Key insights from genomic epidemiology
Trevor Bedford
Fred Hutchinson Cancer Center / Howard Hughes Medical Institute
25 Aug 2023
CDC-PGCoE visit
Northwest PGCoE
Pathogen genomes may reveal
- Evolution of new adaptive variants
- Epidemic origins
- Patterns of geographic spread
- Animal-to-human spillover
- Transmission chains
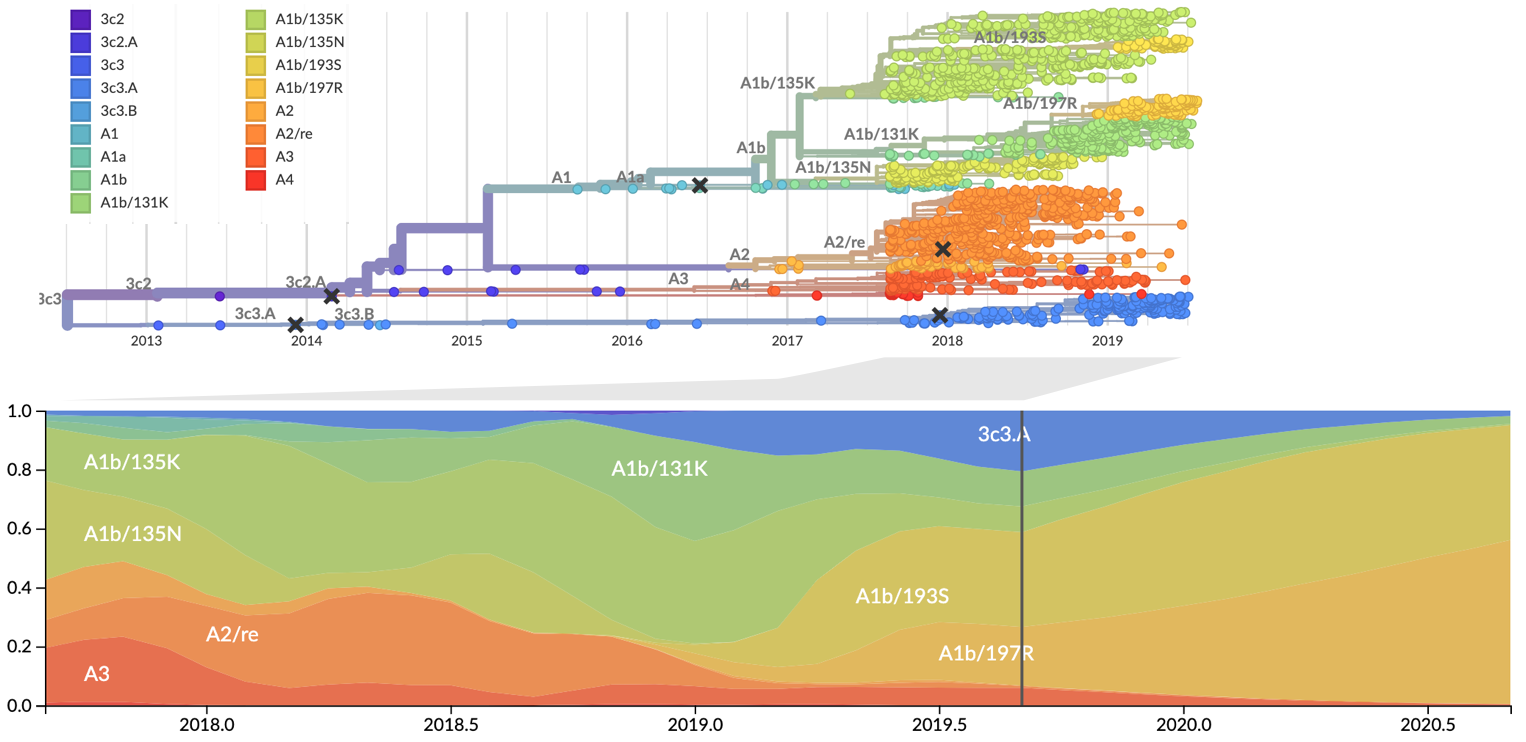
Influenza: Forecasting spread of new variants for vaccine strain selection
Zika: Uncovering origins of the epidemic in the Americas
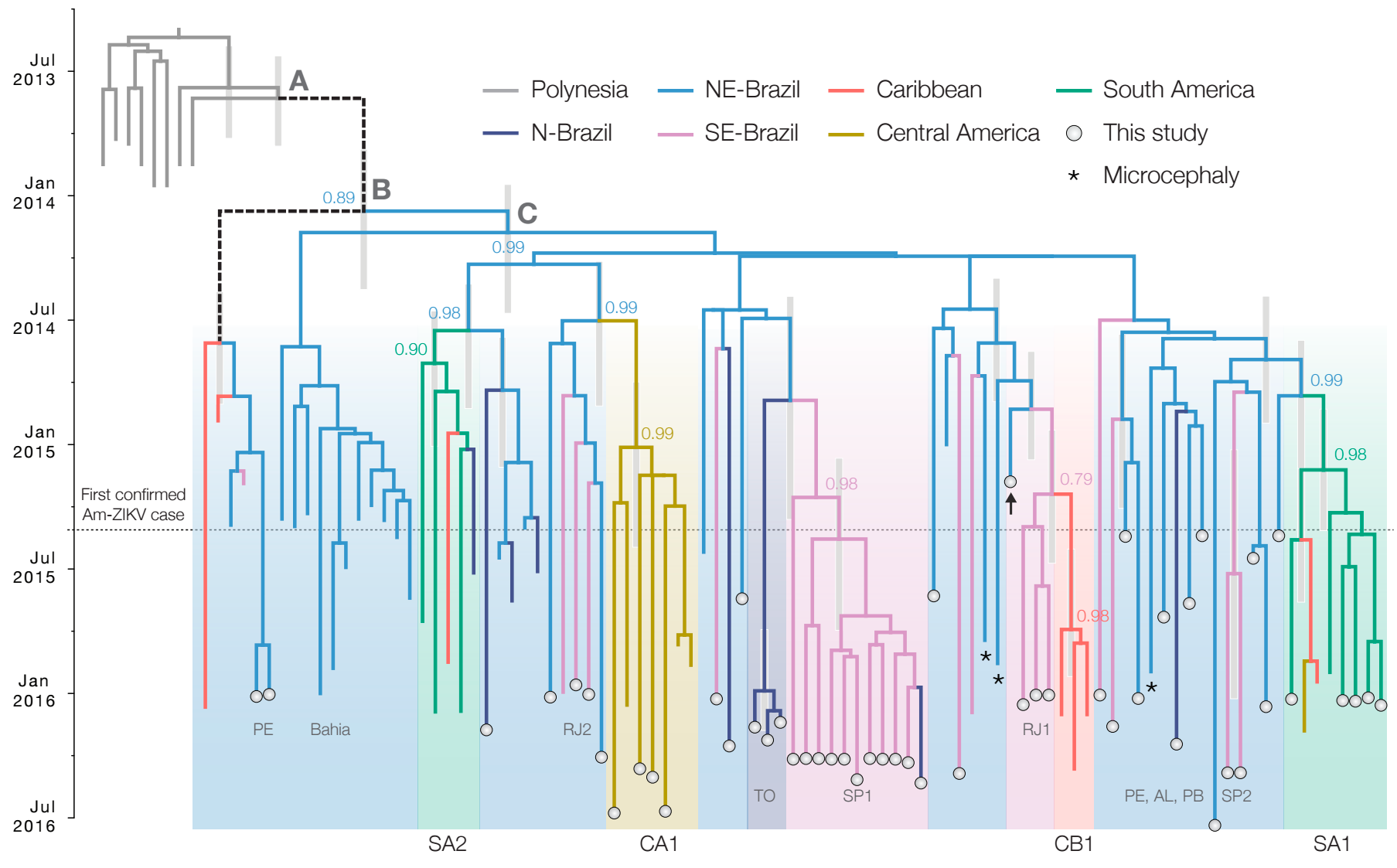
Ebola: Revealing spatial spread and persistence in West Africa
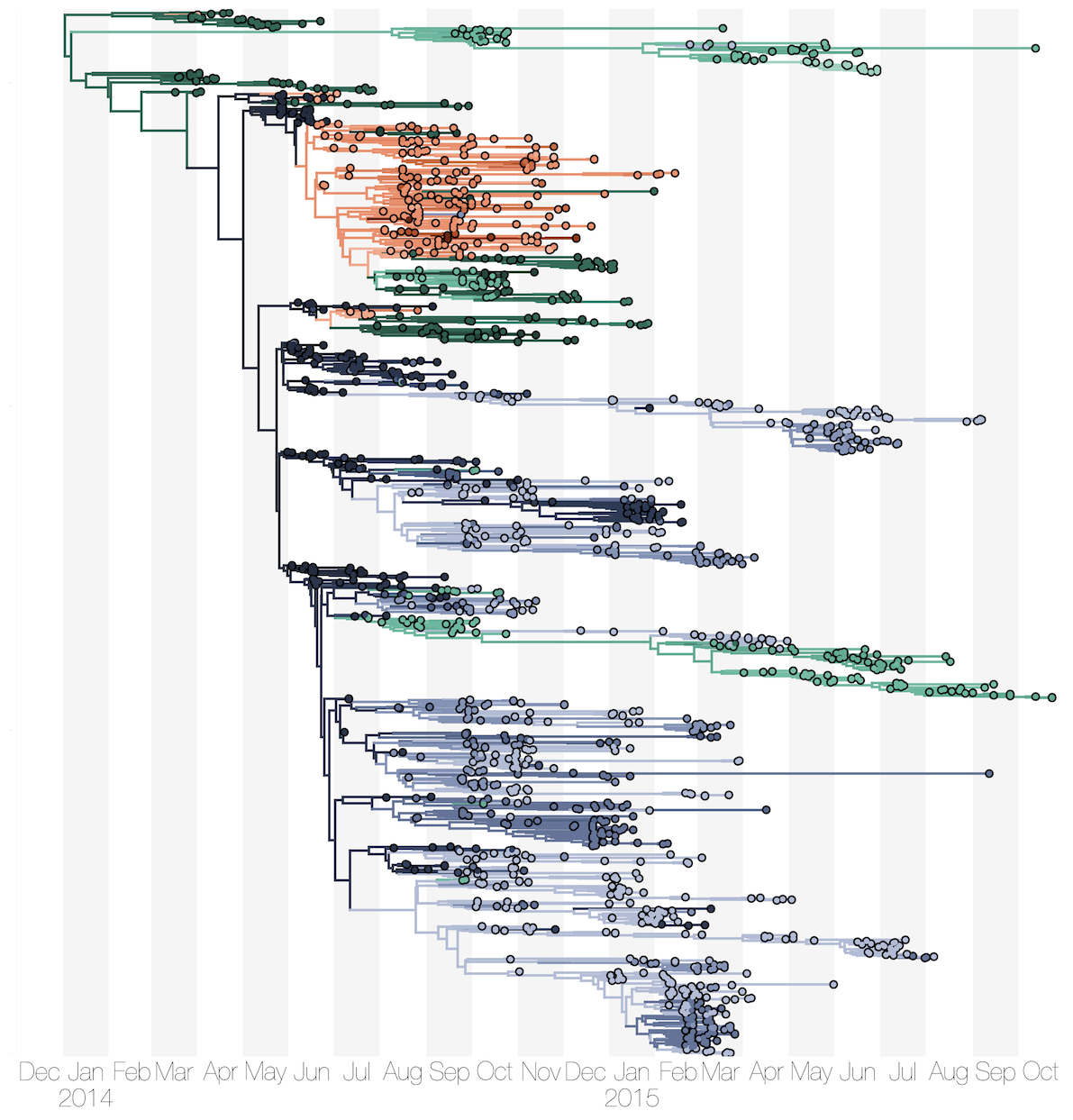
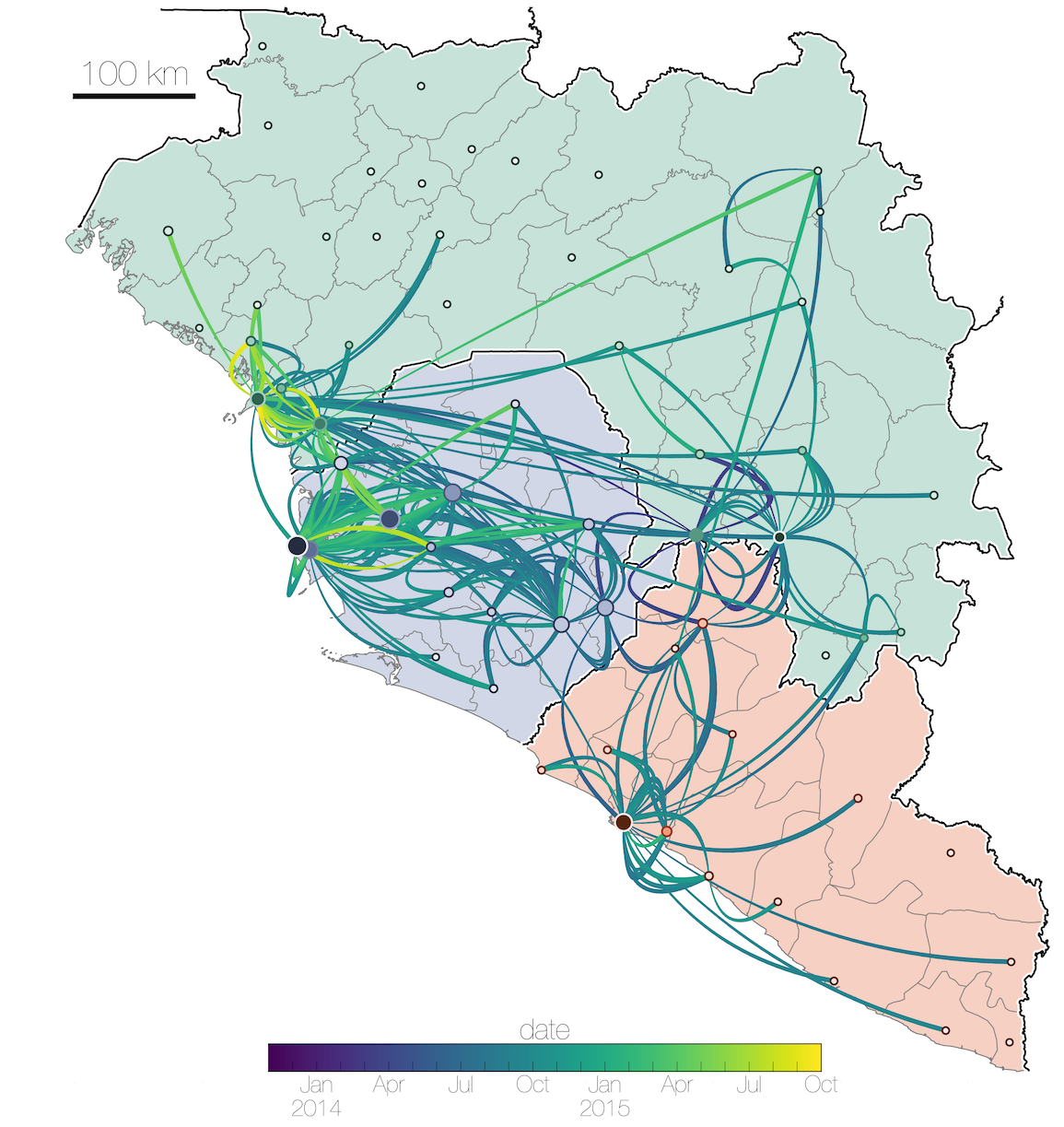
MERS: Repeated spillover into the human population from camel reservoir
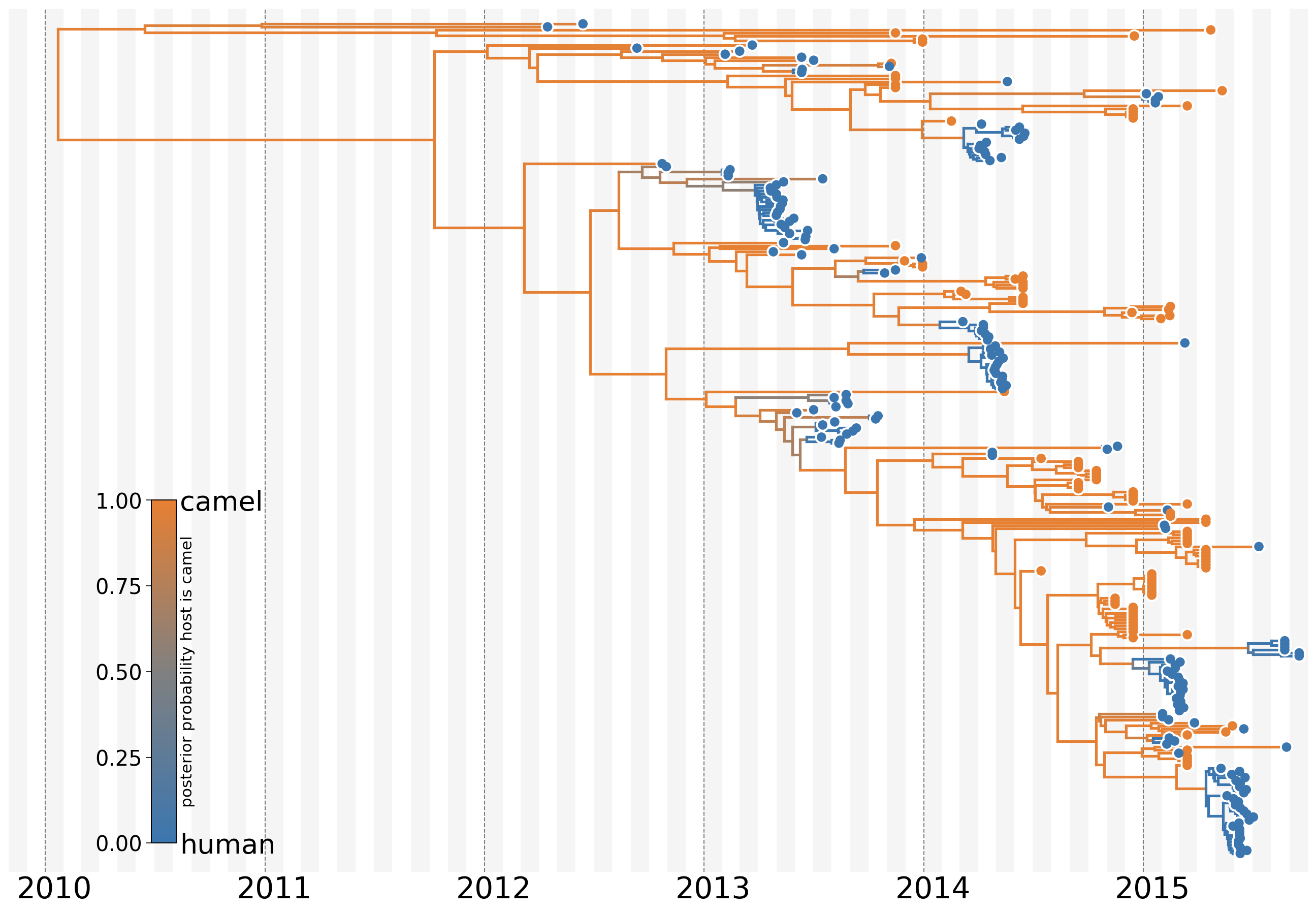
Actionable inferences
Nextstrain
Project to conduct real-time genomic epidemiology and evolutionary analysis of emerging epidemics
Nextstrain architecture
Central aims: (1) rapid and flexible bioinformatic workflows, (2) interactive visualization and (3) always up-to-date analyses at nextstrain.org
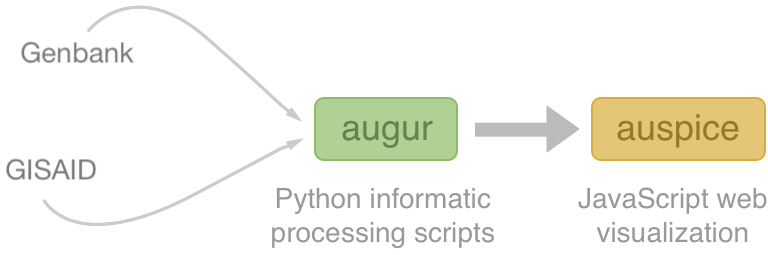
All code open source at github.com/nextstrain
Operationalized during the 2018-2020 outbreak of Ebola in the Democratic Republic of the Congo

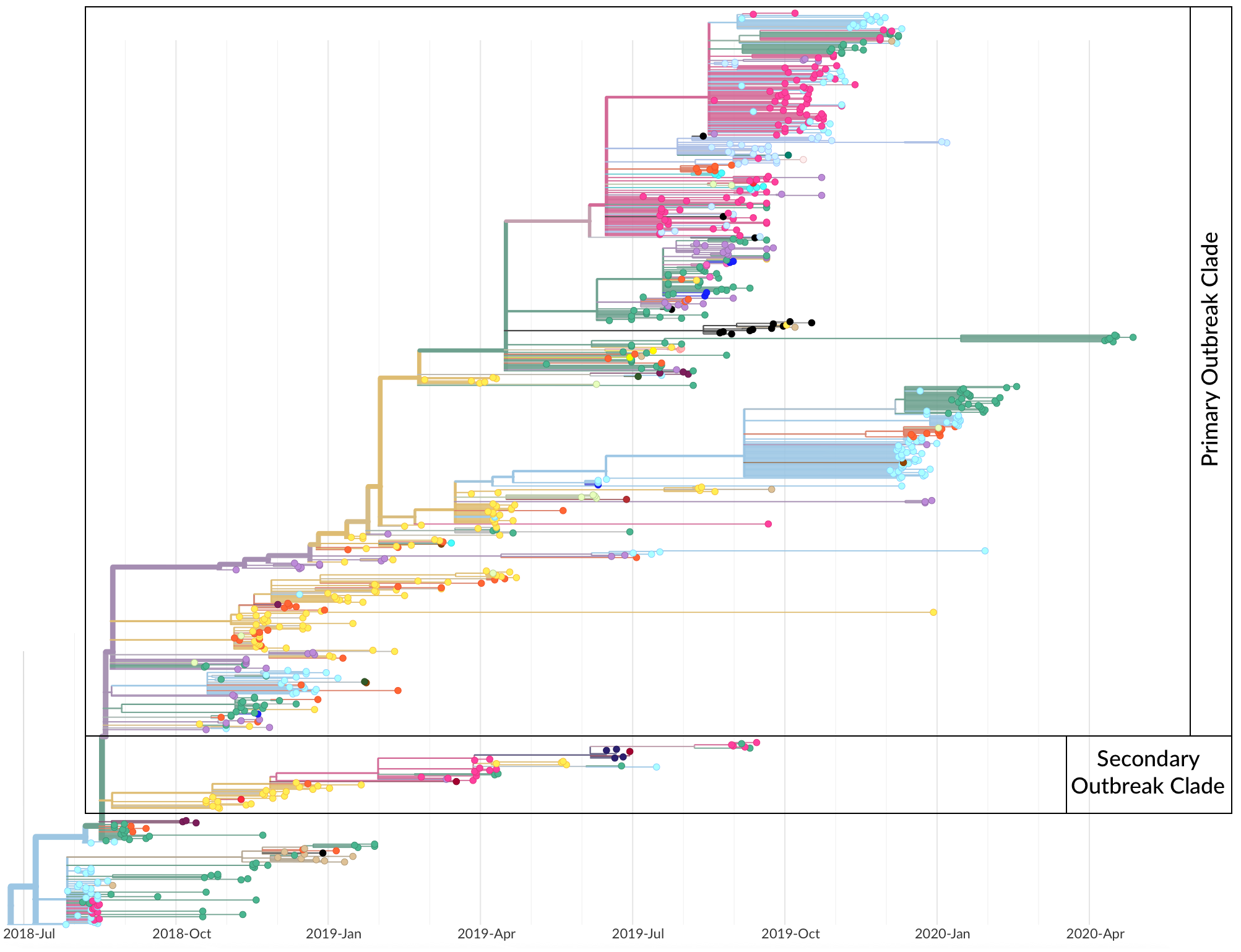
Genomic epidemiology during the COVID-19 pandemic
Over 15M SARS-CoV-2 genomes shared to GISAID and evolution tracked in real-time at nextstrain.org
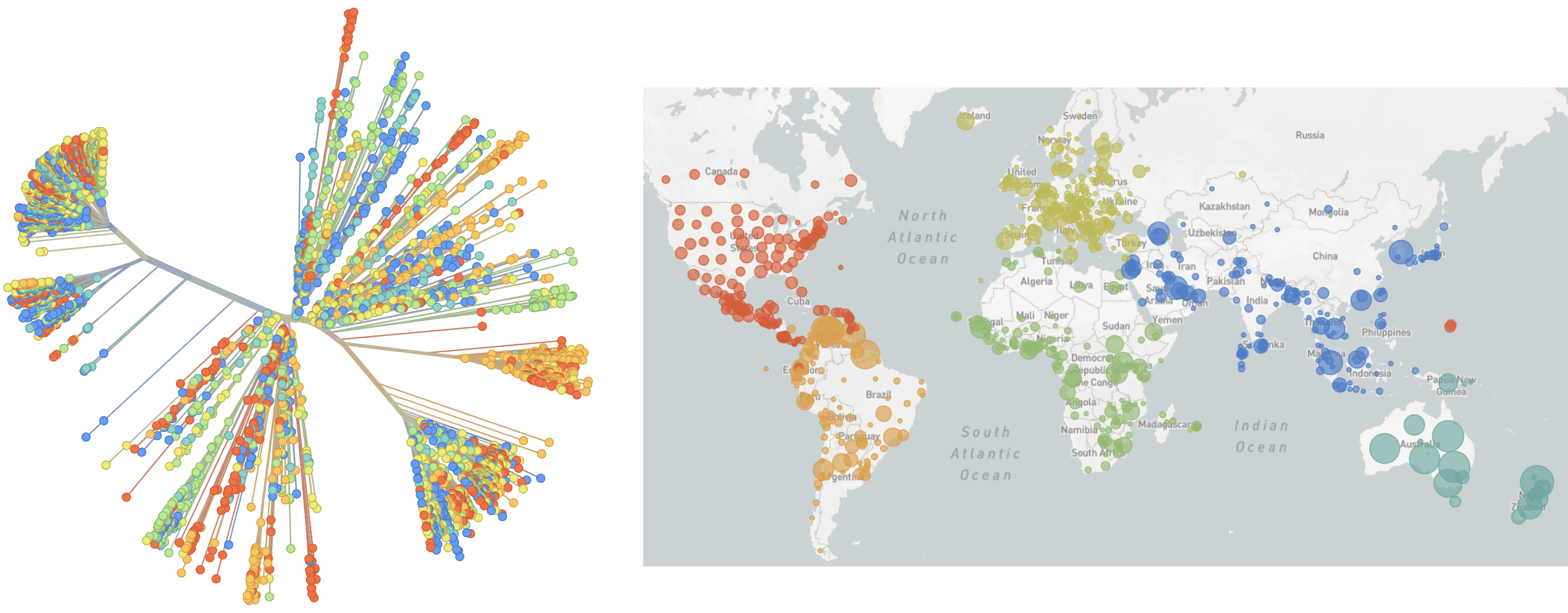
![]() Richard Neher,
Richard Neher,
![]() Ivan Aksamentov,
Ivan Aksamentov,
![]() Jennifer Chang
Jennifer Chang
![]() James Hadfield,
James Hadfield,
![]() Emma Hodcroft,
Emma Hodcroft,
![]() John Huddleston,
John Huddleston,
![]() Jover Lee,
Jover Lee,
![]() Victor Lin,
Victor Lin,
![]() Cornelius Roemer,
Cornelius Roemer,
![]() Thomas Sibley
Thomas Sibley
Three key insights that genomic epi provided during pandemic
- Rapid human-to-human spread in Wuhan beyond initial market outbreak
- Extensive local transmission while testing was rare
- Identification of variants of concern and mapping of increased transmission rates
Jan 11: First five genomes showed a novel SARS-like coronavirus
Initially thought clustering due to epi investigation of linked cases at Huanan seafood market
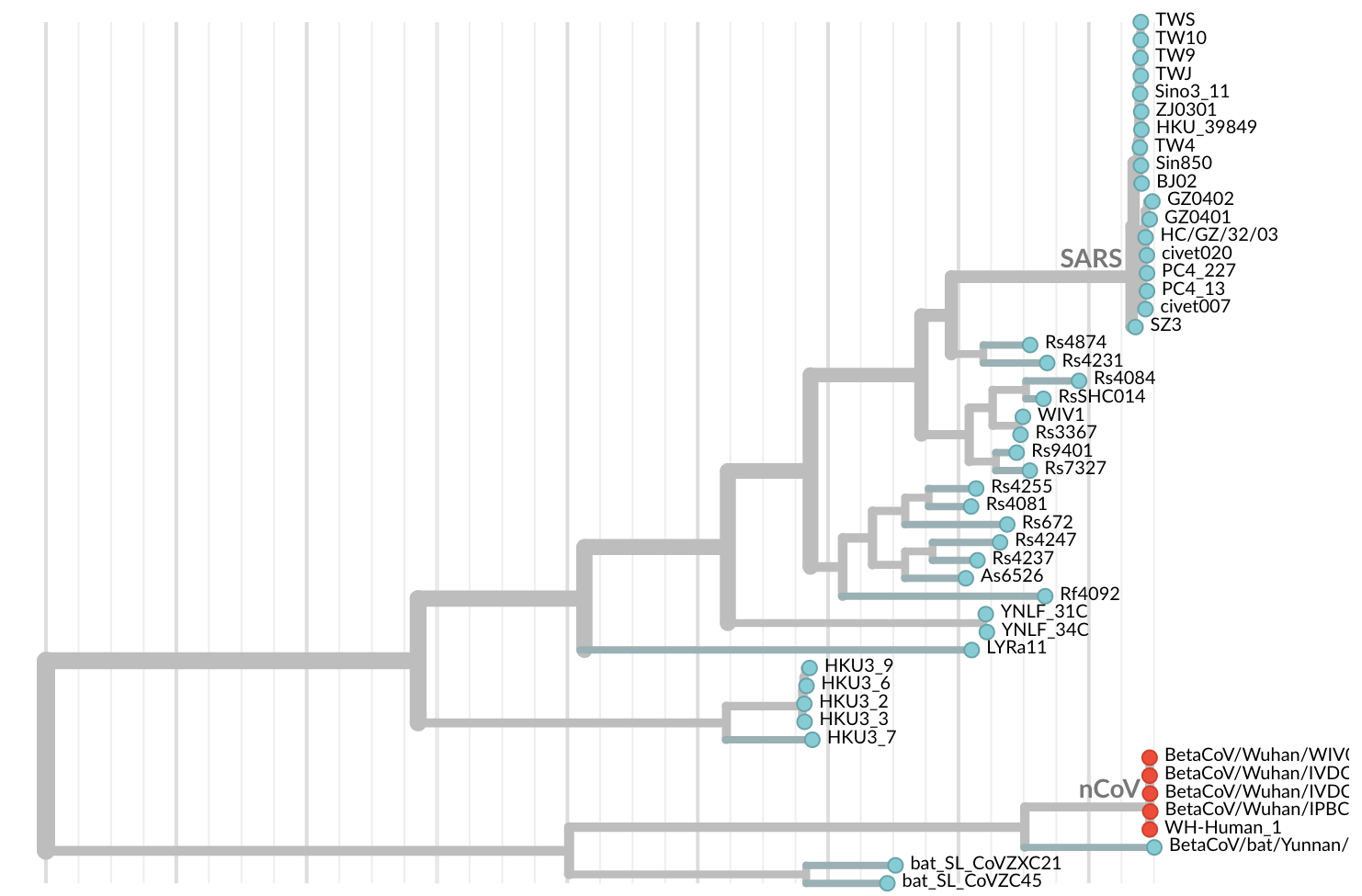
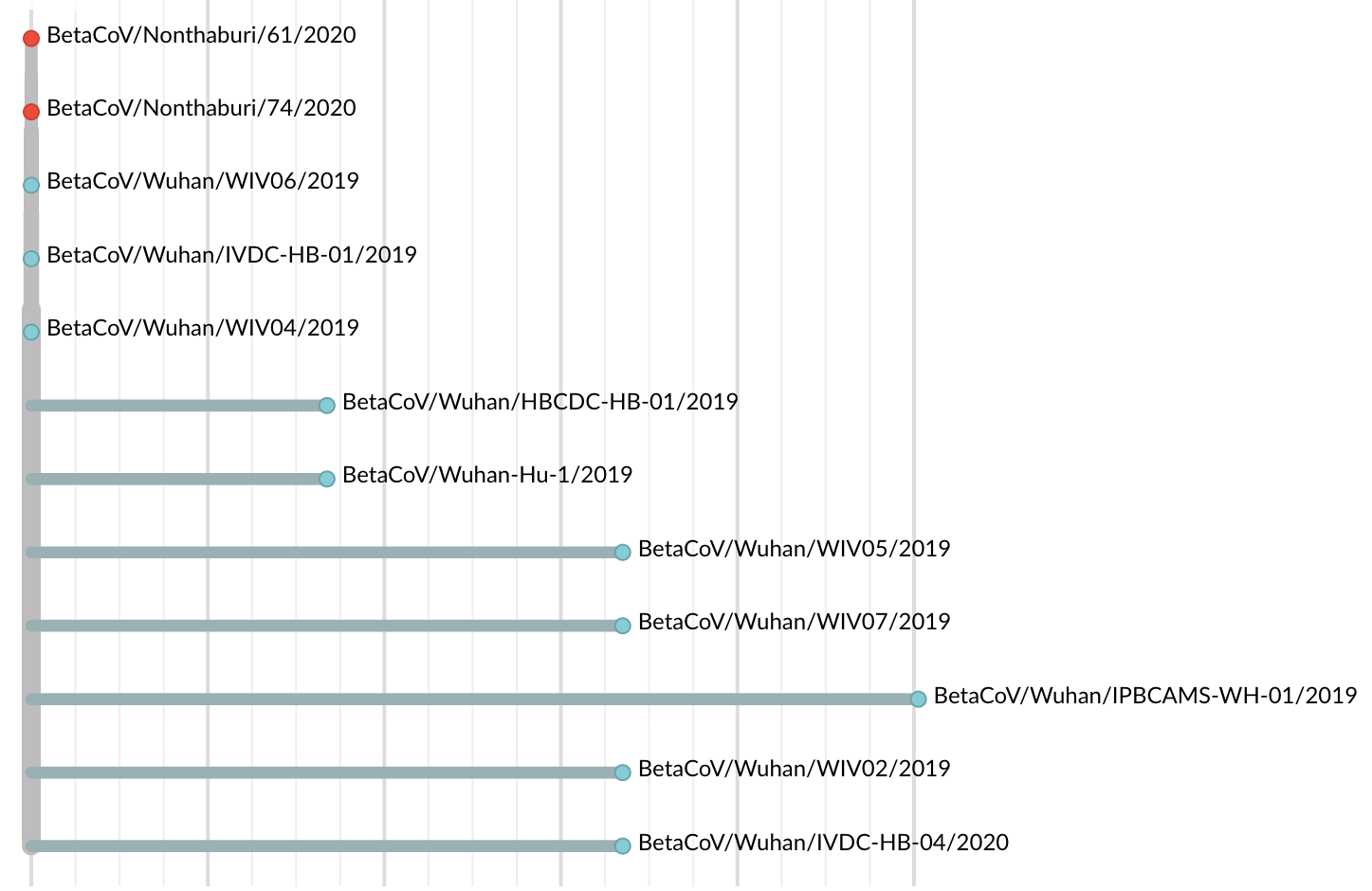
Jan 19: First 12 genomes from Wuhan (blue) and Bangkok (red) showed lack of genetic diversity
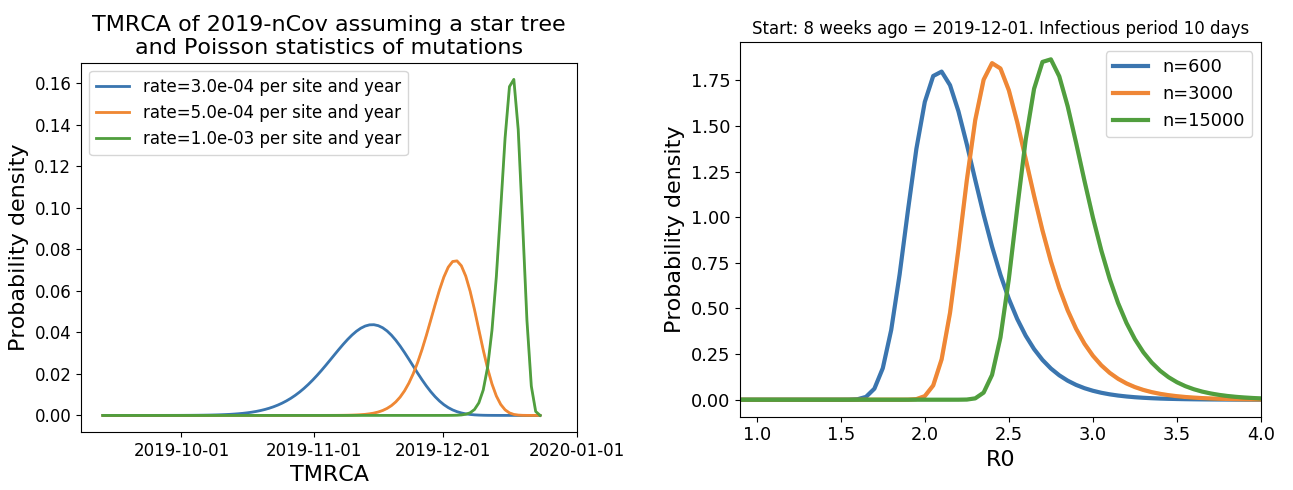
Jan 23: Introduction into the human population between Nov 15 and Dec 15 and subsequent rapid human-to-human spread
Rapid global epidemic spread from China
Epidemic in the USA was introduced from China in late Jan and from Europe during Feb
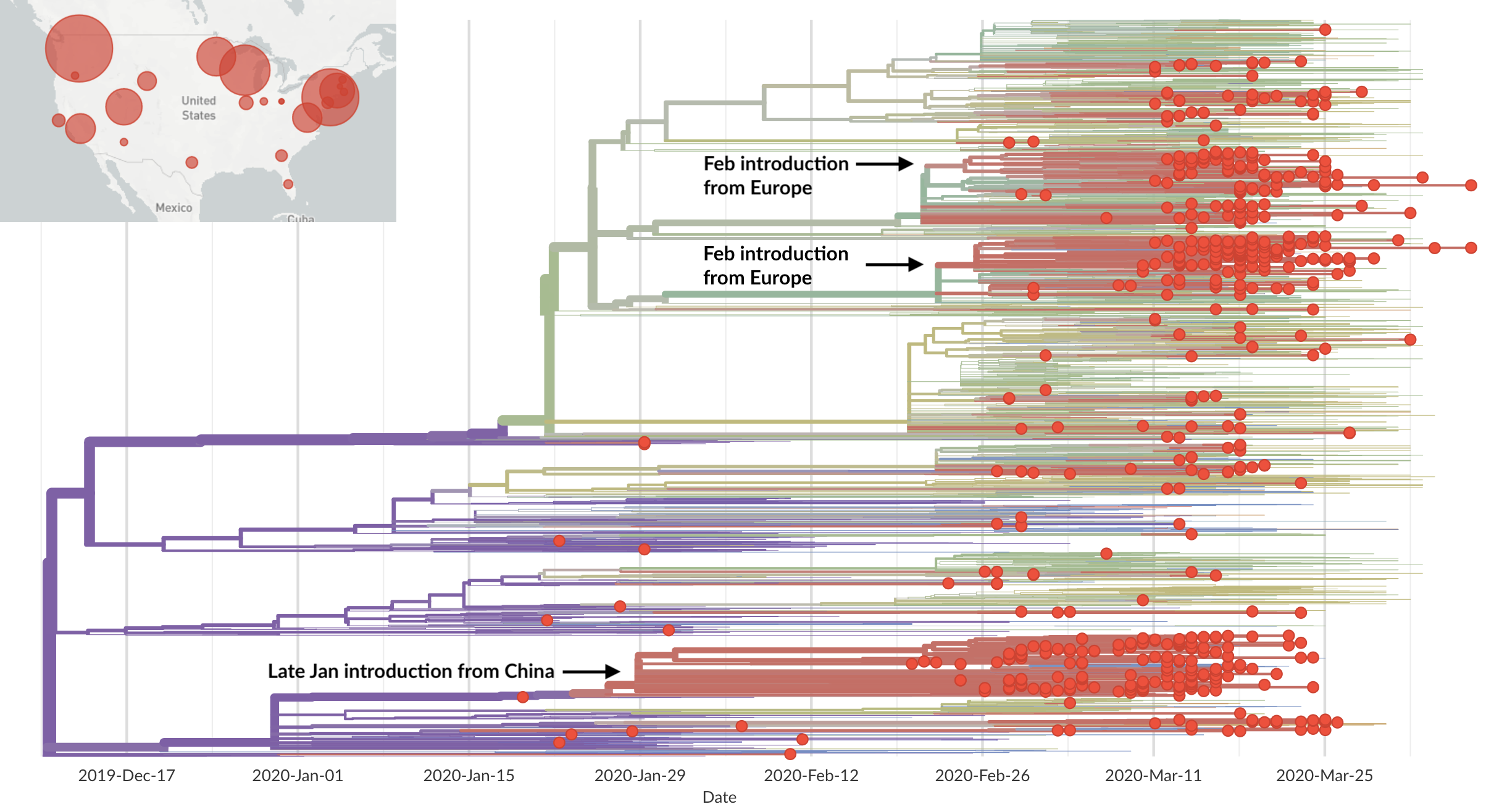
Early sequencing provided best estimate of extent of local outbreak
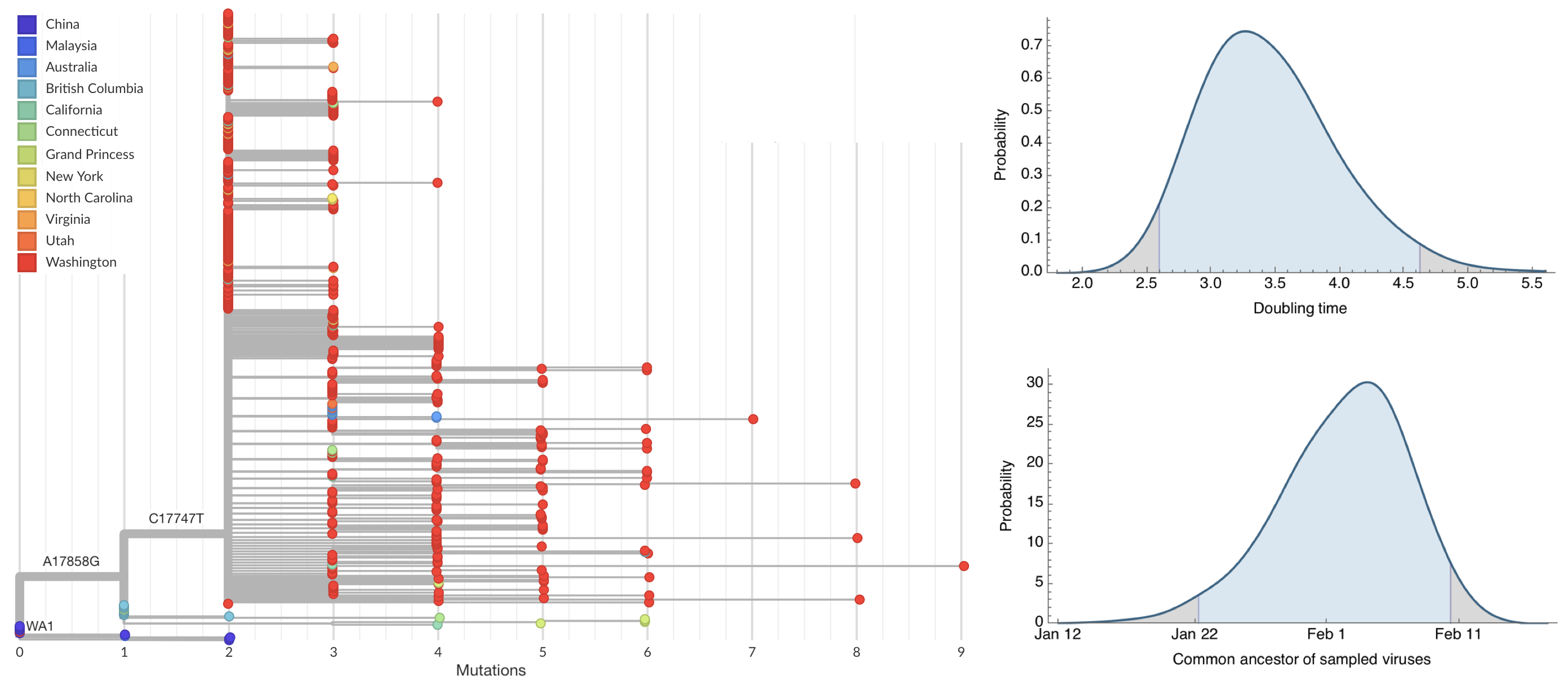
After initial wave, with mitigation
efforts and decreased travel,
regional clades emerge
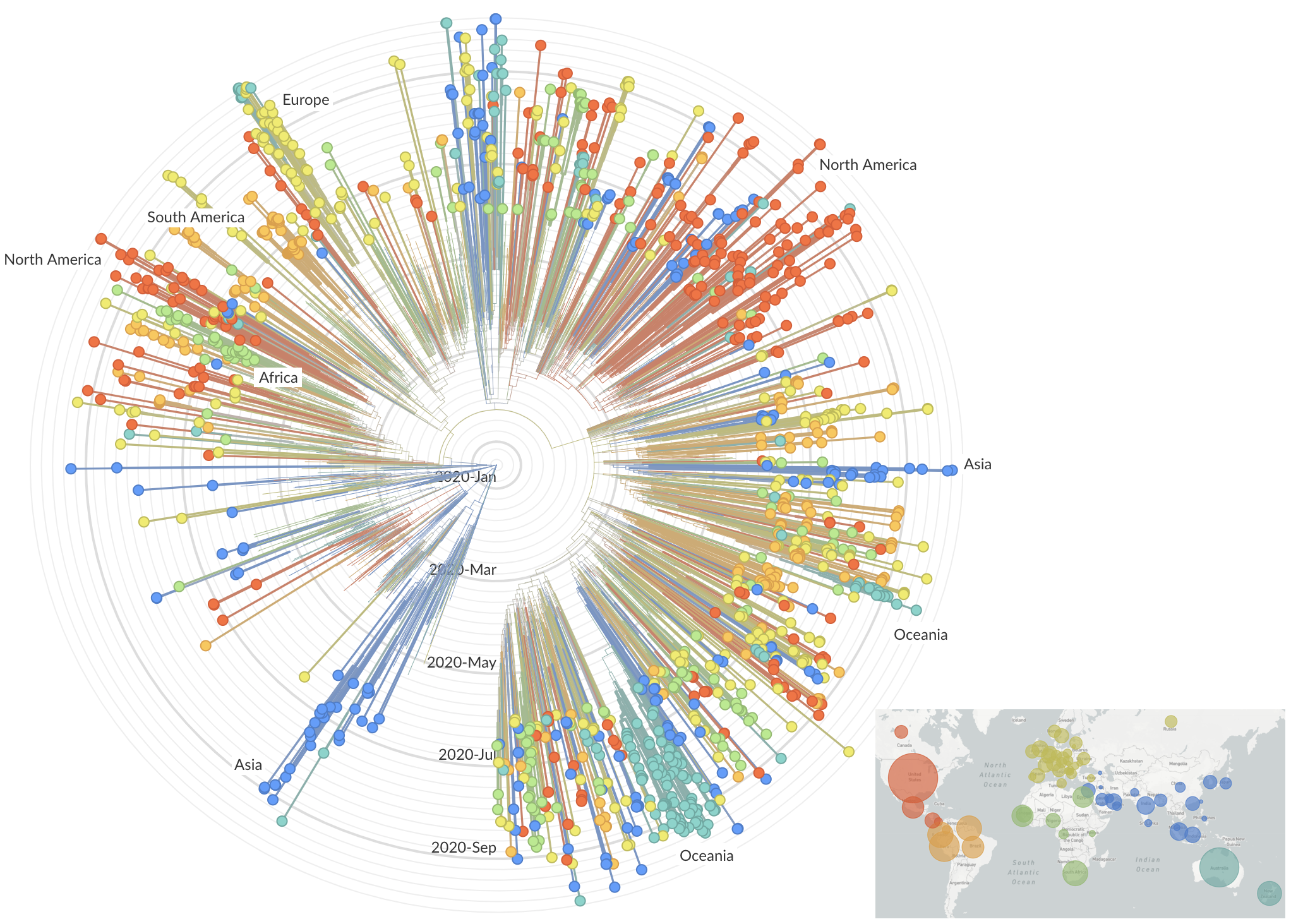
Emergence of Alpha in the UK with excess spike mutations
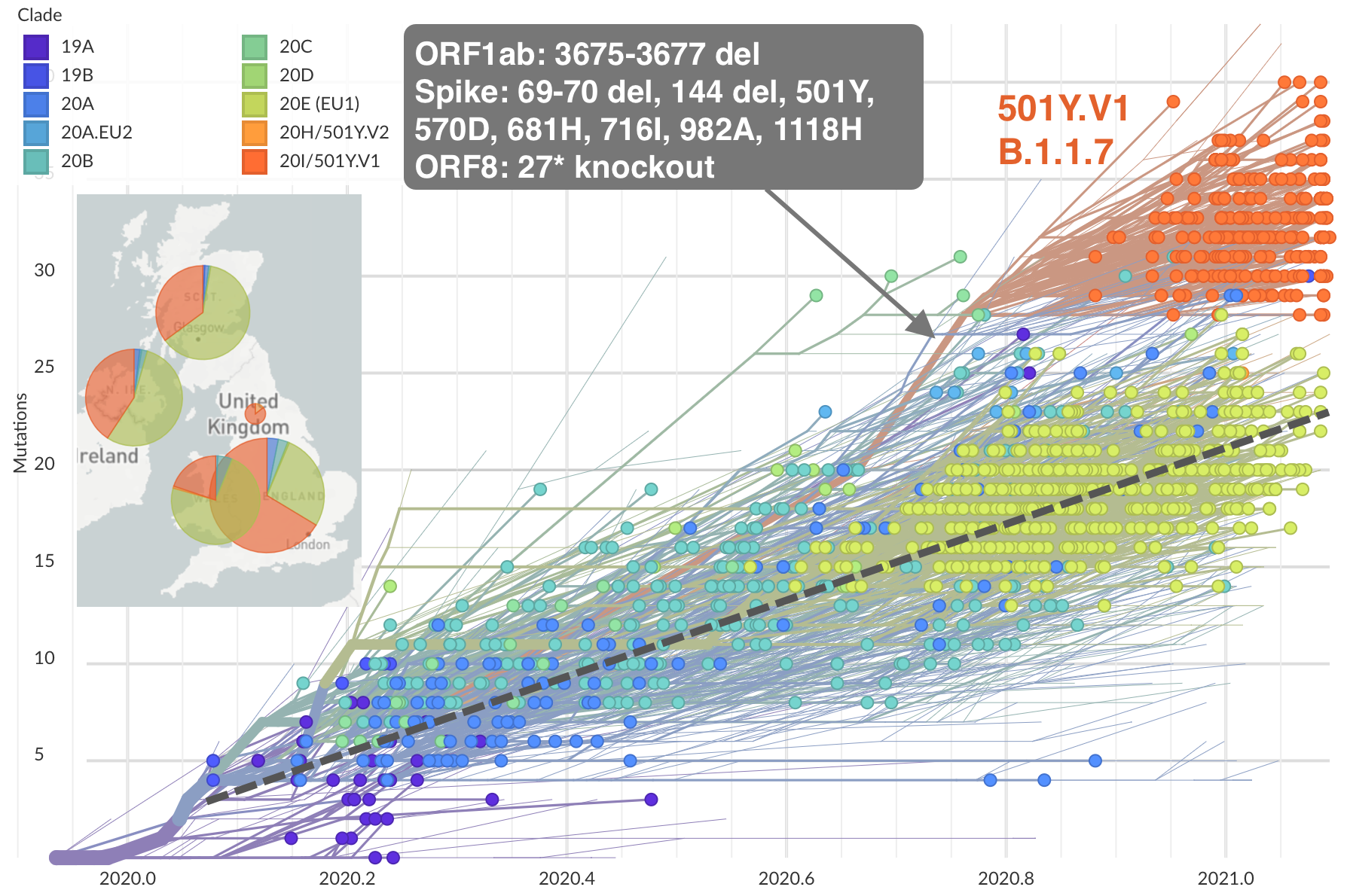
Further emergence of variants with increased transmissibility
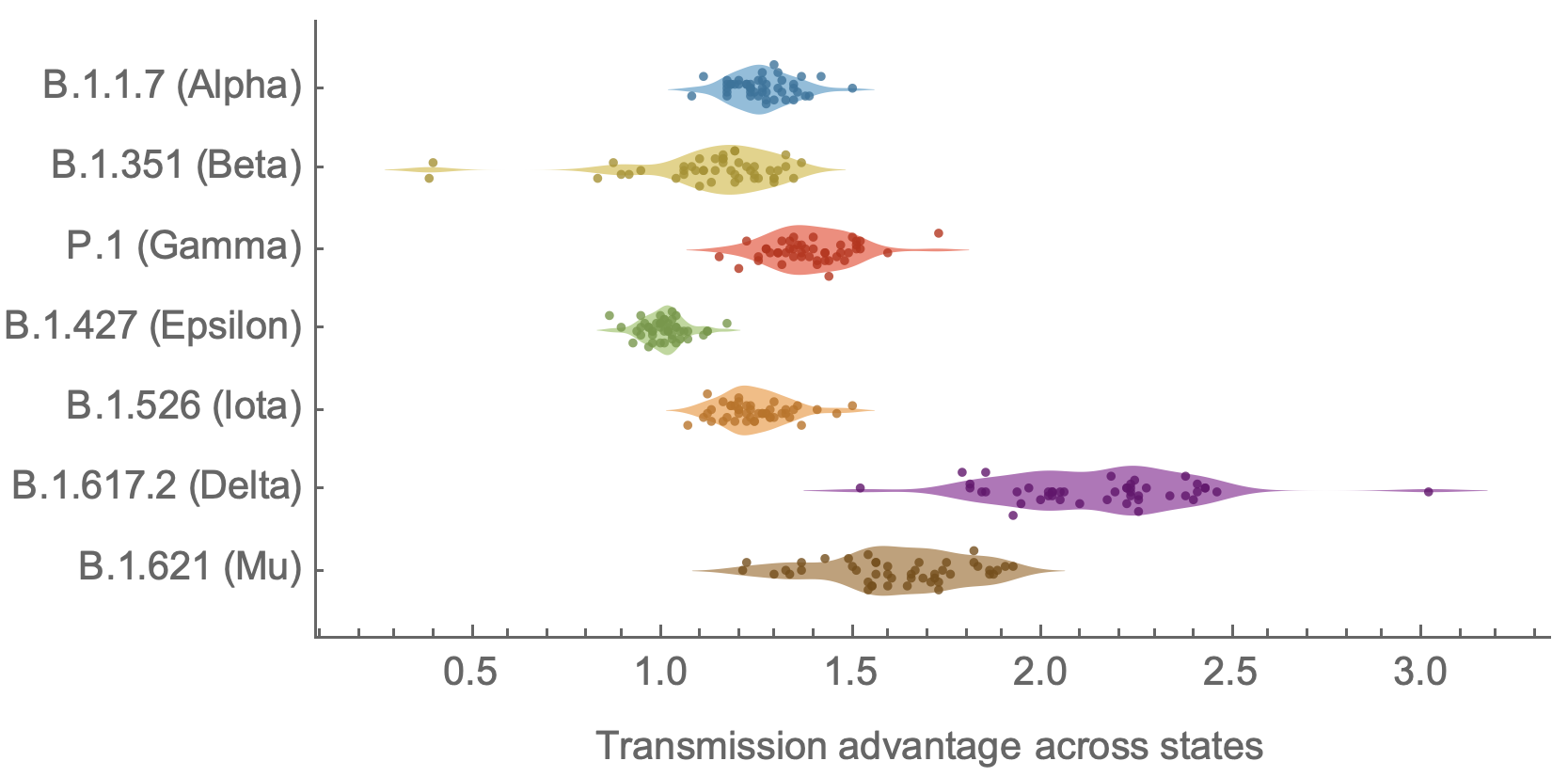
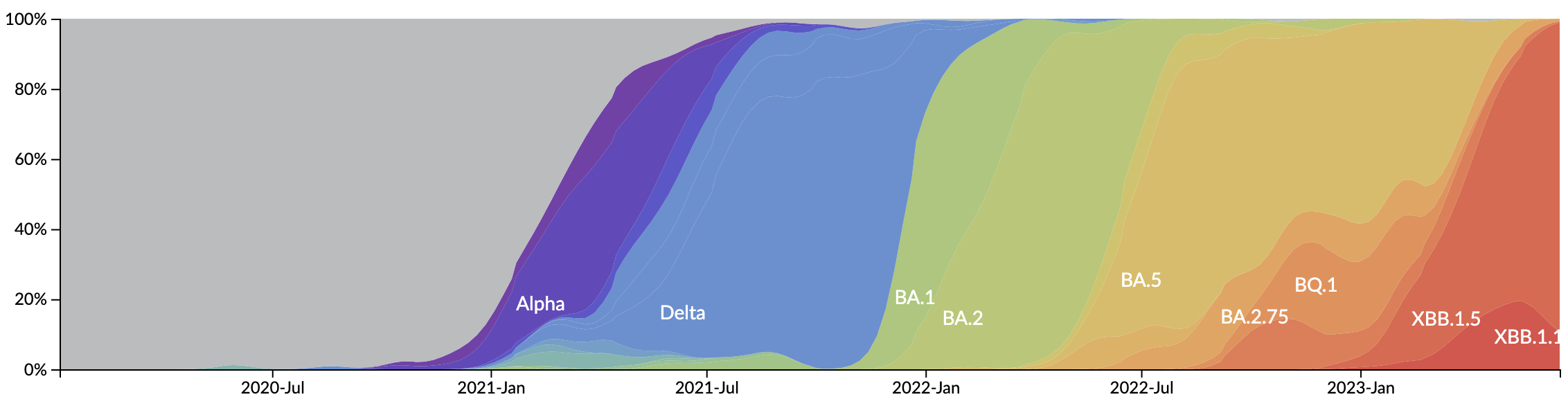
Variant emergence and spread has continued
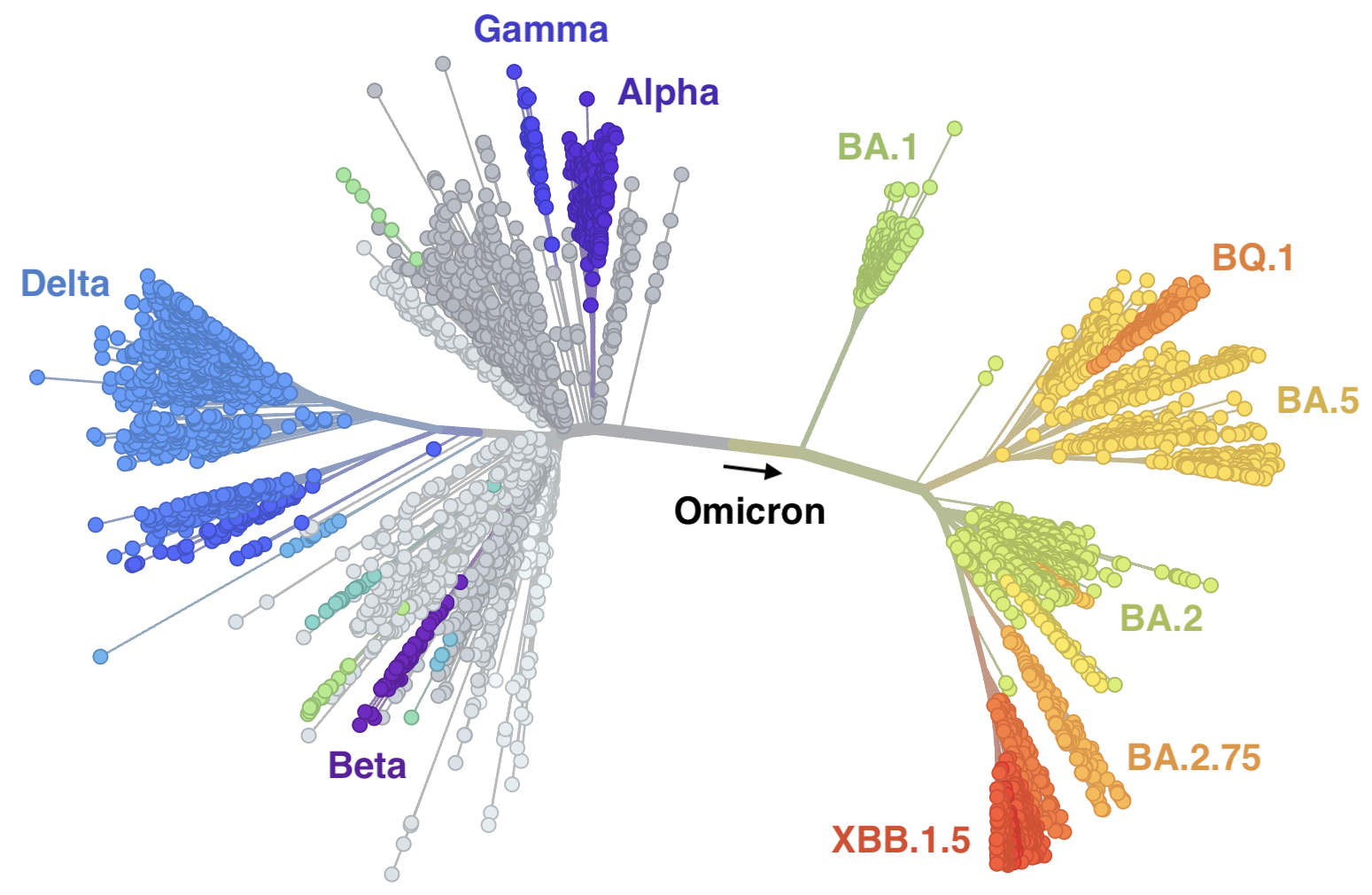
In which new variants emerge that escape from existing population immunity and spread rapidly
Future of genomic epidemiology
The COVID-19 pandemic has pushed the field perhaps ~5 years into the future
| 2013-16 Ebola in West Africa | 29k confirmed cases | 1610 genomes |
| 2015-17 Zika in the Americas | 223k confirmed cases | 942 genomes |
| 2018-19 seasonal flu in US | 290k confirmed cases | 8864 genomes |
| 2020-22 COVID-19 pandemic | 732M confirmed cases | 14.5M genomes |

Current research program and work with NW PGCoE
- Continue two major threads
- Evolutionary forecasting: focus on seasonal influenza and SARS-CoV-2 for impact on vaccine strain selection
- Genomic outbreak investigation: across pathogens with focus on spatial dynamics and recontructing spread
- Continue to build out the Nextstrain software platform for broader use by the community
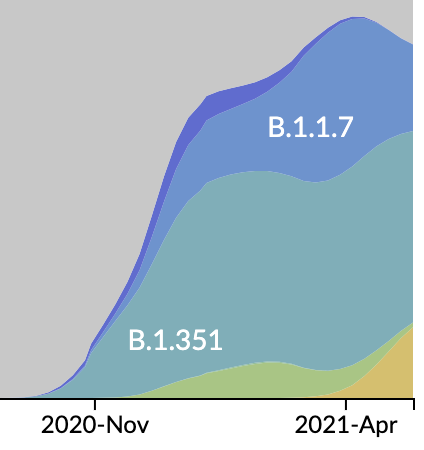
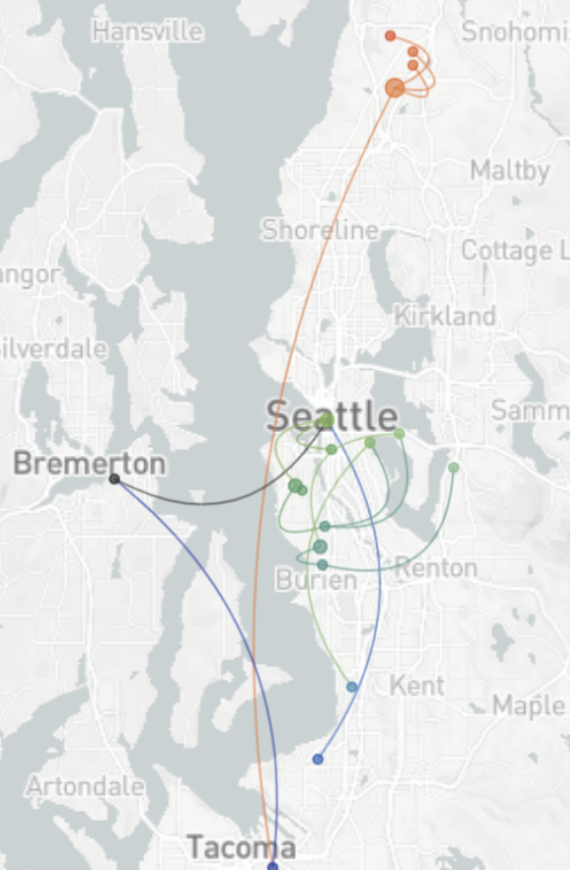
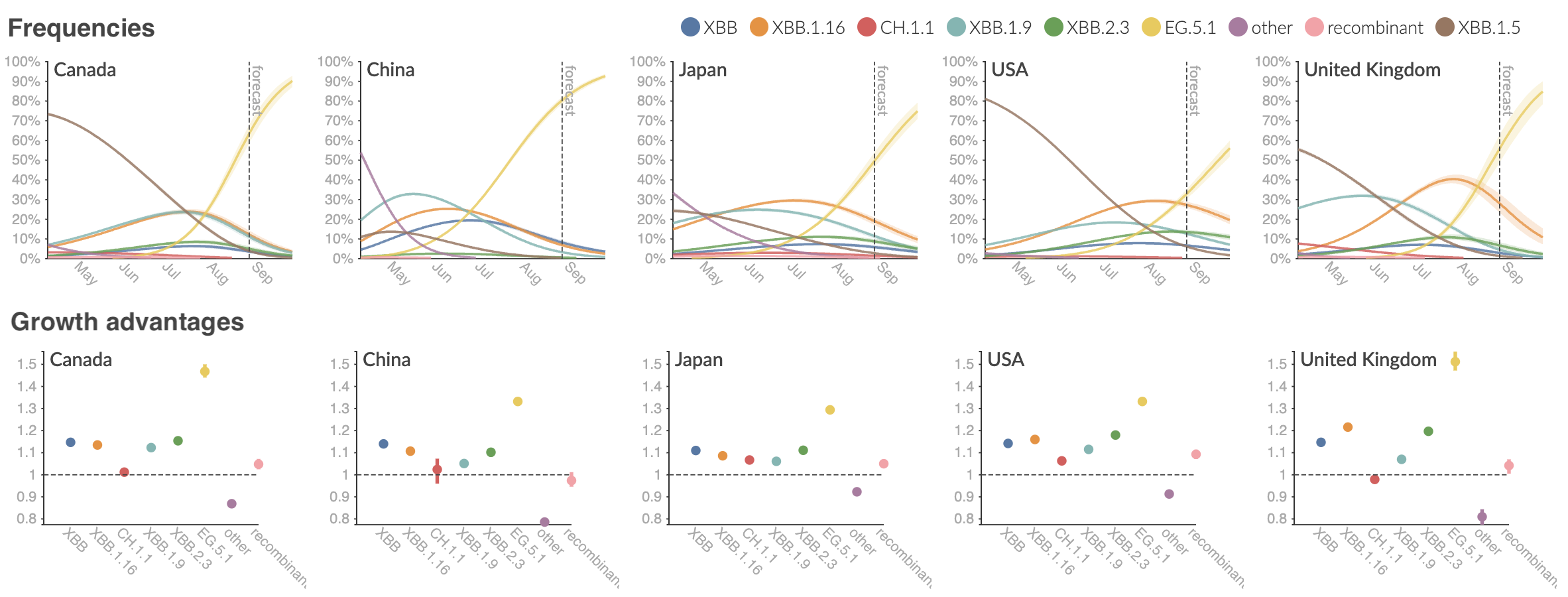
Clade and lineage forecasts continuously updated
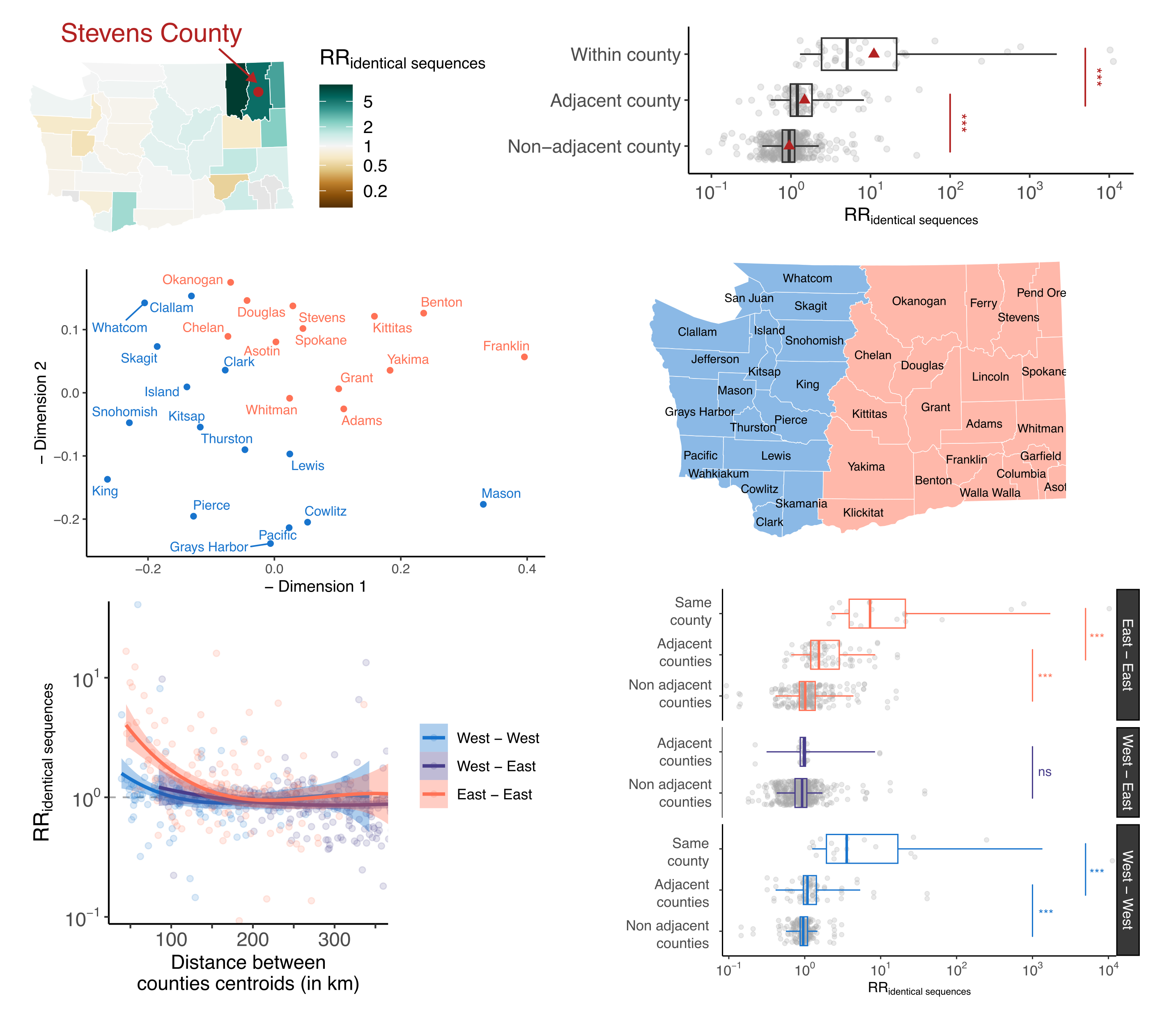
Detailed geographic transmission patterns from identical sequences
Lay ground-work for near ubiquitous pathogen sequencing
Acknowledgements
SARS-CoV-2 genomic epi: Data producers from all over the world, GISAID and the Nextstrain team
Bedford Lab:
![]() John Huddleston,
John Huddleston,
![]() James Hadfield,
James Hadfield,
![]() Katie Kistler,
Katie Kistler,
![]() Thomas Sibley,
Thomas Sibley,
![]() Jover Lee,
Jover Lee,
![]() Cassia Wagner,
Cassia Wagner,
![]() Miguel Paredes,
Miguel Paredes,
![]() Nicola Müller,
Nicola Müller,
![]() Marlin Figgins,
Marlin Figgins,
![]() Victor Lin,
Victor Lin,
![]() Jennifer Chang,
Jennifer Chang,
![]() Allison Li,
Allison Li,
![]() Eslam Abousamra,
Eslam Abousamra,
![]() Donna Modrell,
Donna Modrell,
![]() Nashwa Ahmed,
Nashwa Ahmed,
![]() Cécile Tran Kiem
Cécile Tran Kiem







