Real-time evolutionary forecasting for influenza vaccine strain selection
Trevor Bedford (@trvrb)
2 Dec 2015
Epidemics5
Clearwater Beach, FL
Slides at bedford.io/talks/
Influenza H3N2 vaccine updates
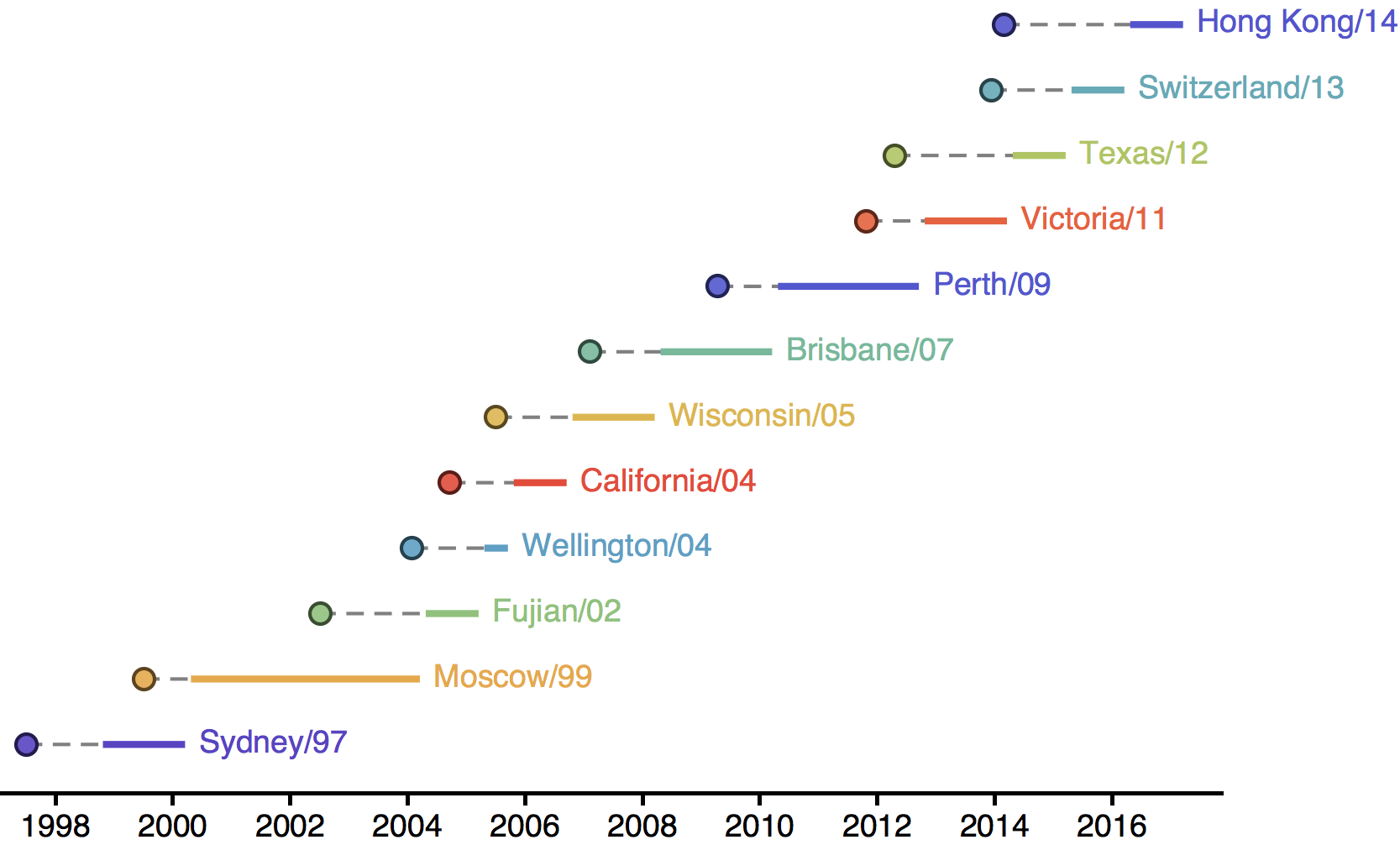
H3N2 phylogeny showing antigenic drift
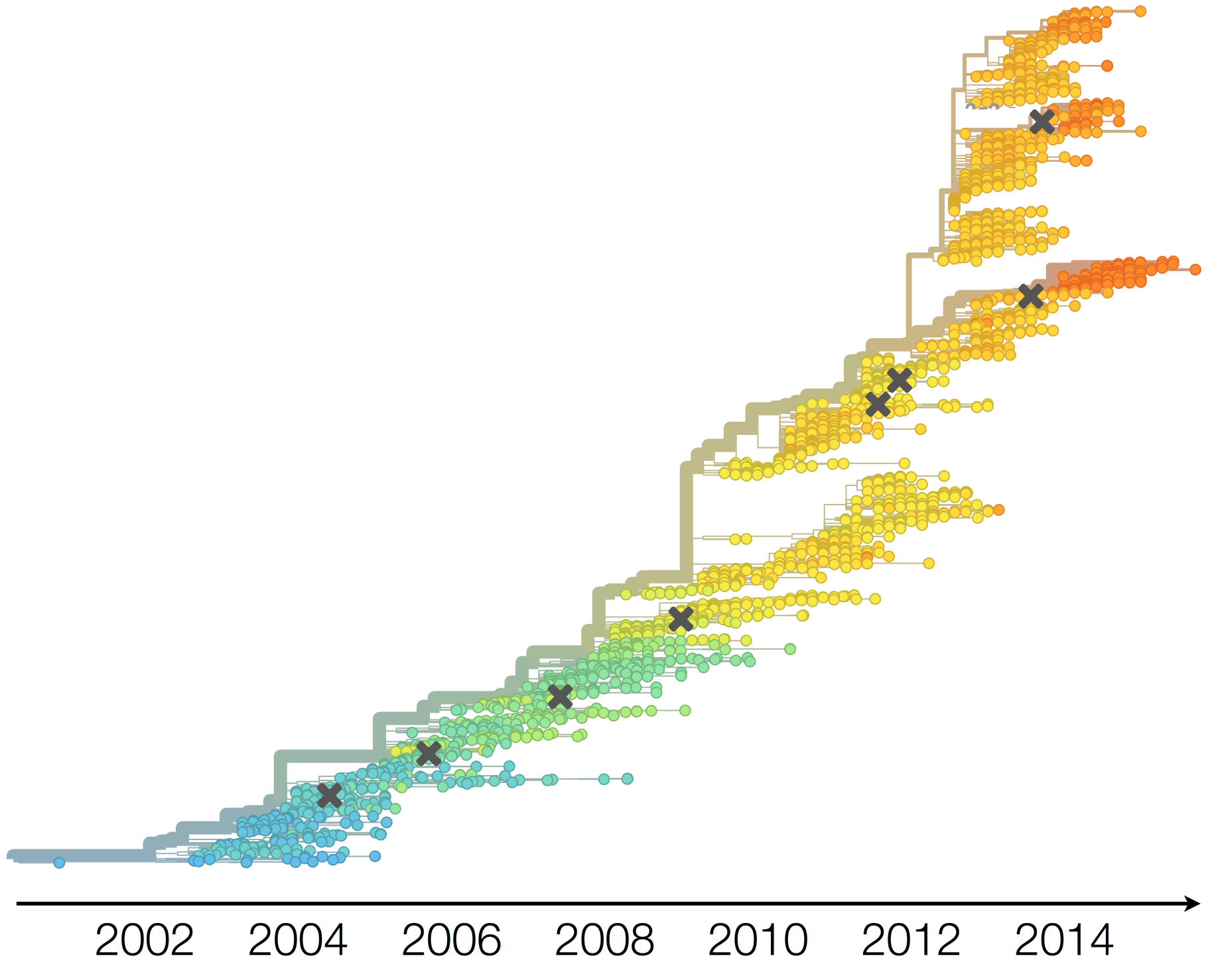
H3N2 phylogeny showing antigenic drift
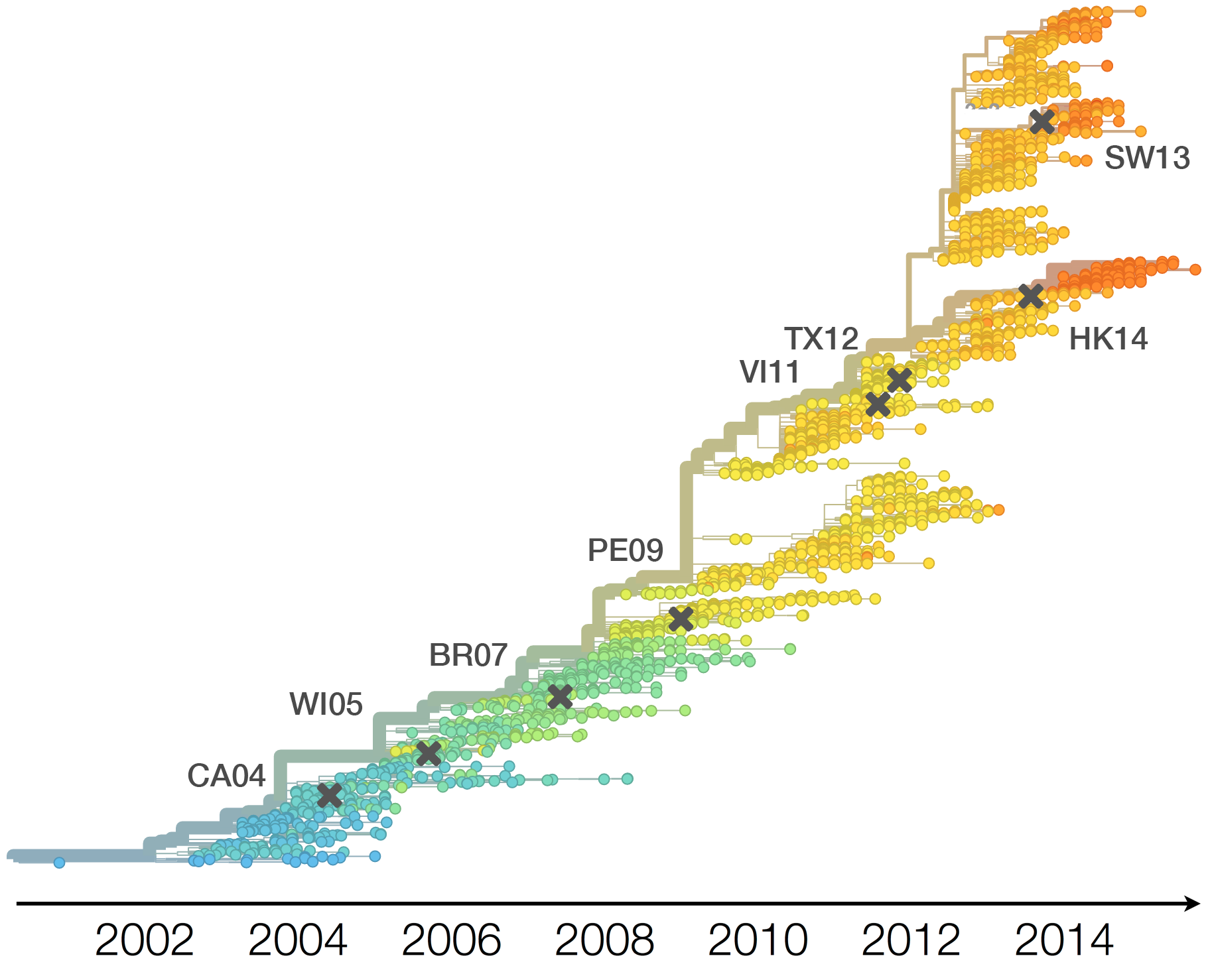
Drift variants rapidly take over the virus population
Timely surveillance and rapid analysis essential to understand ongoing influenza evolution
nextflu
Project to provide a real-time view of the evolving influenza population
All in collaboration with Richard Neher

nextflu pipeline
- Download all recent HA sequences from GISAID
- Filter to remove outliers
- Subsample across time and space
- Align sequences
- Build tree
- Estimate frequencies
- Export for visualization
Up-to-date analysis publicly available at:
nextflu.org
Forecasting
The future is here, it's just not evenly distributed yet
— William Gibson
USA music industry, 2011 dollars per capita
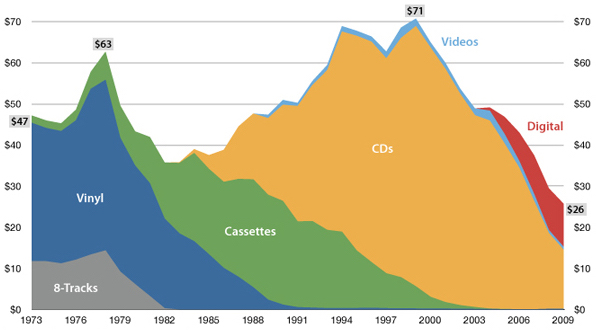
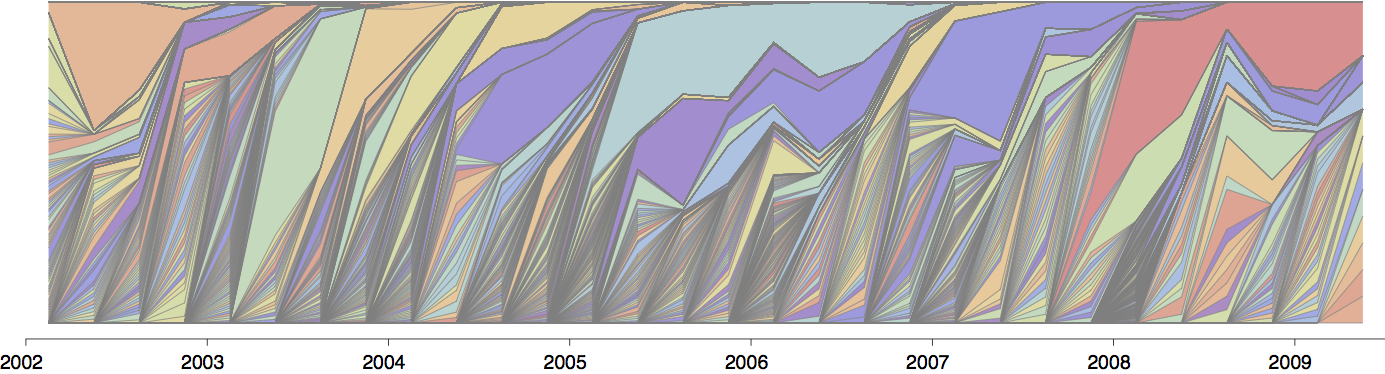
Influenza population turnover
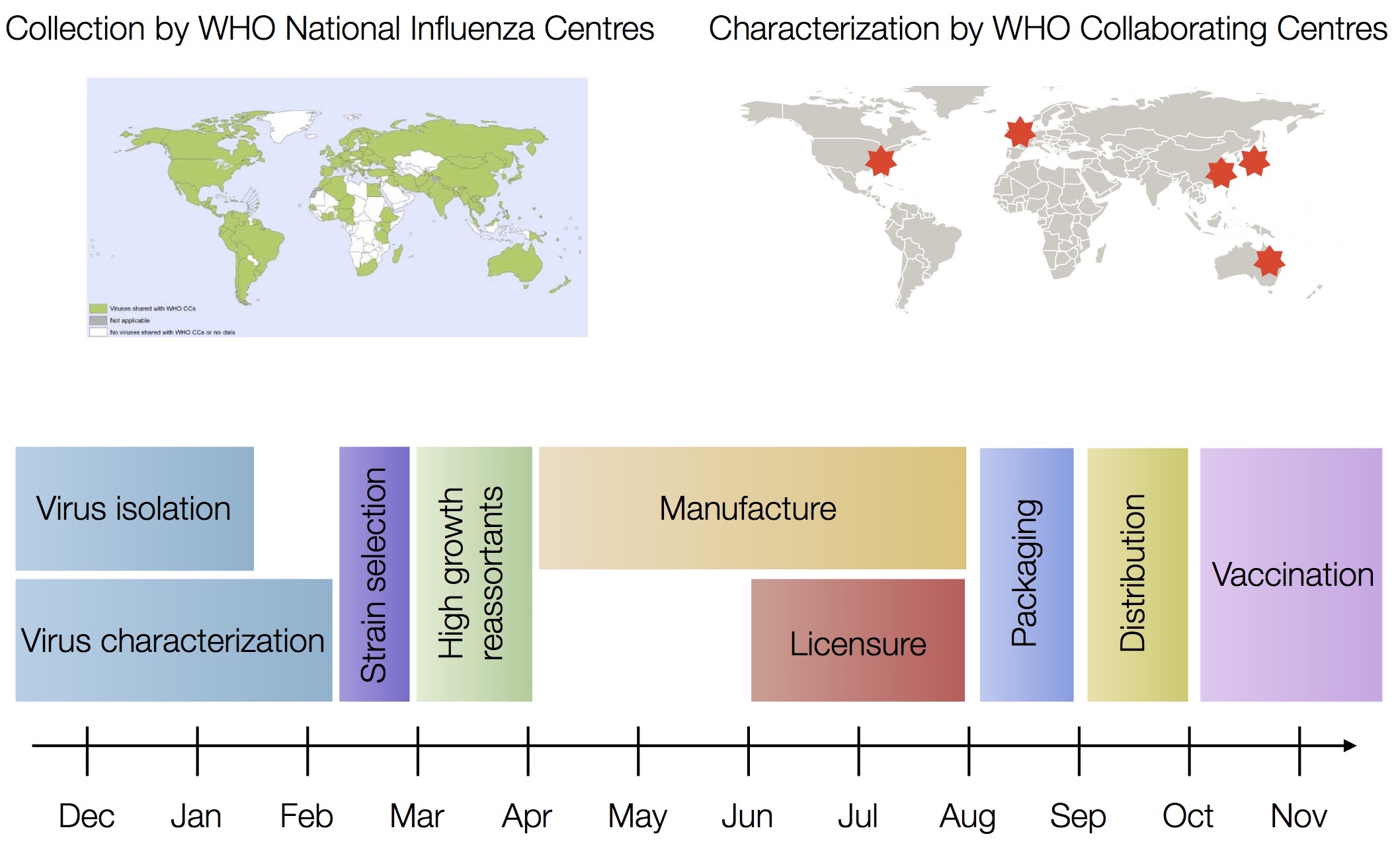
Vaccine strain selection timeline
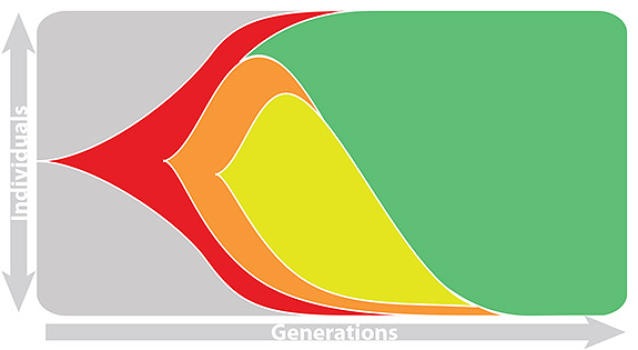
Seek to explain change in clade frequencies over 1 year
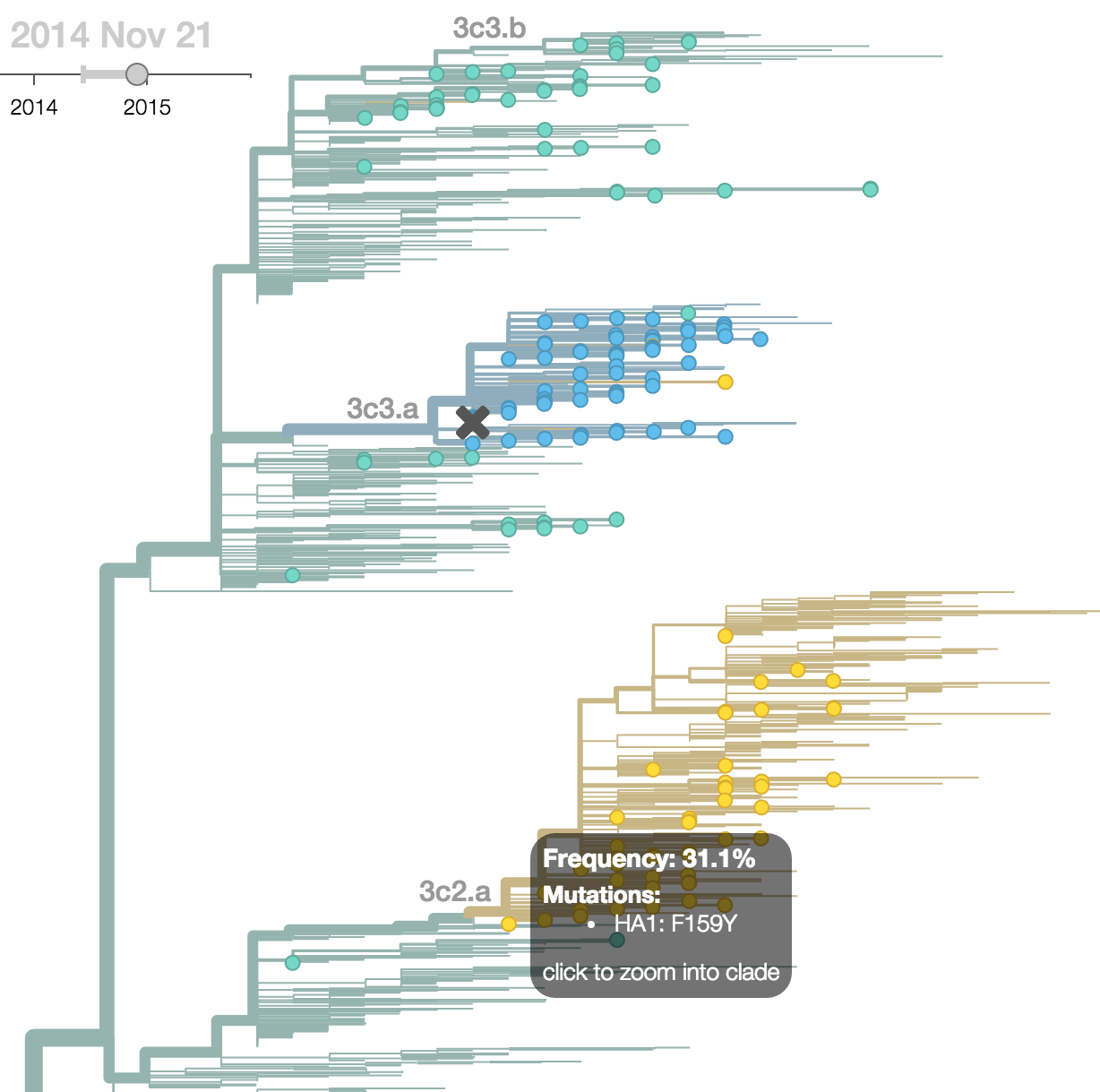
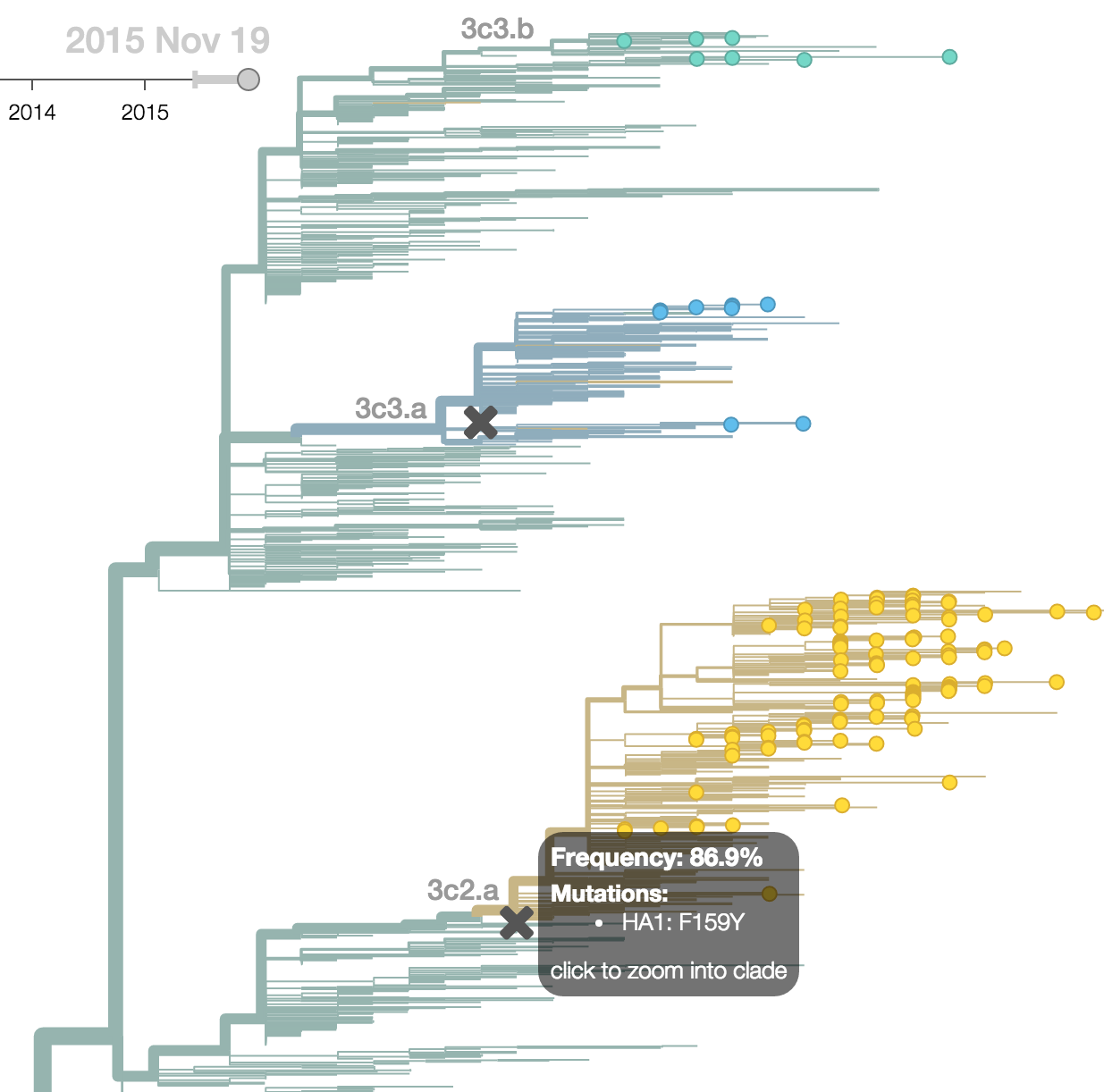
Fitness models can project clade frequencies
Clade frequencies $X$ derive from the fitnesses $f$ and frequencies $x$ of constituent viruses, such that
$$\hat{X}_v(t+\Delta t) = \sum_{i:v} x_i(t) \, \mathrm{exp}(f_i \, \Delta t)$$
This captures clonal interference between competing lineages
Predictive fitness models
A simple predictive model estimates the fitness $f$ of virus $i$ as
$$\hat{f}_i = \beta^\mathrm{ep} \, f_i^\mathrm{ep} + \beta^\mathrm{ne} \, f_i^\mathrm{ne}$$
where $f_i^\mathrm{ep}$ measures cross-immunity via substitutions at epitope sites and $f_i^\mathrm{ep}$ measures mutational load via substitutions at non-epitope sites
We implement a similar model based on two predictors
- Clade frequency change
- Antigenic advancement
Project frequencies forward,
growing clades have high fitness
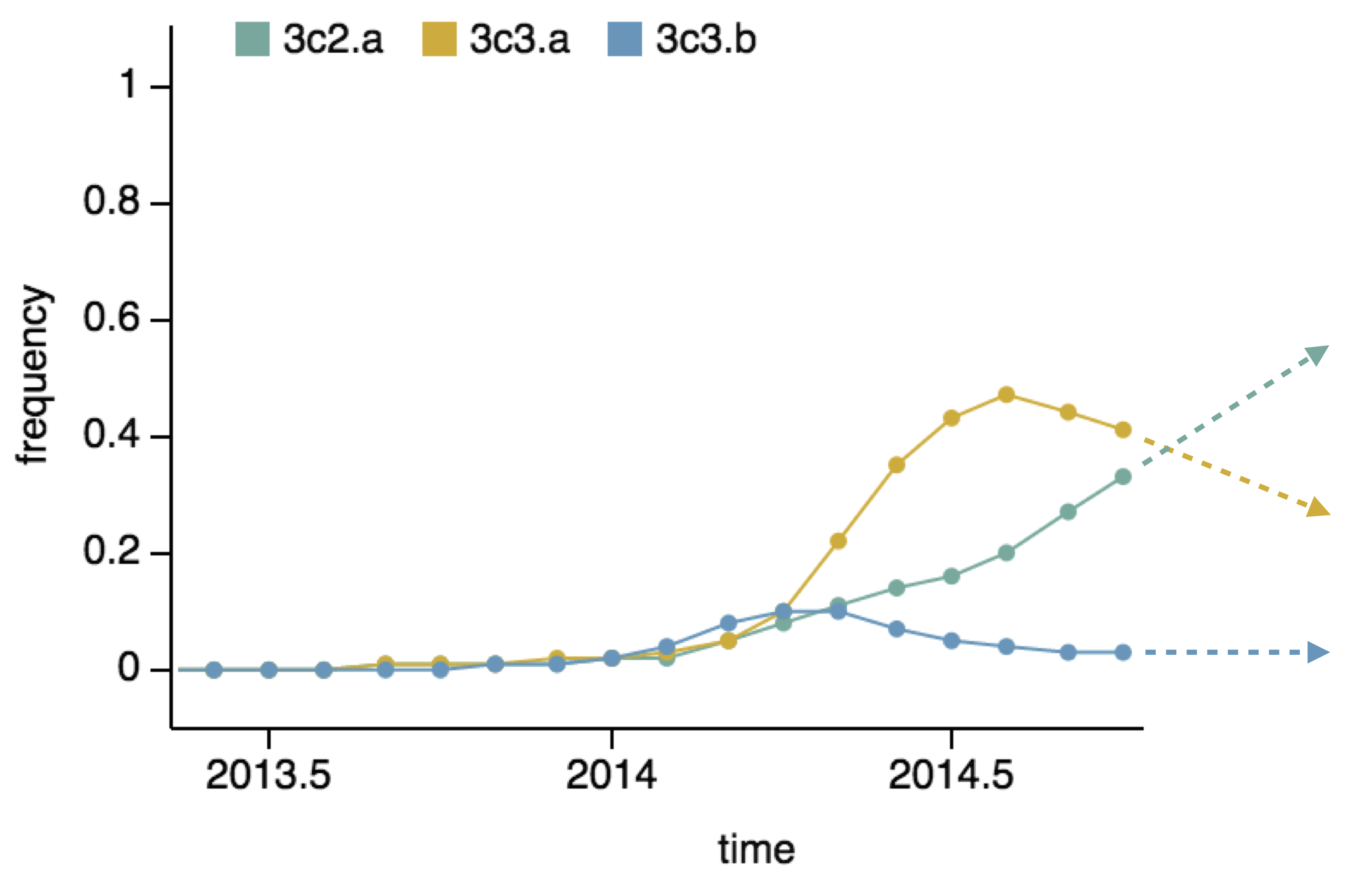
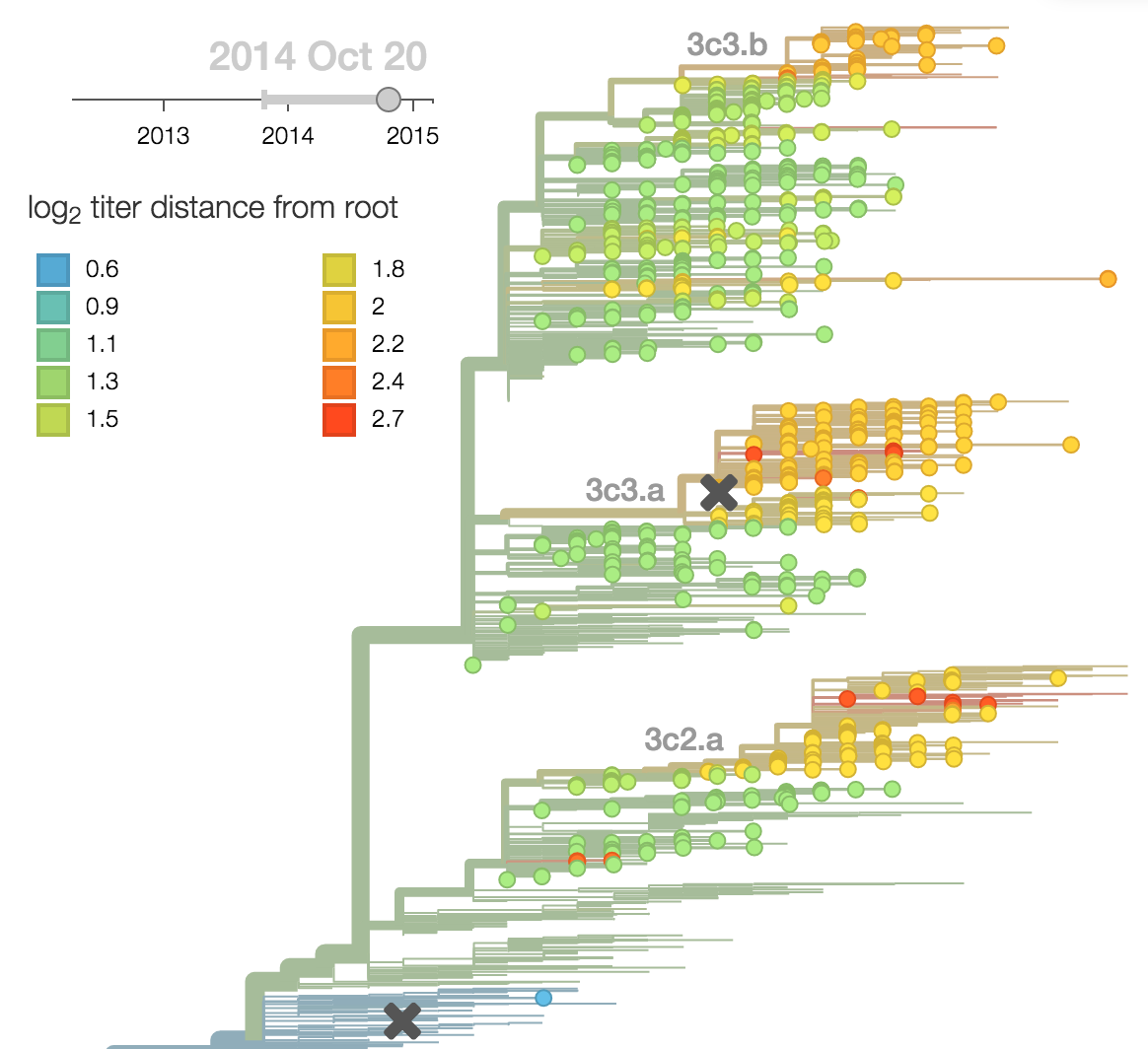
Calculate HI drop from ancestor,
drifted clades have high fitness
Fitness model parameterization
Our predictive model estimates the fitness $f$ of virus $i$ as
$$\hat{f}_i = \beta^\mathrm{freq} \, f_i^\mathrm{freq} + \beta^\mathrm{HI} \, f_i^\mathrm{HI}$$
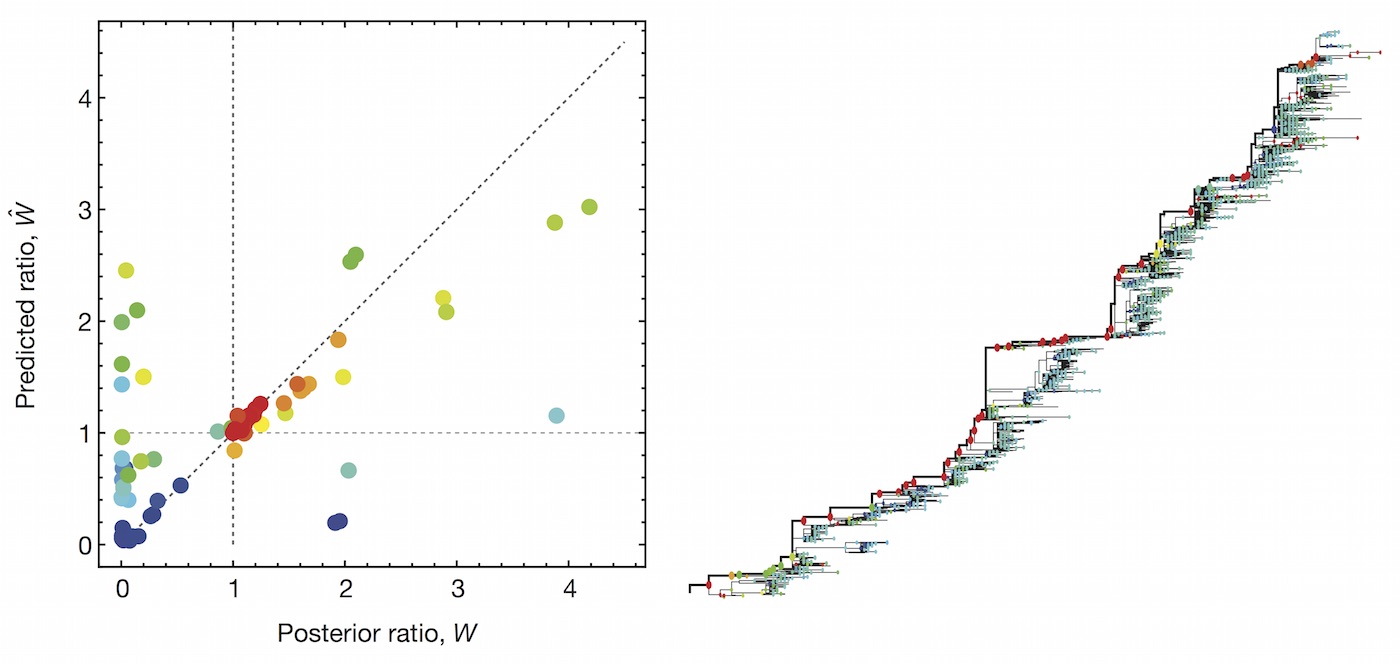
We learn coefficients and validate model based on previous 15 H3N2 seasons
Clade growth and decline is well predicted
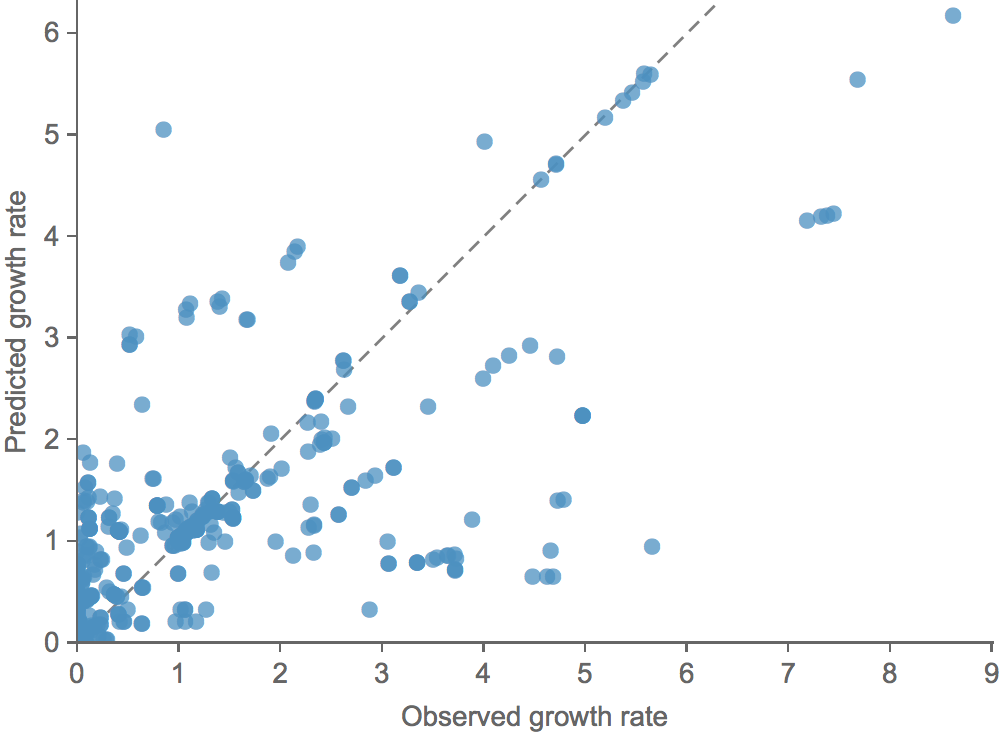
Correct in 81% of cases
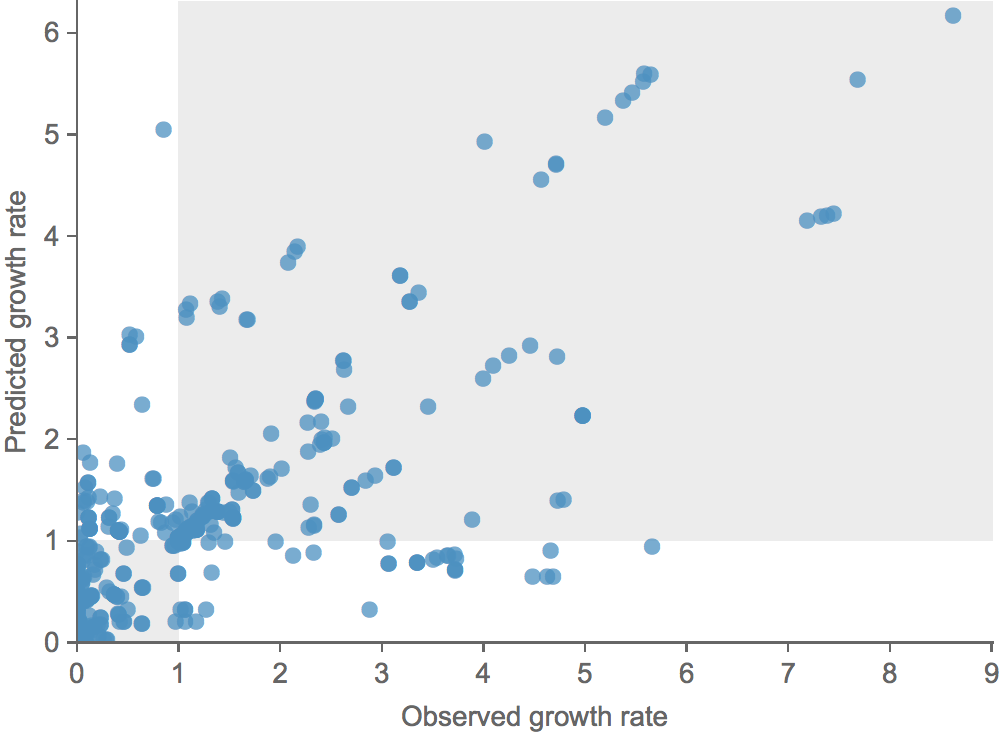
Prototype up at
dev.nextflu.org
Further work on predictive modeling
- Integrate data predictors and data sources, e.g. plan to investigate a geographic predictor
- Possible to build predictive models for H1N1 and B and to forecast NA evolution
Evolutionary analyses can inform influenza vaccine strain selection
Analyses must be rapid and widely available
Predictive models can flag clades for experimental follow-up and creation of vaccine candidates
Acknowledgements
WHO Global Influenza Surveillance Network, GISAID, Richard Neher, Andrew Rambaut, John McCauley, Rod Daniels, Ian Barr, Masato Tashiro, Colin Russell, Sebastian Maurer-Stroh, Michael Lässig, Marta Łuksza



Contact
- Website: bedford.io
- Twitter: @trvrb
- Slides: bedford.io/talks/real-time-forecasting-epidemics/