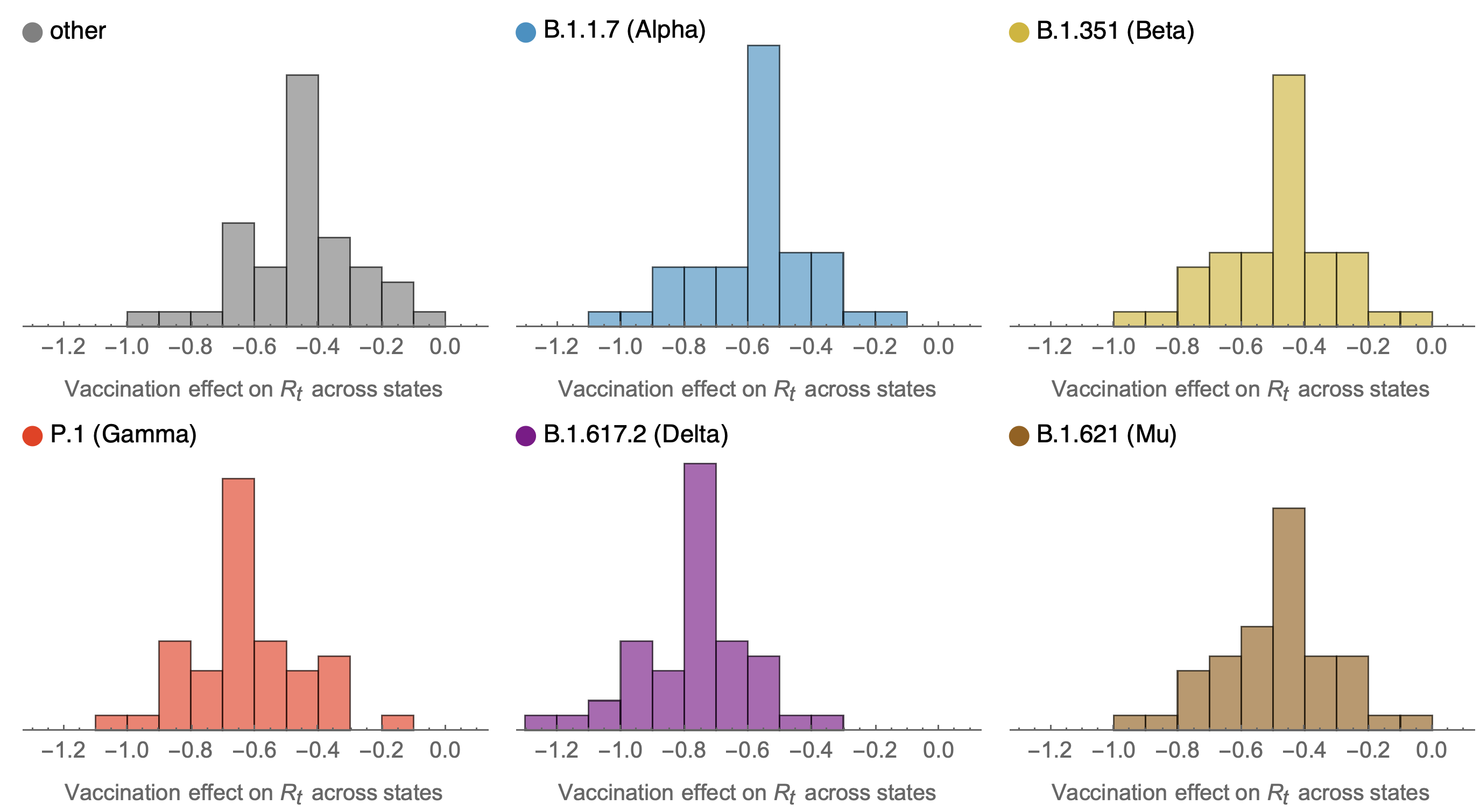
Future work would ideally tie together granular empirical estimates of viral fitness from frequency data together with mutational and phenotypic predictors to learn what is driving variant success
Omicron variant
Lineage B.1.1.539 / clade 21K / Omicron variant emerging from basal diversity
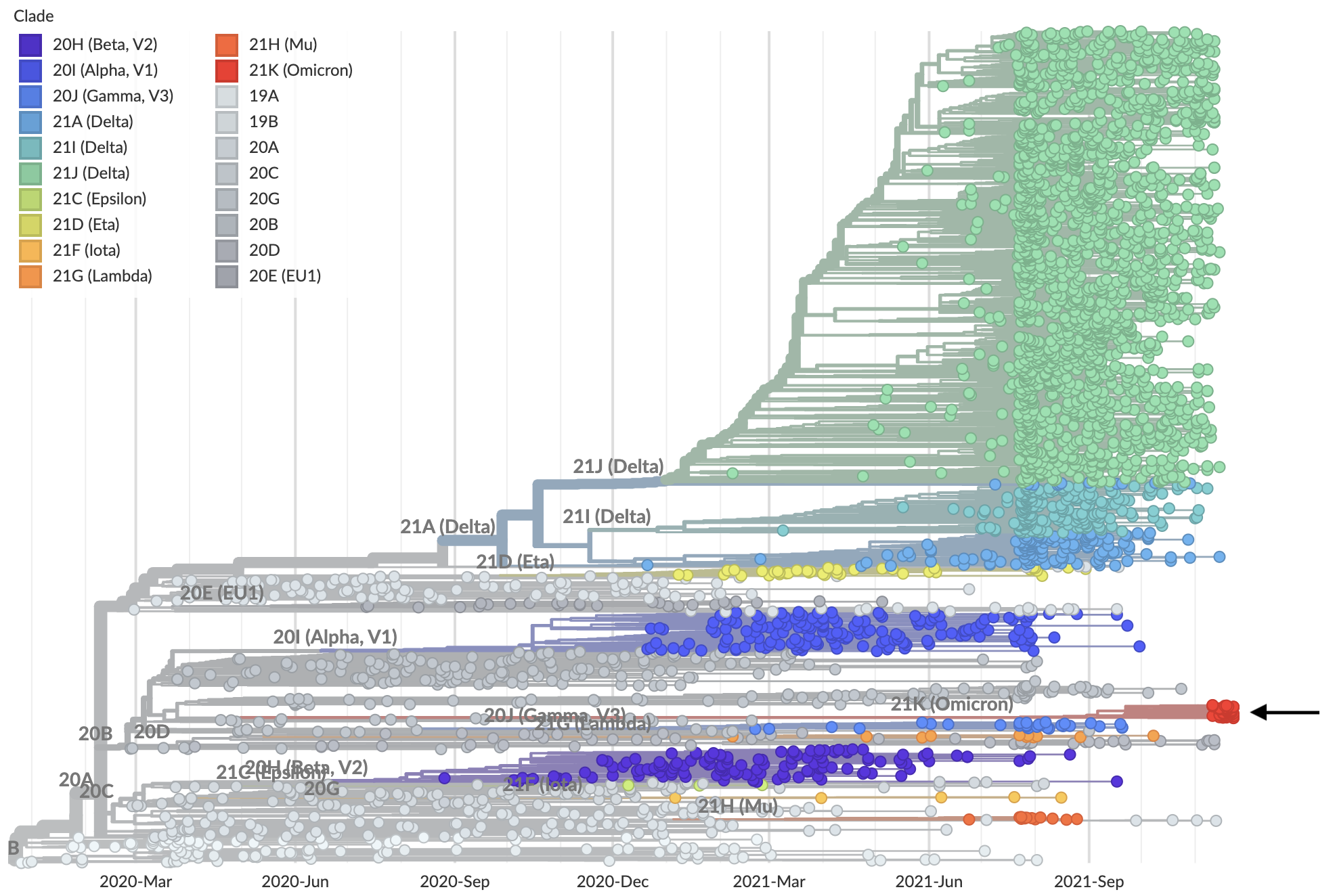
Long branch connecting closest sequenced viruses
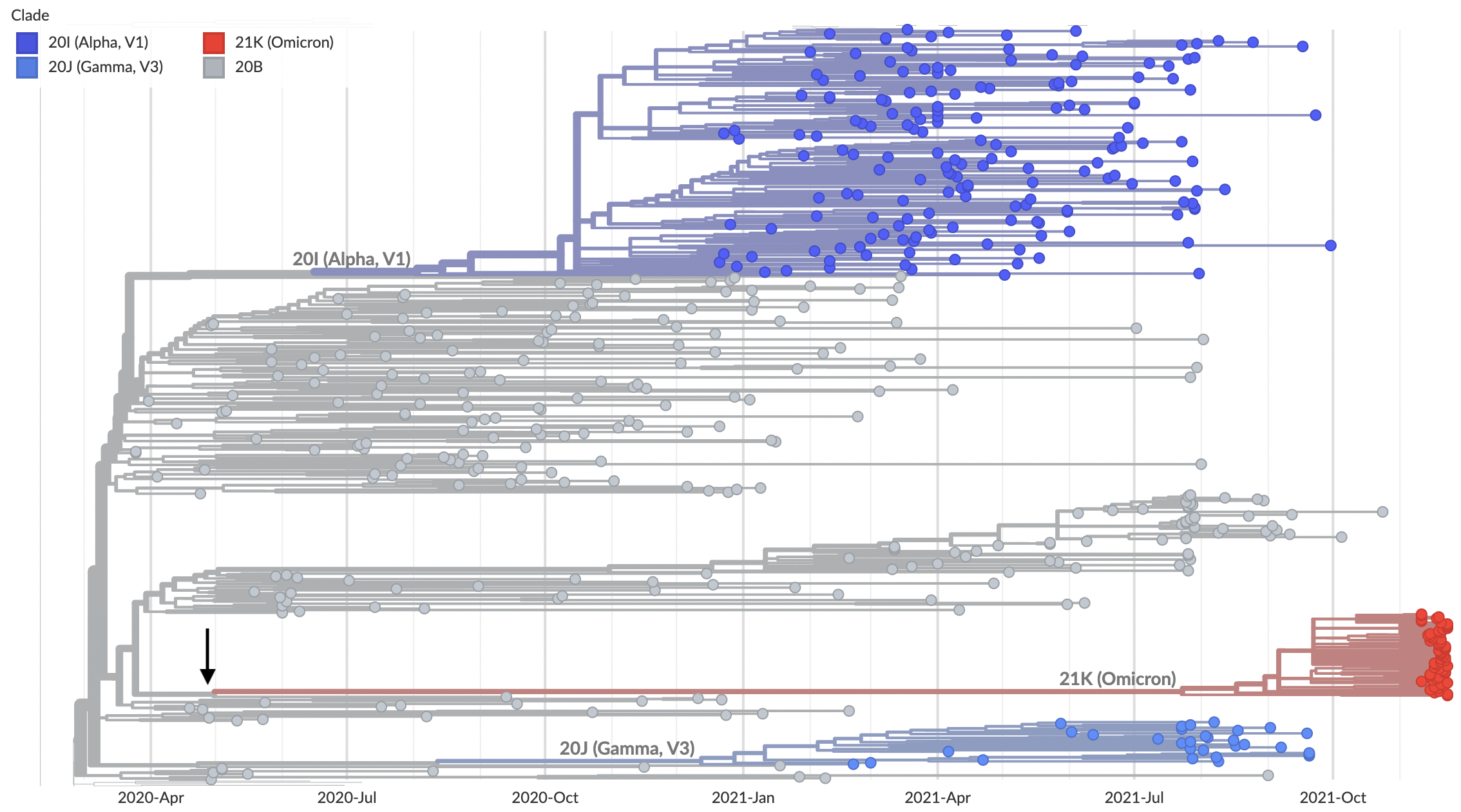
Omicron viruses with huge excess of mutations in S1
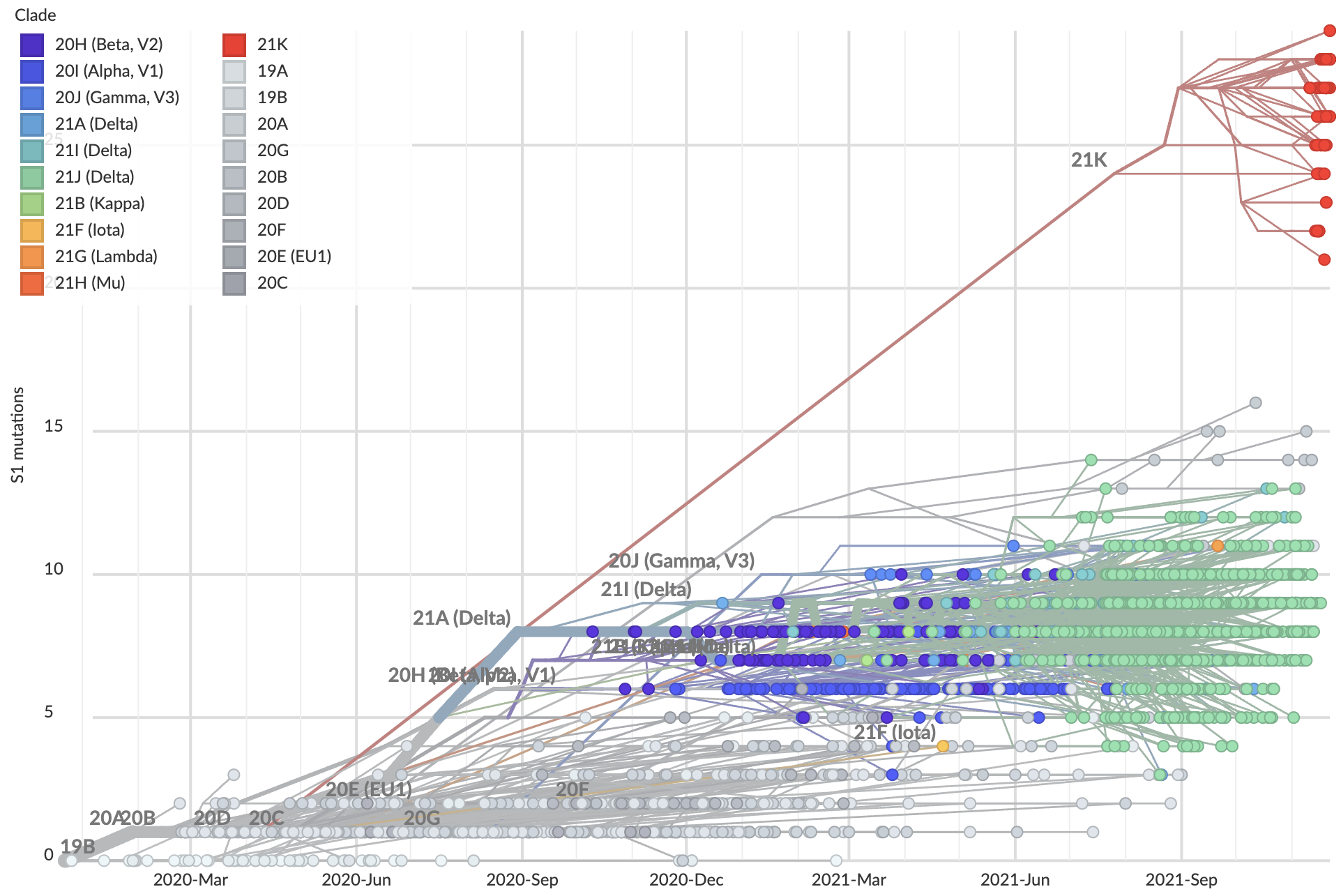
But a slight paucity of divergence in the rest of the genome
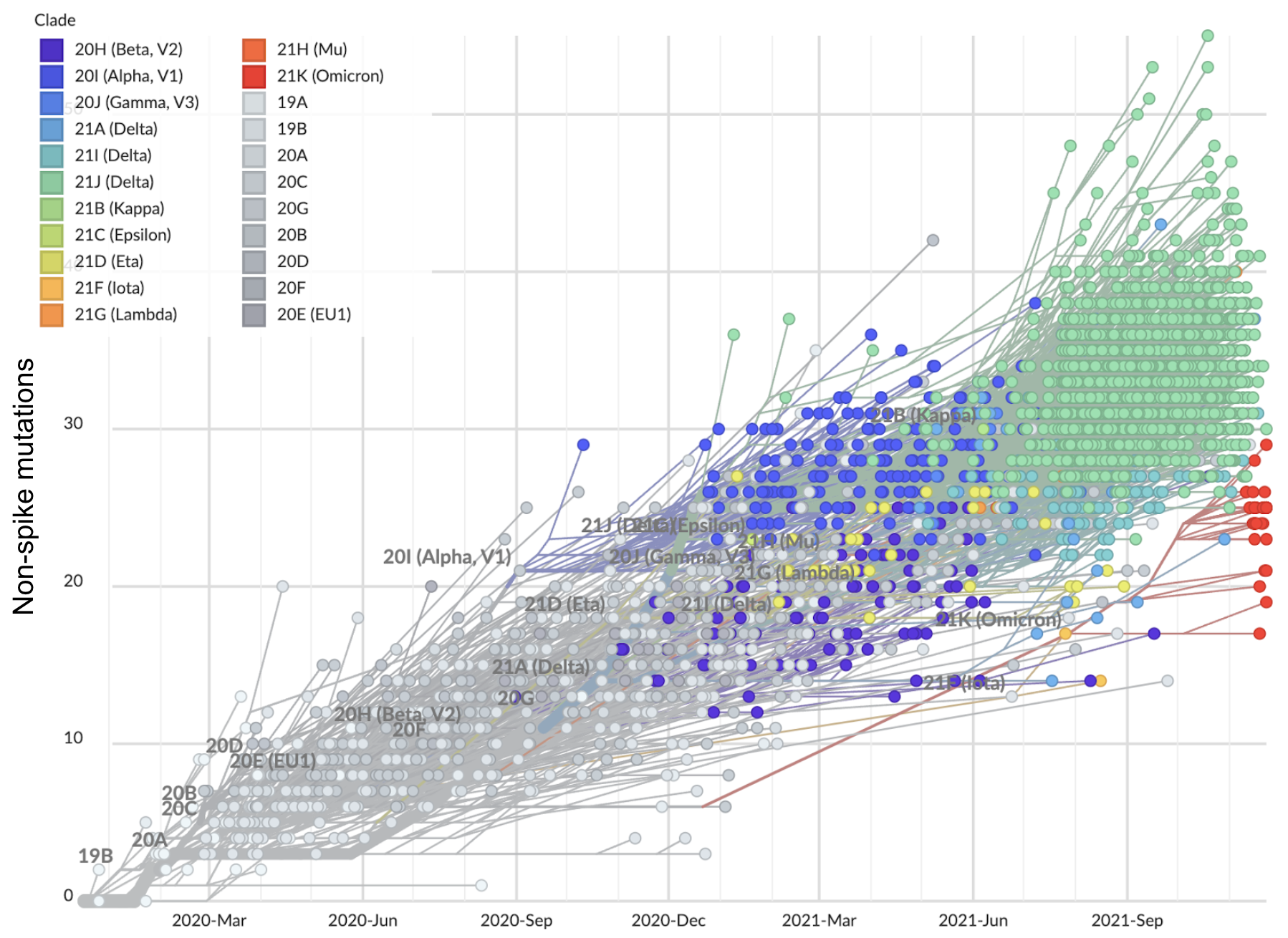
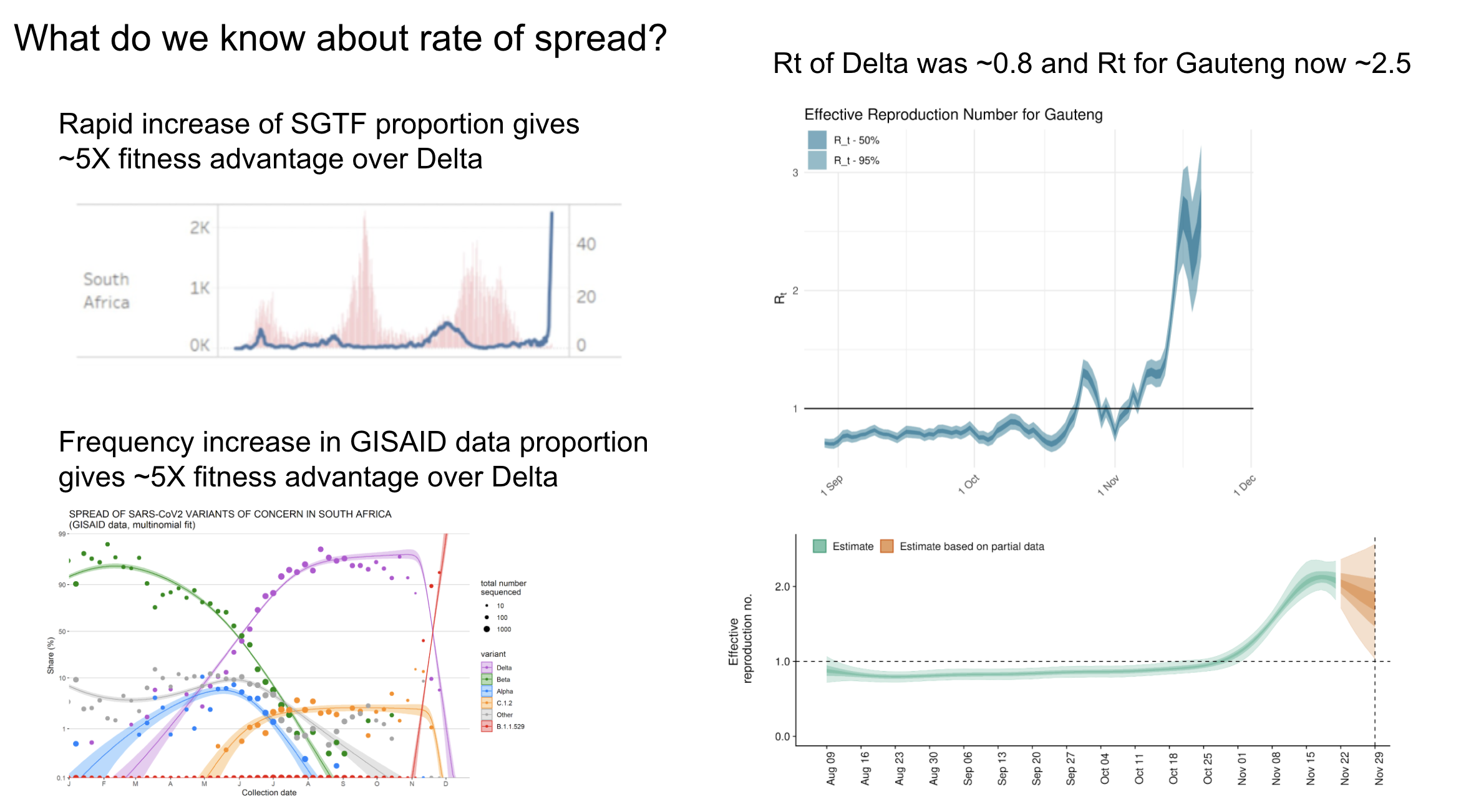
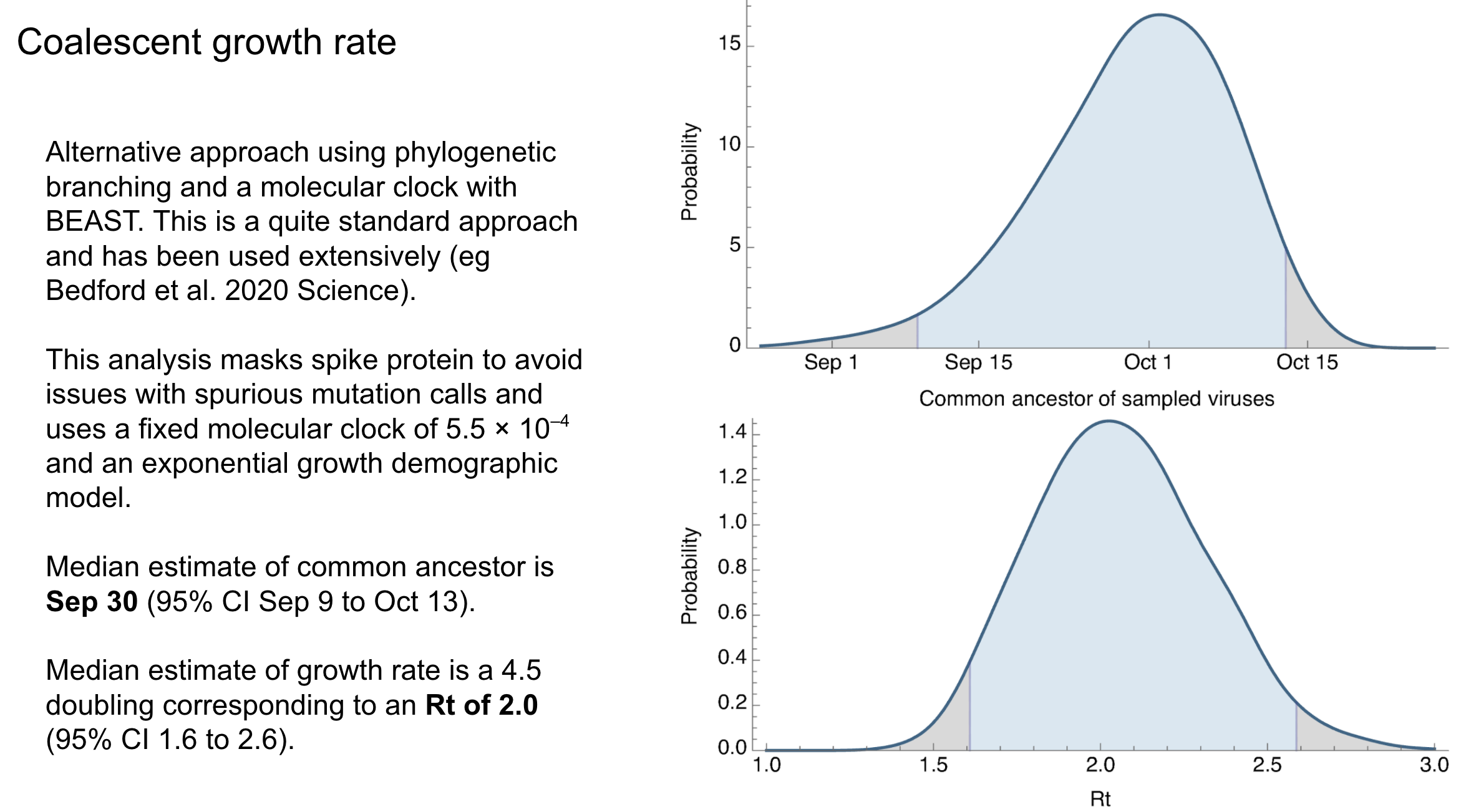
Closing thoughts

Acknowledgements
SARS-CoV-2 genomic epi: Data producers from all over the world, GISAID and the Nextstrain team
Omicron: Tulio de Oliveira and team, Centre for Epidemic Response & Innovation, National Institute For Communicable Diseases in South Africa and the Botswana Harvard HIV Reference Laboratory
Bedford Lab:
![]() John Huddleston,
John Huddleston,
![]() James Hadfield,
James Hadfield,
![]() Katie Kistler,
Katie Kistler,
![]() Louise Moncla,
Louise Moncla,
![]() Maya Lewinsohn,
Maya Lewinsohn,
![]() Thomas Sibley,
Thomas Sibley,
![]() Jover Lee,
Jover Lee,
![]() Cassia Wagner,
Cassia Wagner,
![]() Miguel Paredes,
Miguel Paredes,
![]() Nicola Müller,
Nicola Müller,
![]() Marlin Figgins,
Marlin Figgins,
![]() Denisse Sequeira,
Denisse Sequeira,
![]() Victor Lin,
Victor Lin,
![]() Jennifer Chang
Jennifer Chang






