Real-time tracking of virus evolution
Trevor Bedford (@trvrb)
23 May 2018
AMD Seminar Series
CDC
We work at the interface of virology, evolution and epidemiology
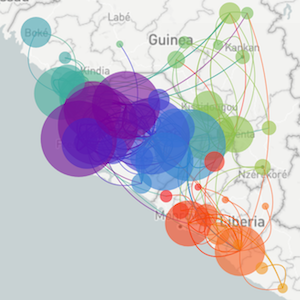

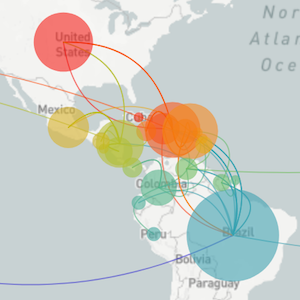
Sequencing to reconstruct pathogen evolution and spread
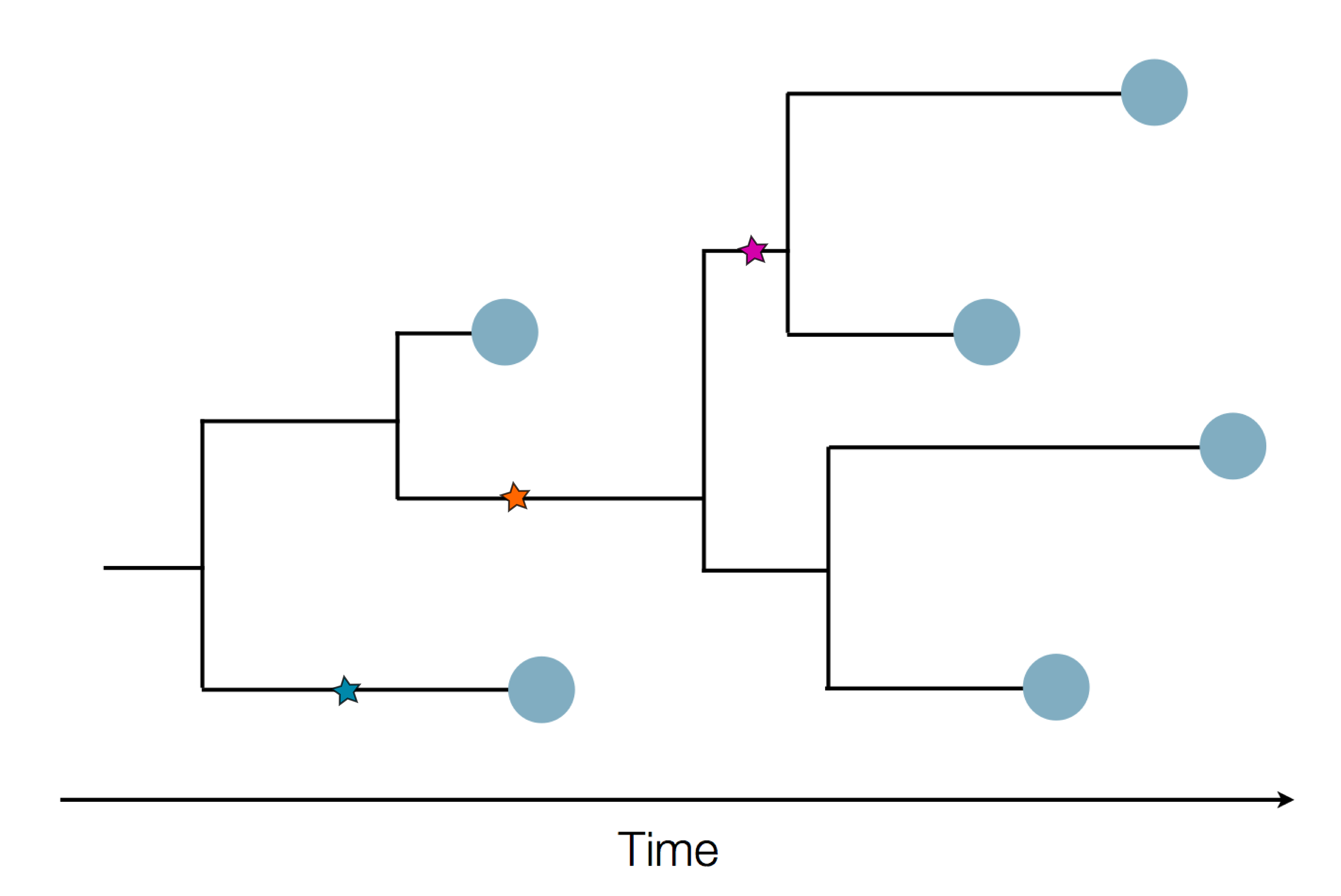
Epidemic process
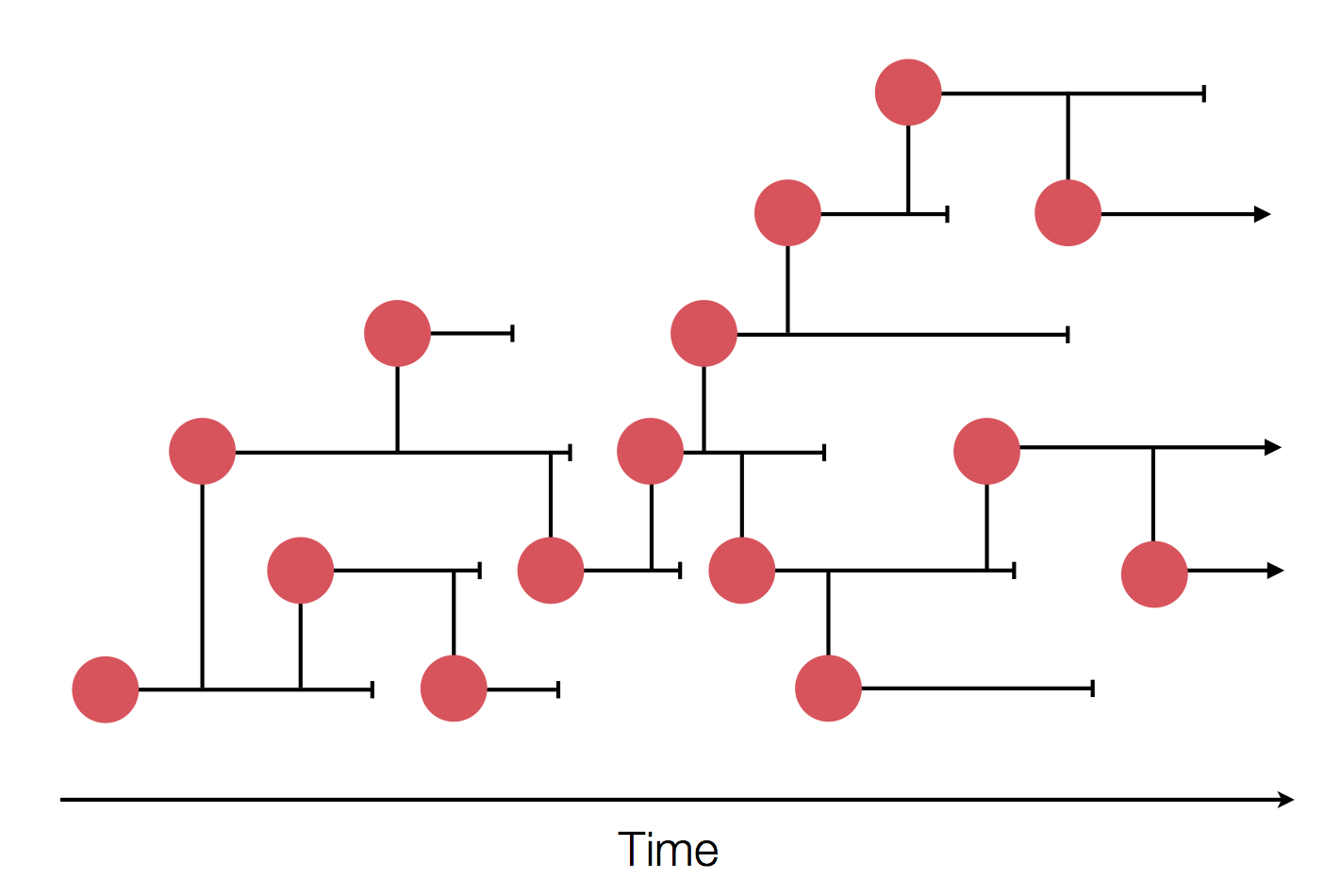
Sample some individuals
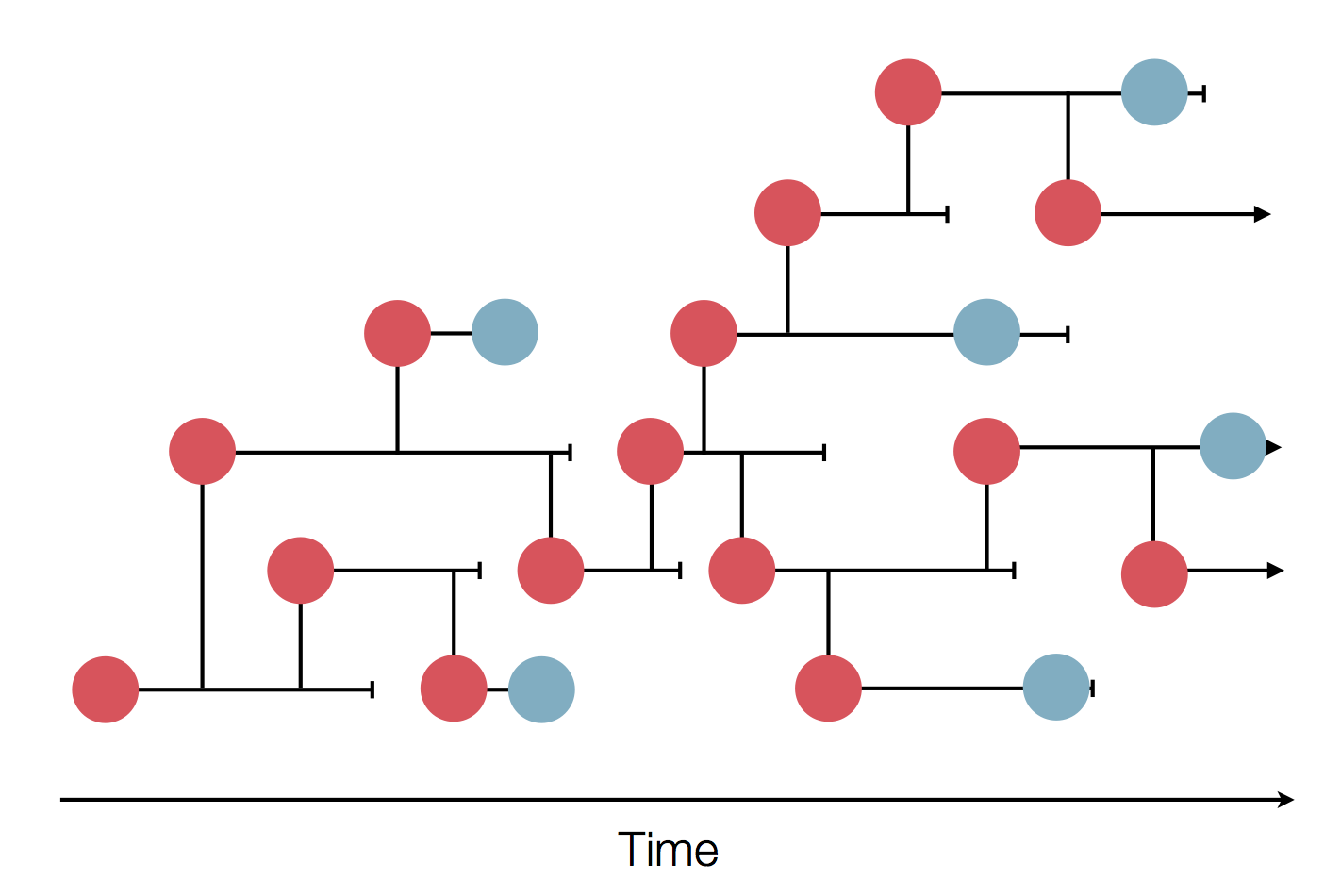
Sequence and determine phylogeny
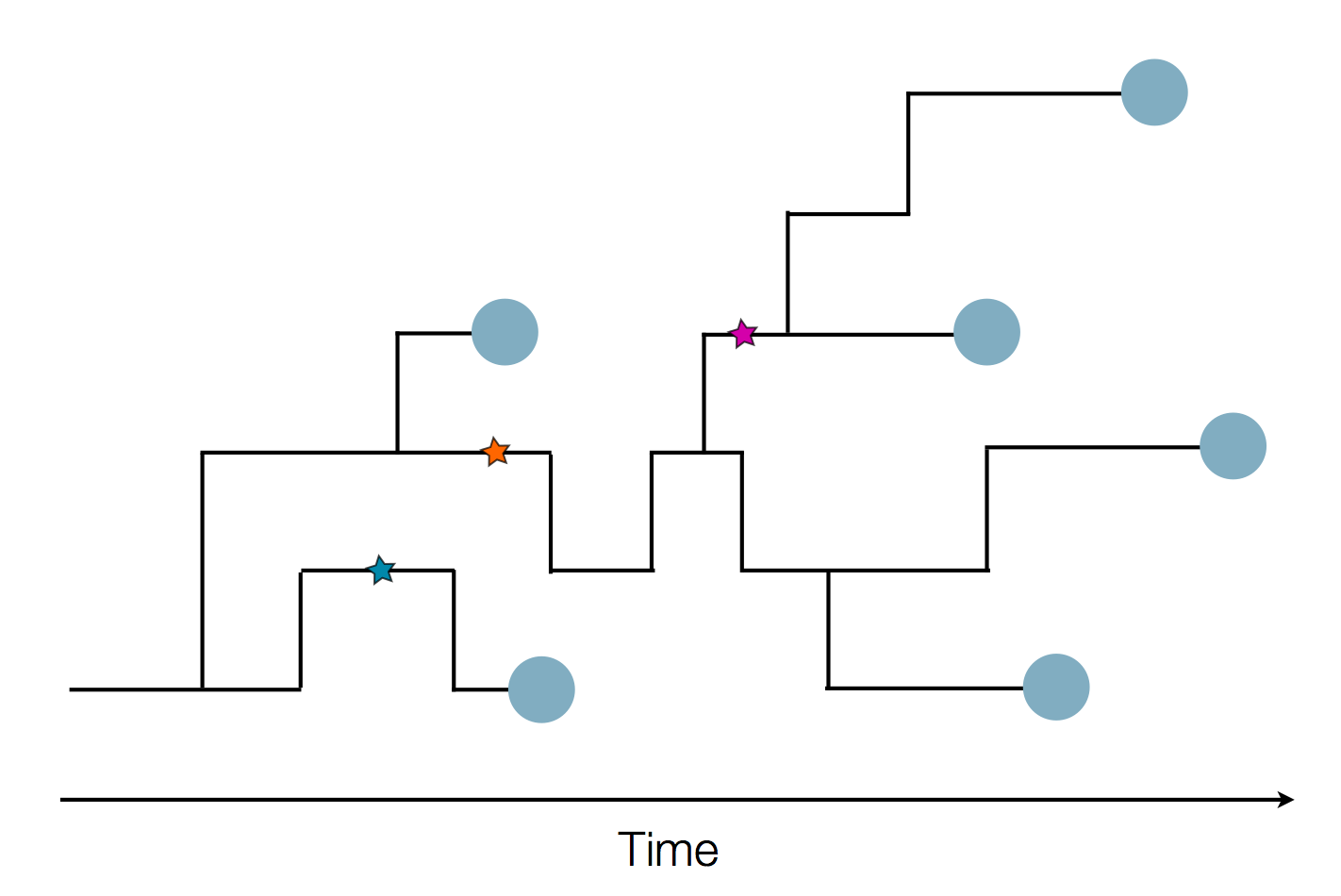
Sequence and determine phylogeny
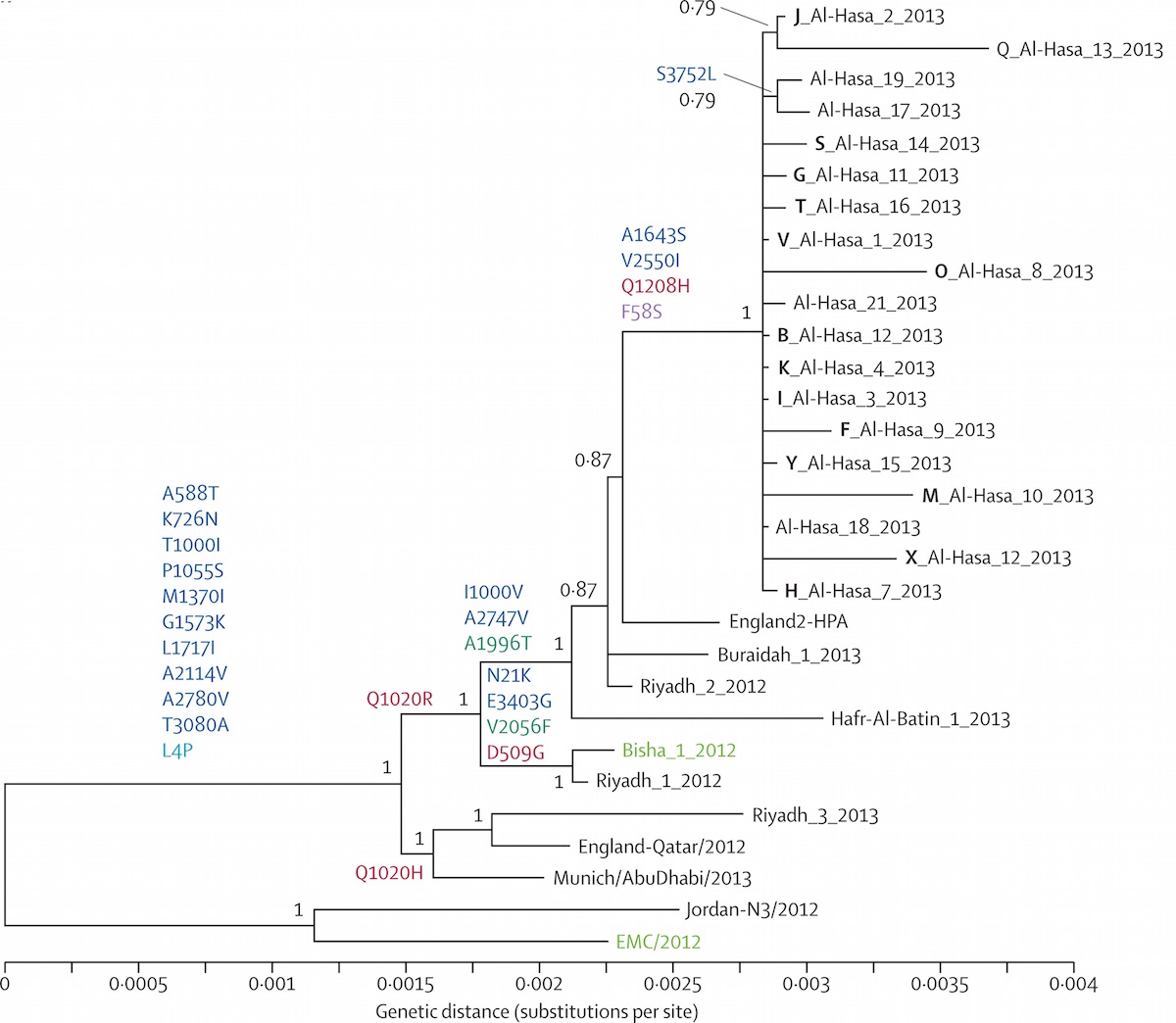
Localized Middle Eastern MERS-CoV phylogeny
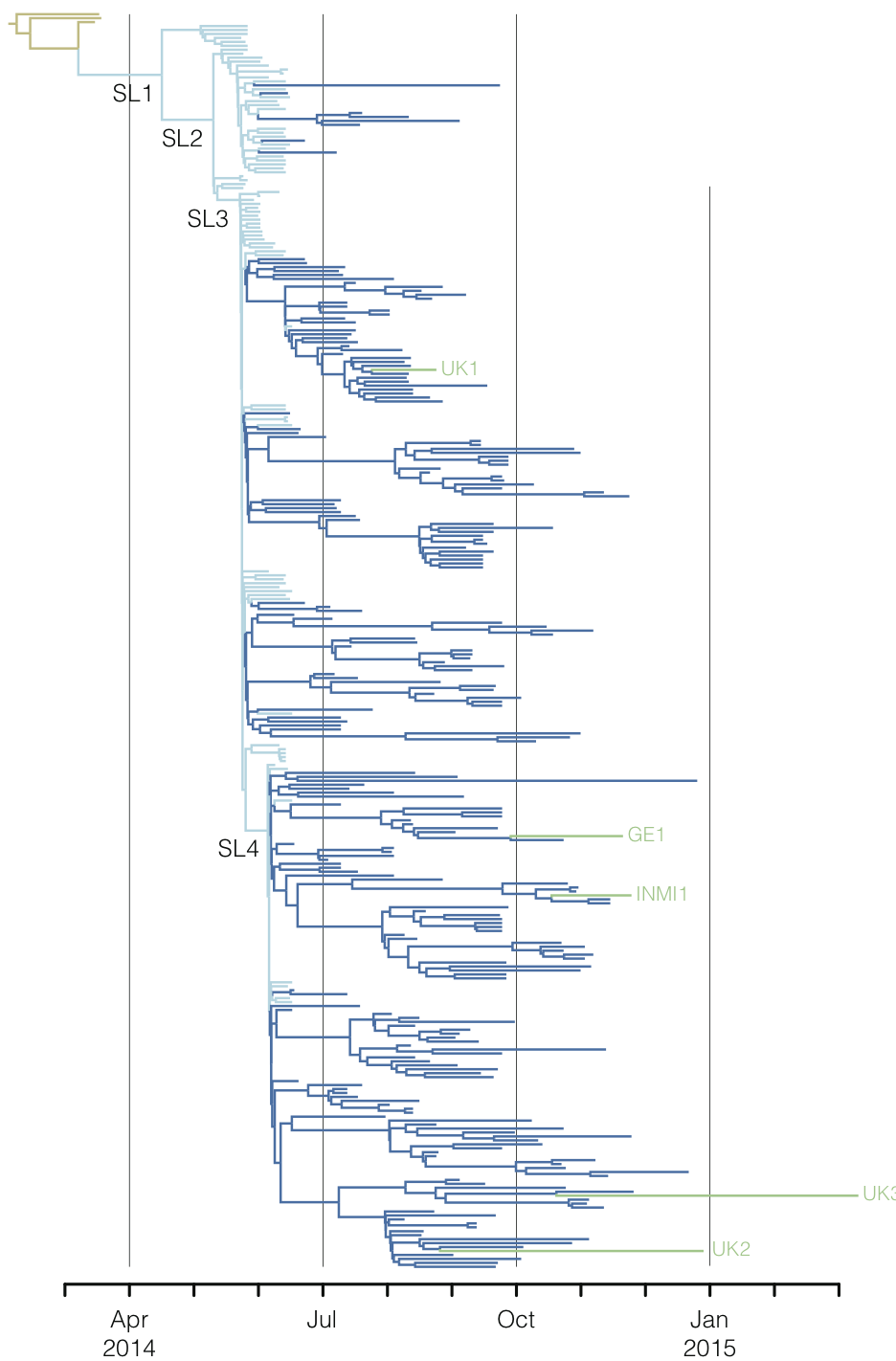
Regional West African Ebola phylogeny
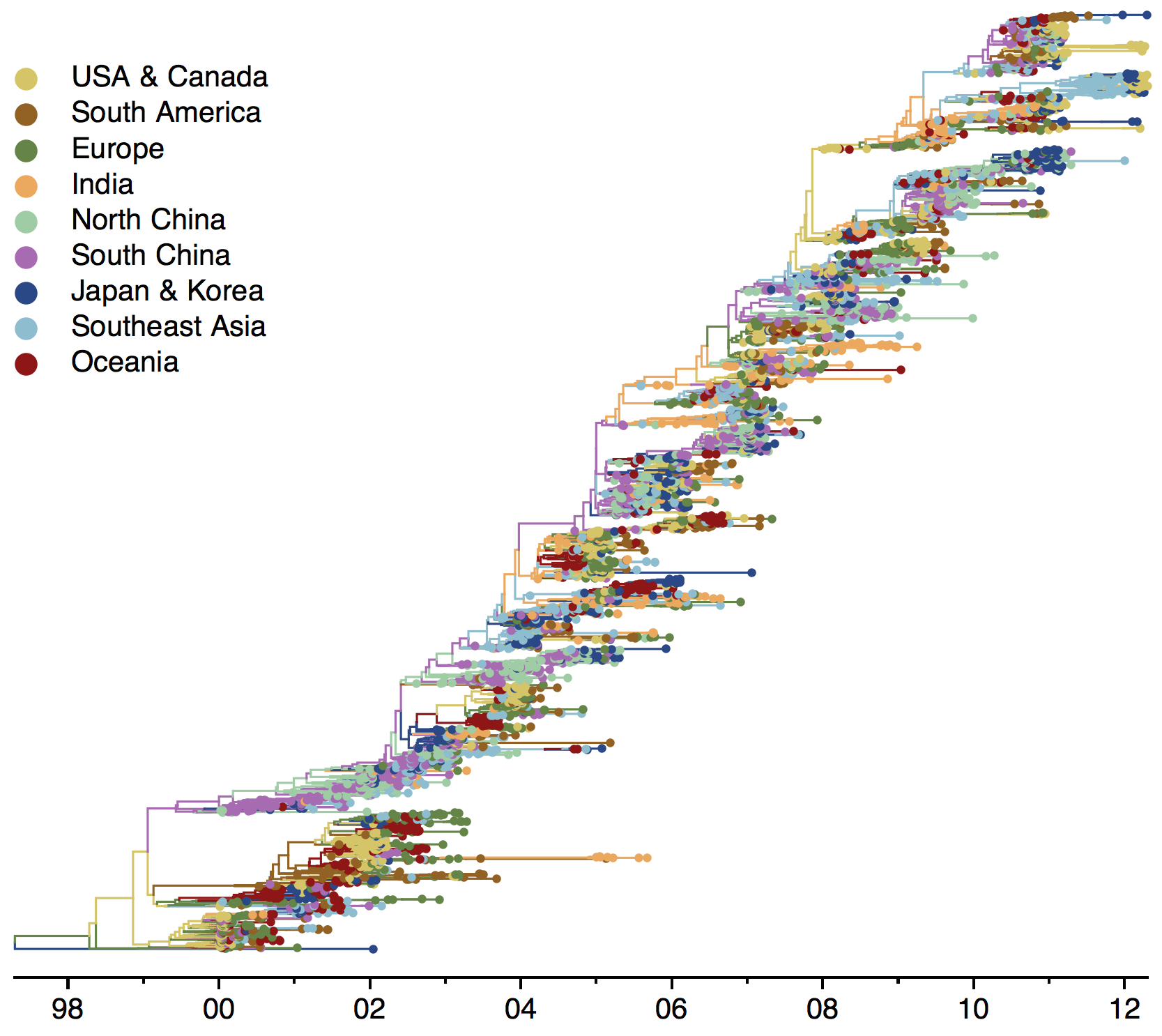
Global influenza phylogeny
Phylogenetic tracking has the capacity to revolutionize epidemiology
Outline
- Influenza circulation and antigenic drift
- Ebola spread in West Africa
- Zika spread in the Americas
- "Real-time" analyses
Influenza
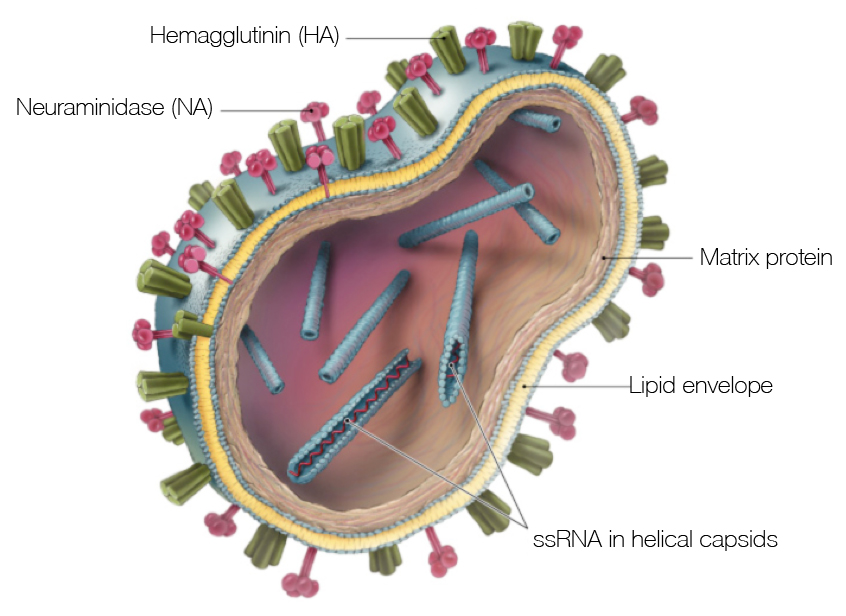
Influenza virion
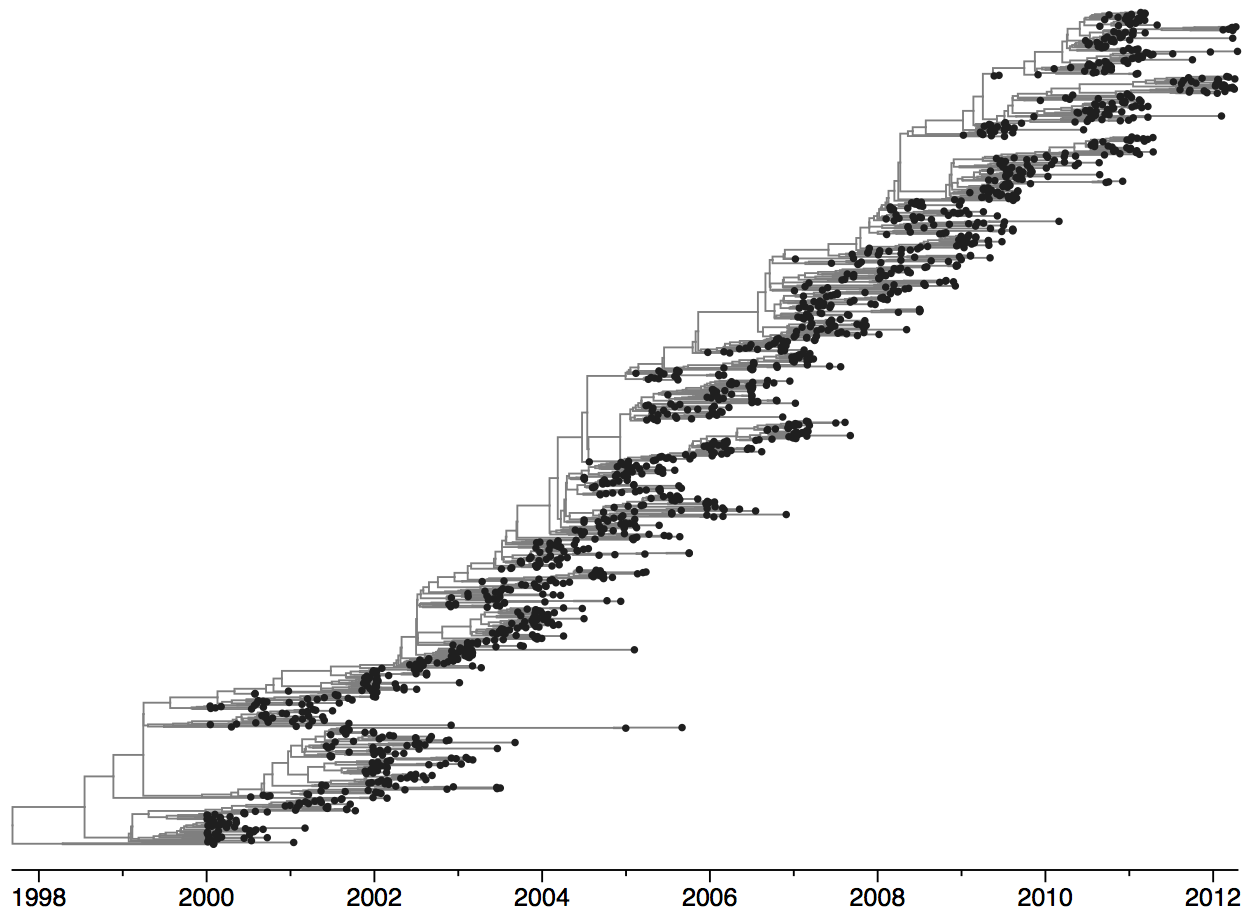
Population turnover is extremely rapid
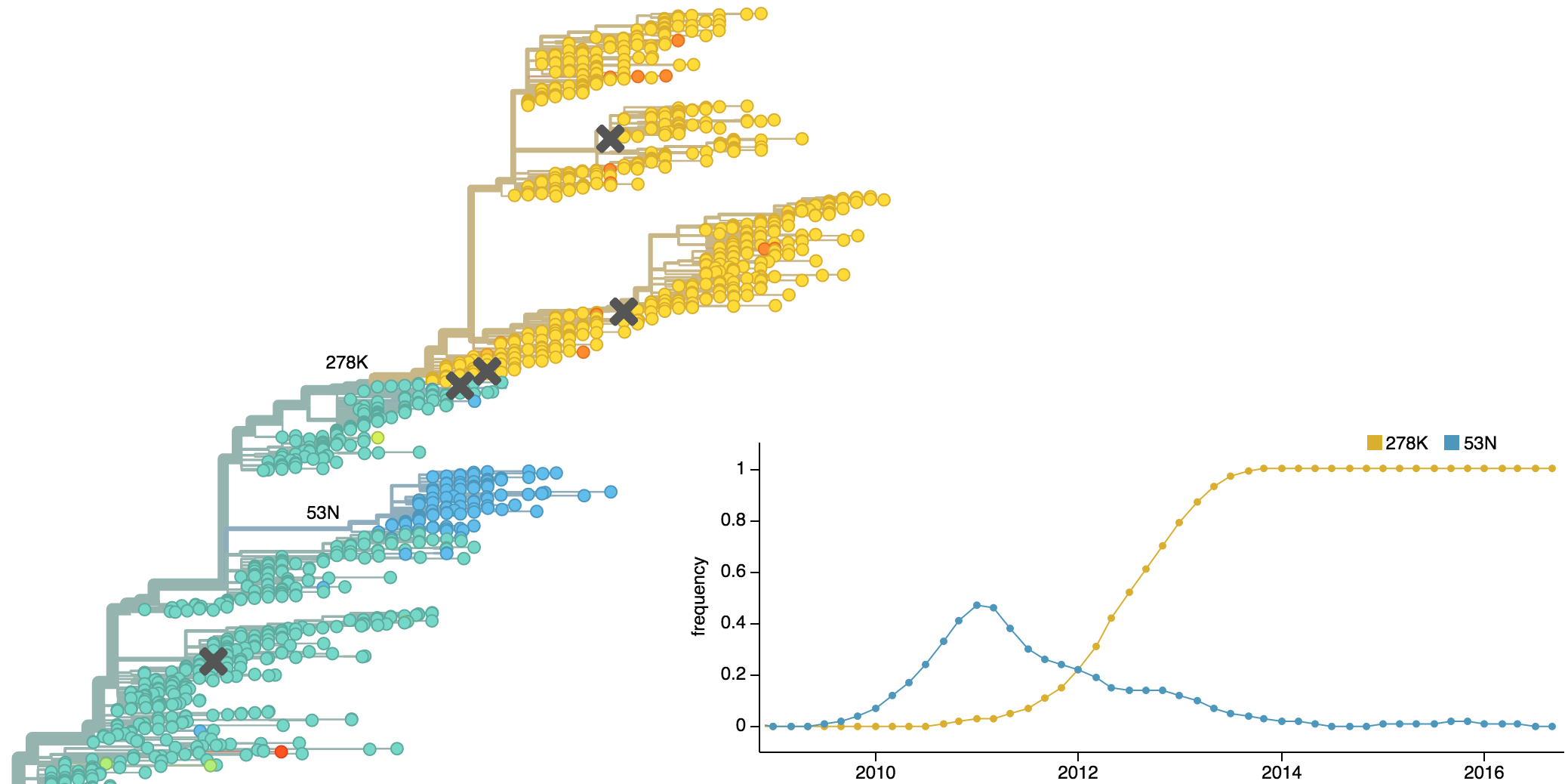
Clades emerge, die out and take over
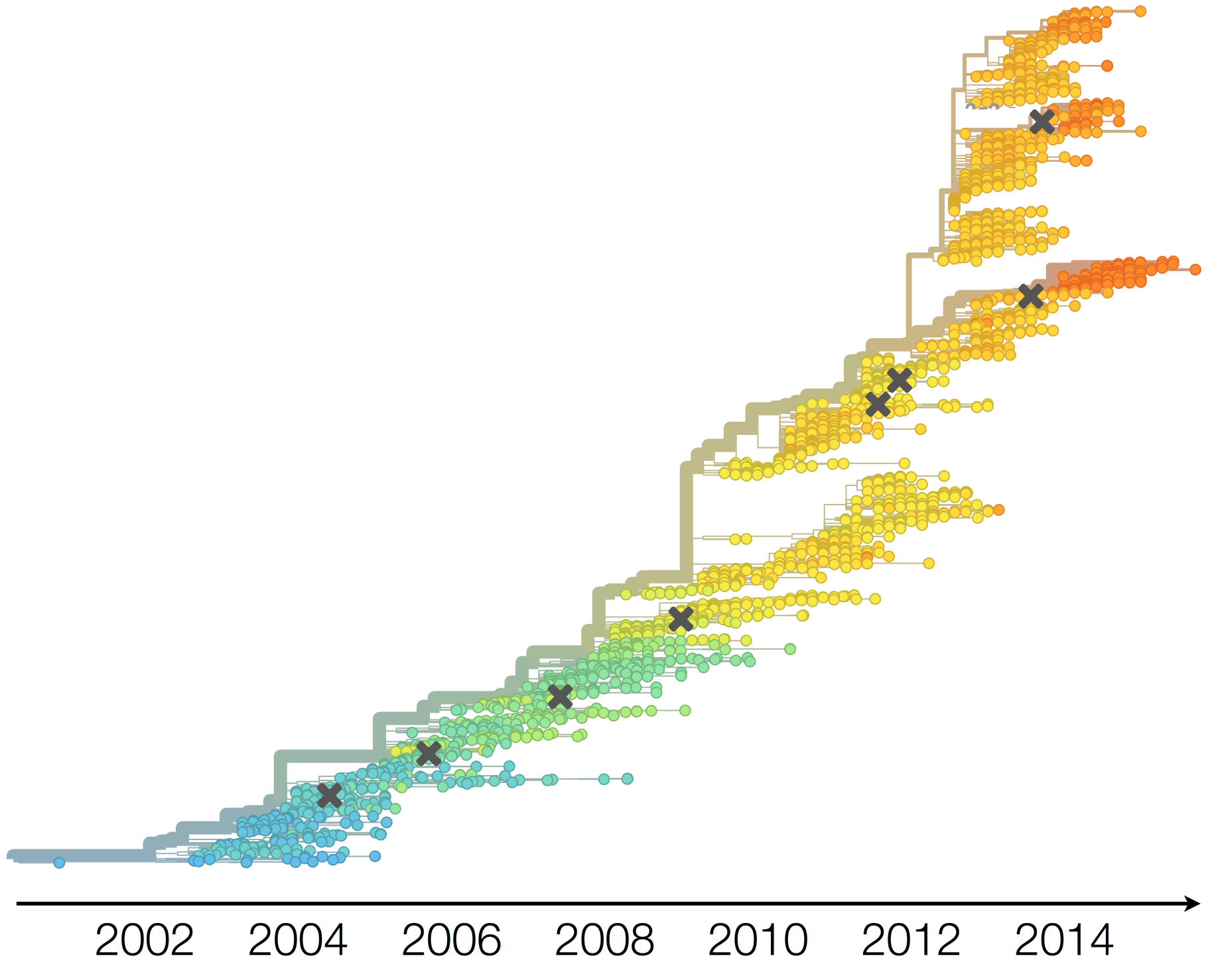
Clades show rapid turnover

Dynamics driven by antigenic drift
Drift necessitates vaccine updates
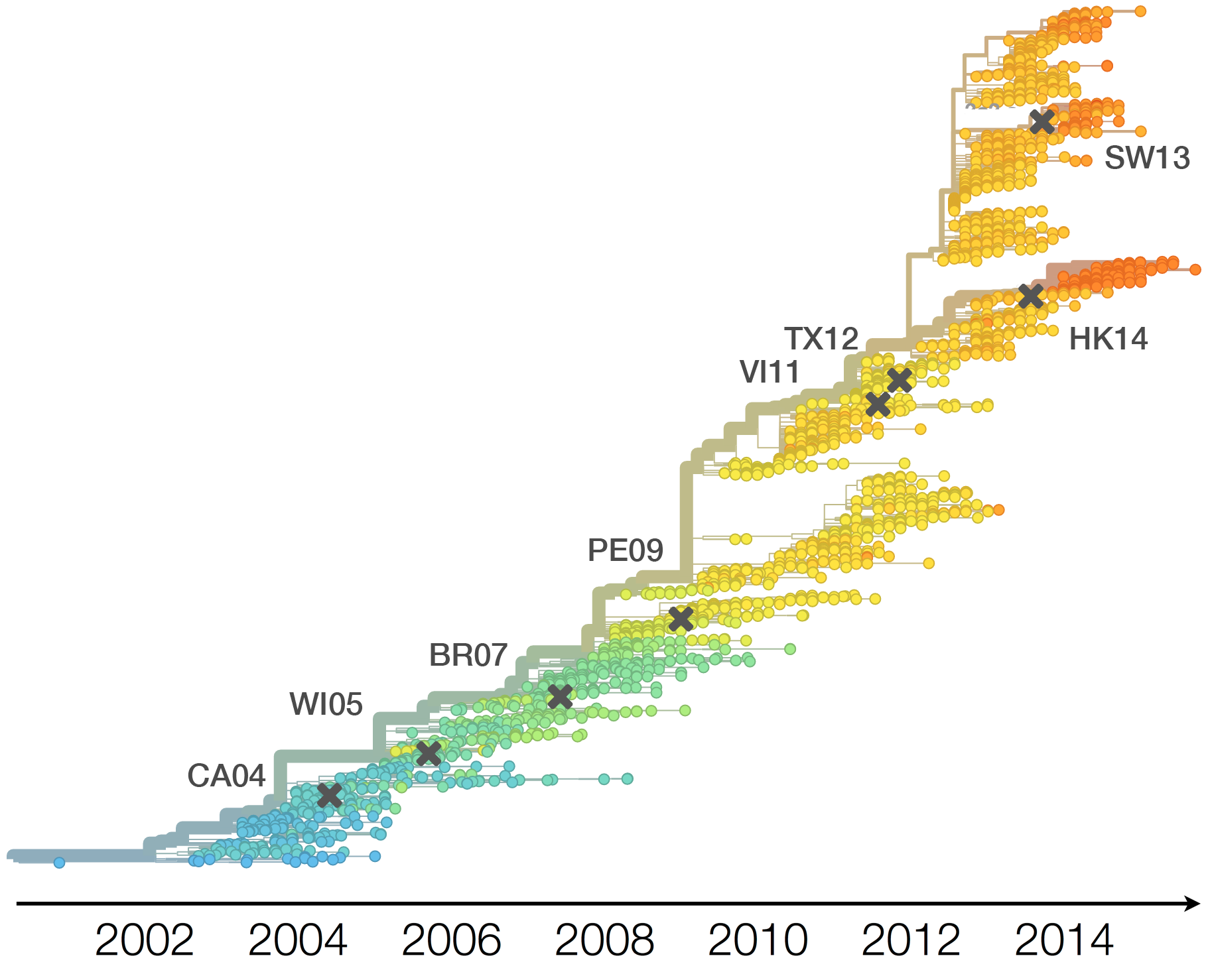
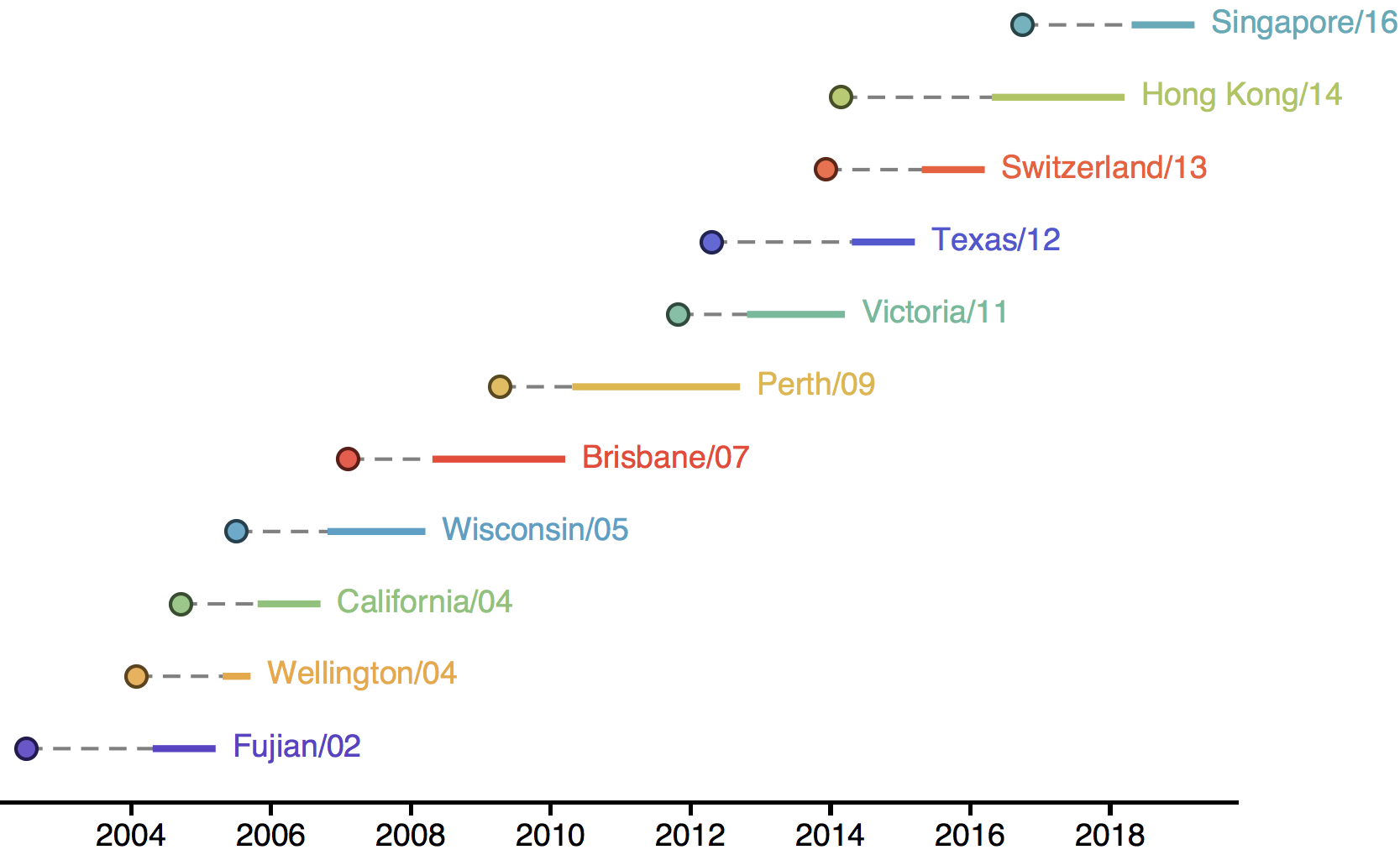
H3N2 vaccine updates occur every ~2 years
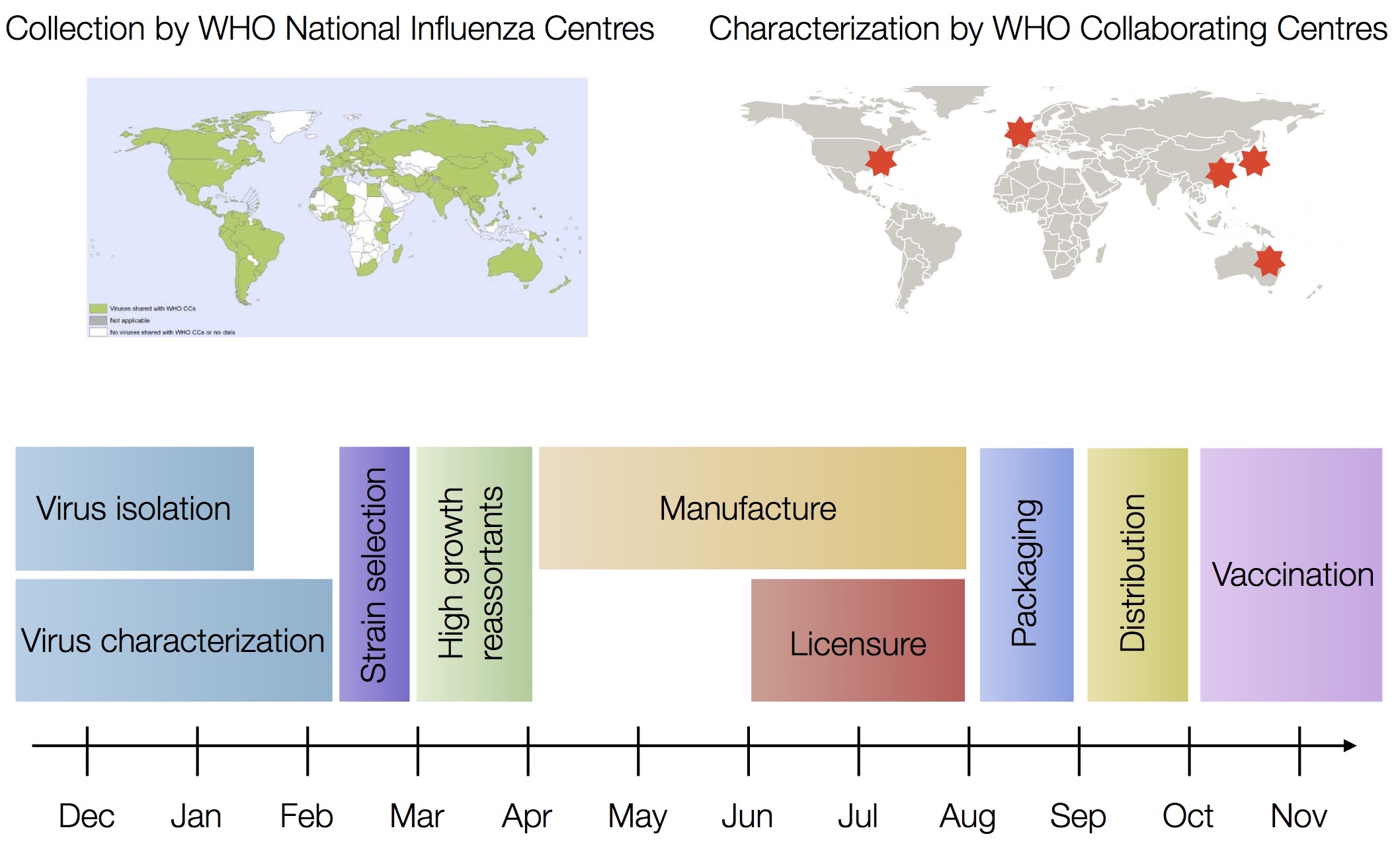
Vaccine strain selection by WHO
Problem of applied evolutionary biology
Every paper in the field...
- "These observations have implications for influenza surveillance and vaccine formulation" (Wolf et al 2006)
- "Our results have implications for the design of vaccines to combat rapidly mutating viral diseases" (Gupta et al 2006)
- "These results may have important implications for influenza vaccine and antiviral research" (Bhatt et al 2011)
- "Needless to say, these results have important implications for the updating of vaccines against influenza" (Zinder et al 2013)
Disconnect between evolutionary studies and information needed by WHO
- WHO needs specific advice, ie this strain is likely to take off, this strain is likely to die out
- Problems of generality and timeliness
2014 workshop on microbial evolution at the Kavli Institute for Theoretical Physics
Decided to tackle this head on and build something that
- Charts behavior of specific strains
- Can be kept continually up to date
Nextflu
Project to provide a real-time view of the evolving influenza population
Nextflu
Project to provide a real-time view of the evolving influenza population
All in collaboration with Richard Neher

Nextflu pipeline
- Download all recent HA sequences from GISAID
- Filter to remove outliers
- Subsample across time and space
- Align sequences
- Build tree
- Estimate clade frequencies
- Infer antigenic phenotypes
- Export for visualization
Up-to-date analysis publicly available at:
Nextflu.org
Current H3N2 diversity
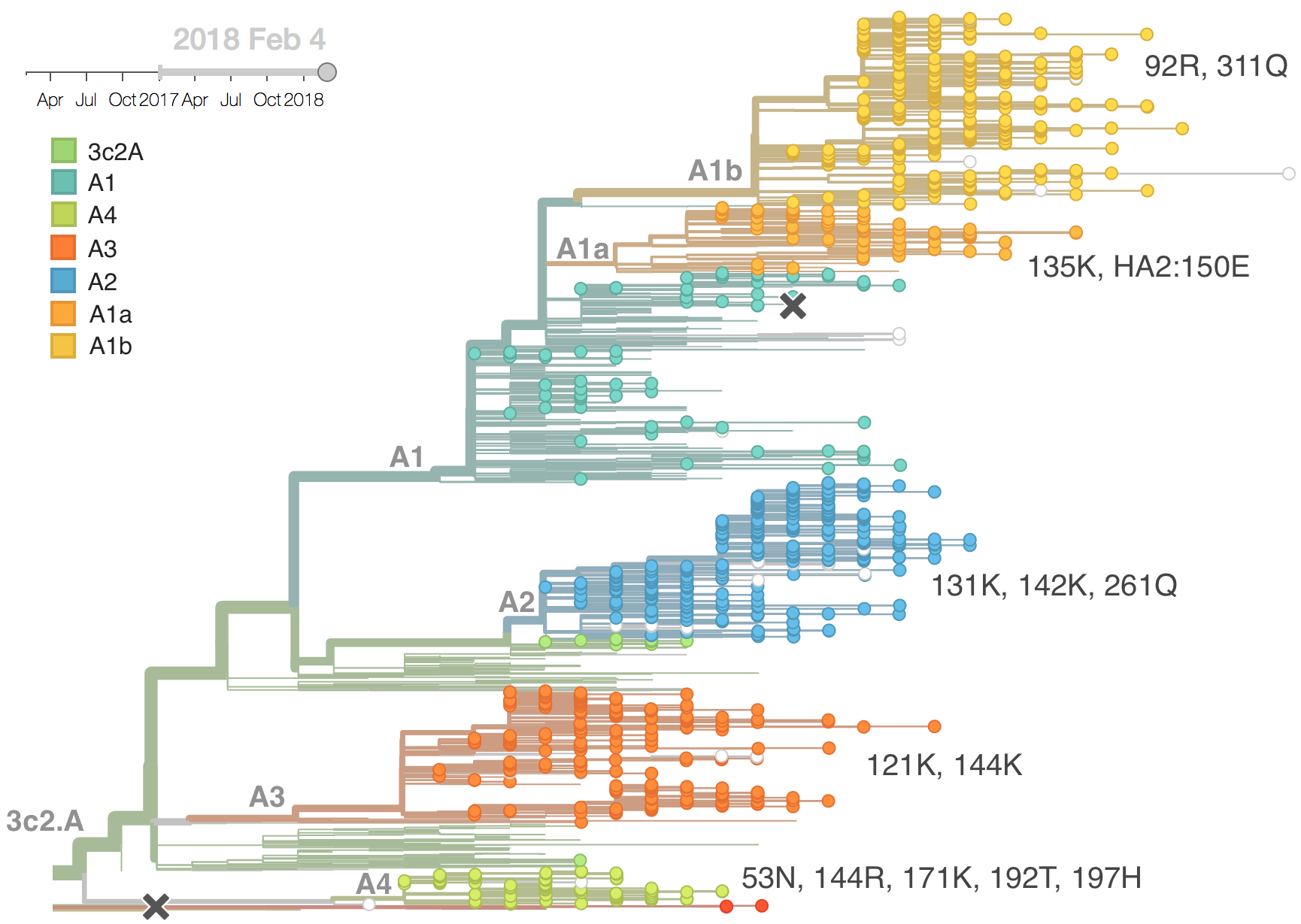
Current H3N2 diversity
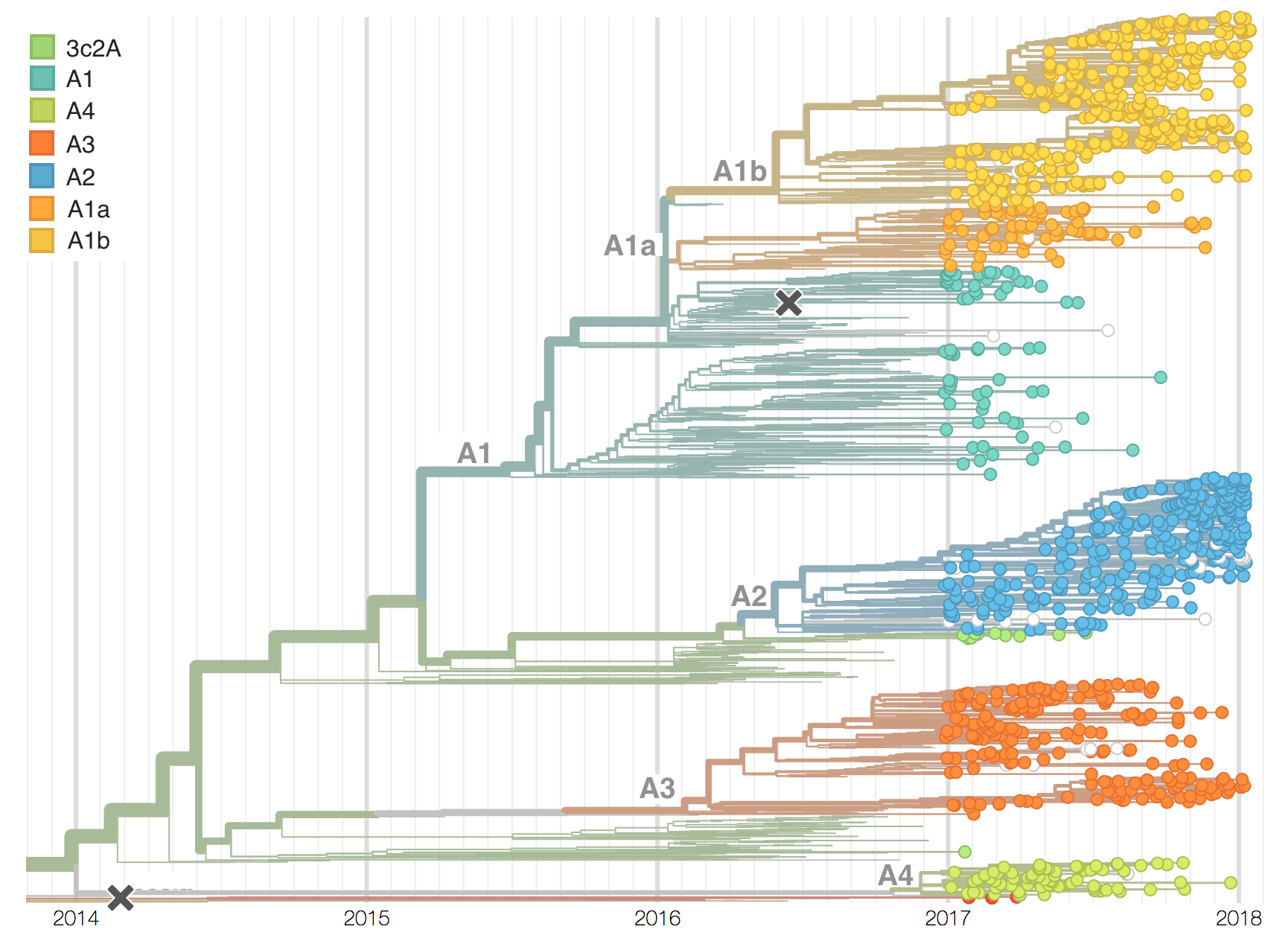
Two clades have been growing rapidly

Clade A2 more recently increasing
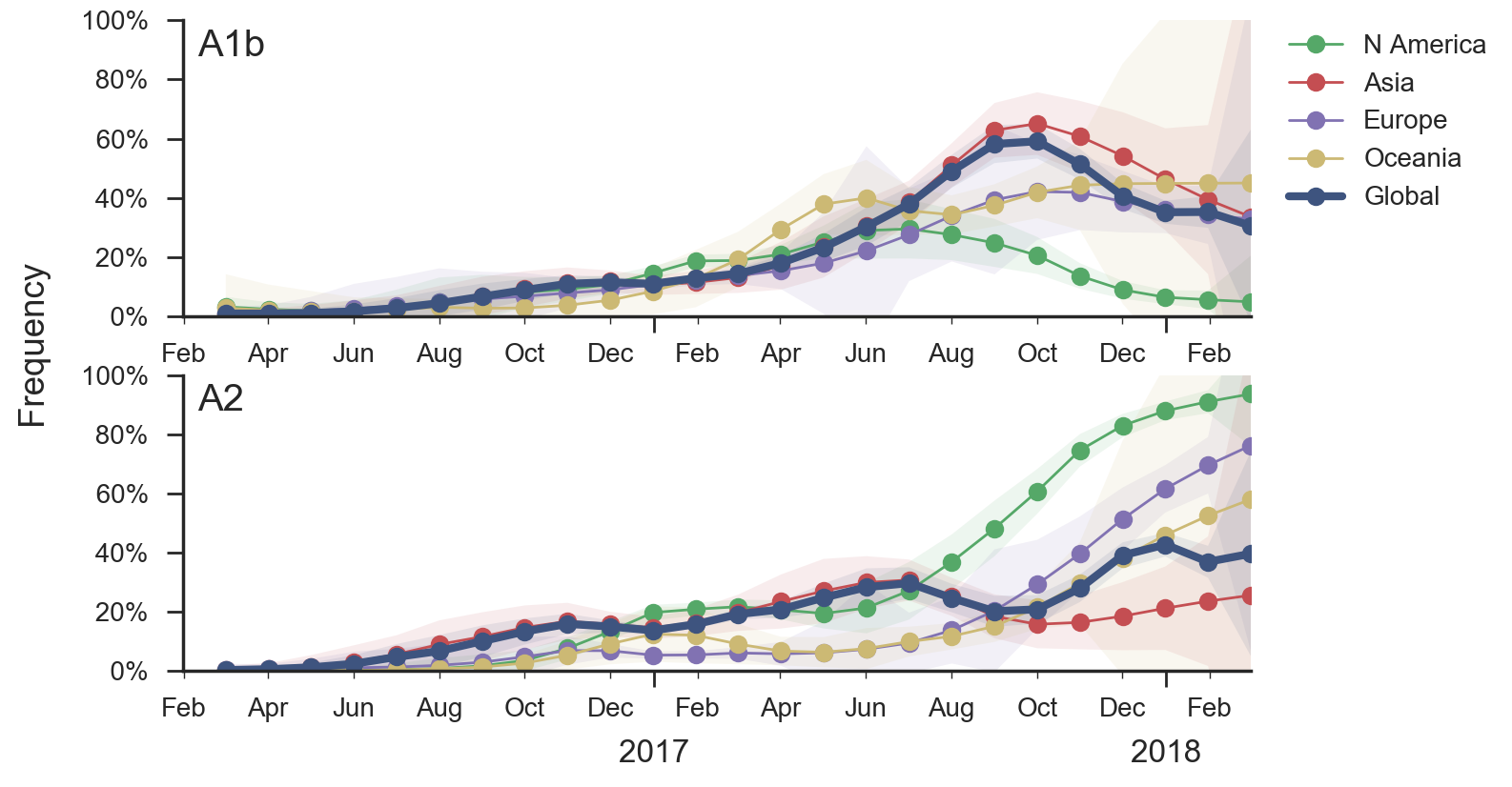
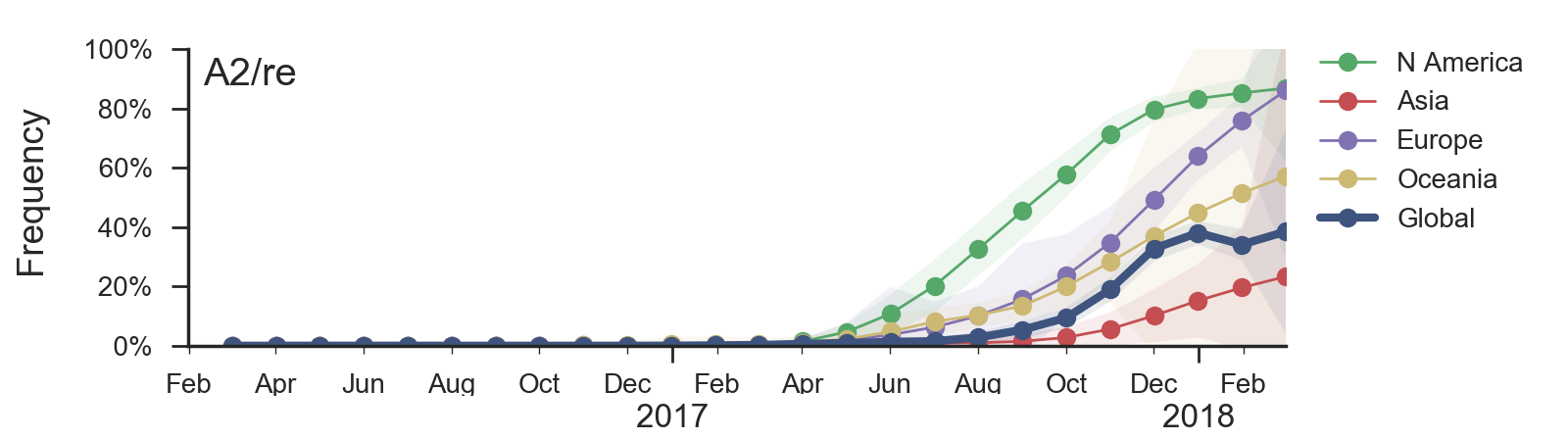
Reassortment event appears to drive success of clade A2

Reassortment event appears to drive success of clade A2
Further methods to integrate serological data and forecast future strain turnover
Ebola
Virus genomes reveal factors that spread and sustained the Ebola epidemic
with ![]() Gytis Dudas, Andrew Rambaut, Luiz Carvalho, Marc Suchard, Philippe Lemey,
Gytis Dudas, Andrew Rambaut, Luiz Carvalho, Marc Suchard, Philippe Lemey,
and many others
Sequencing of 1610 Ebola virus genomes collected during the 2013-2016 West African epidemic
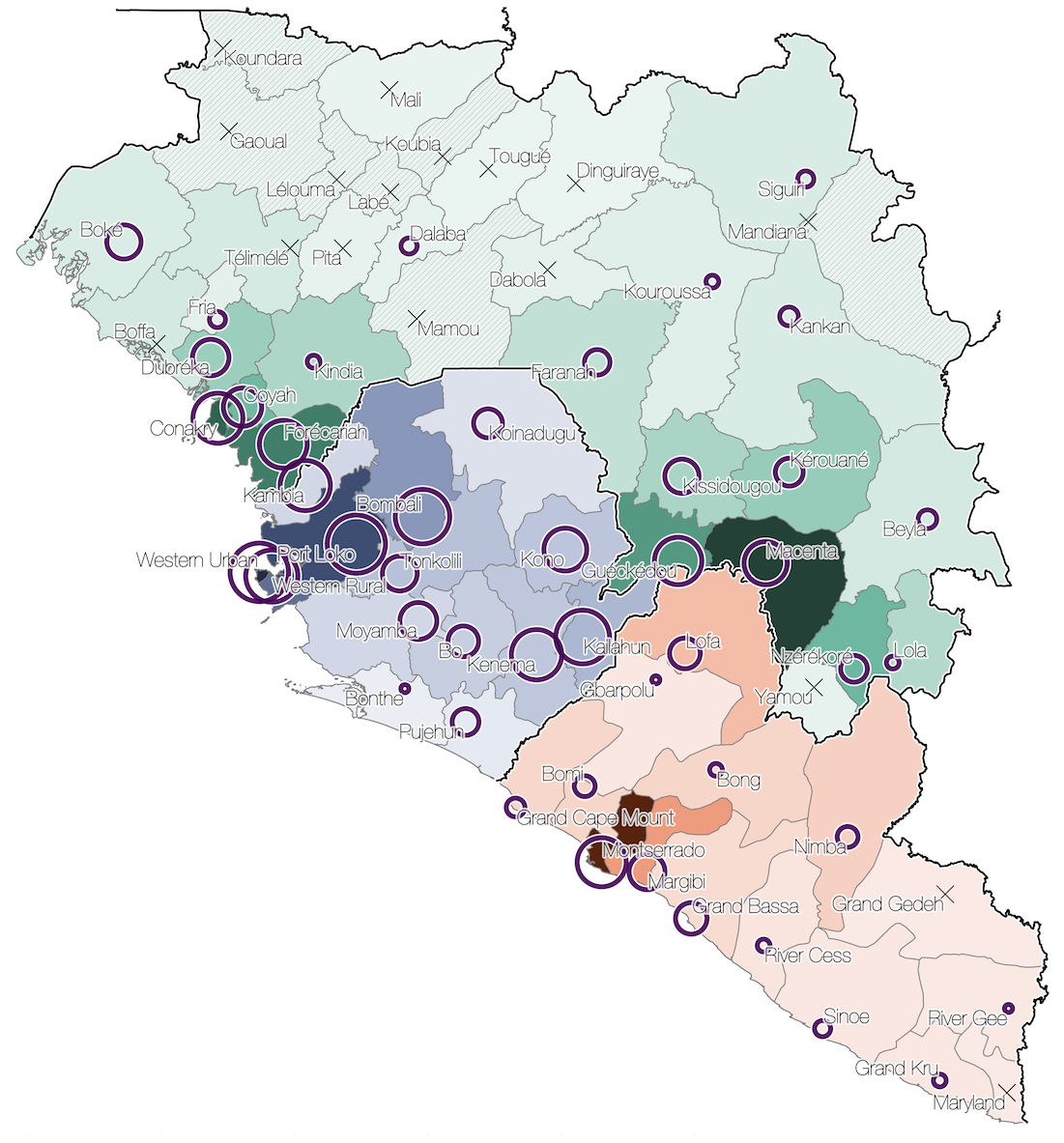
Sequenced genomes were representative of spatiotemporal diversity

Phylogenetic reconstruction of epidemic
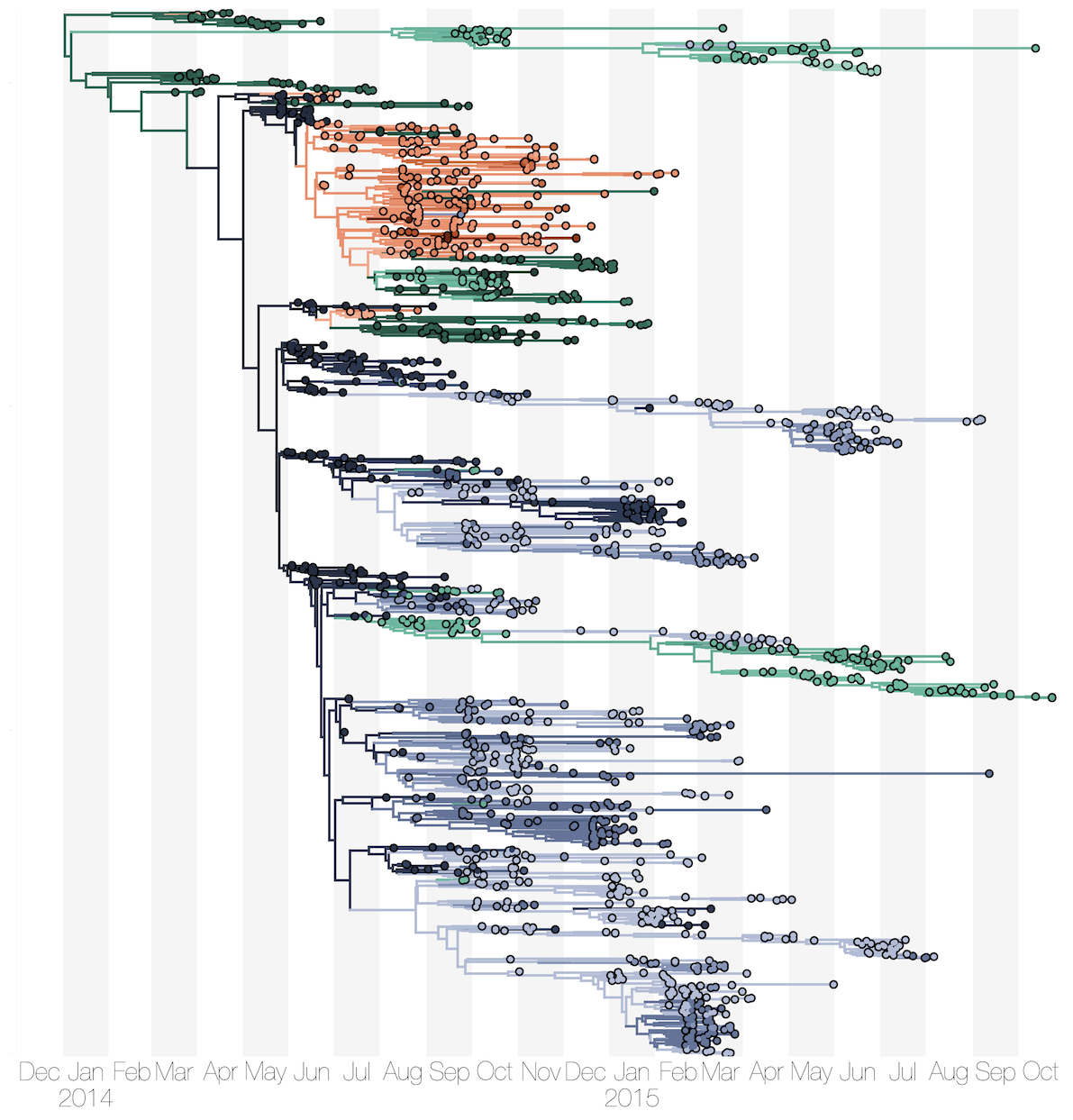
Tracking migration events
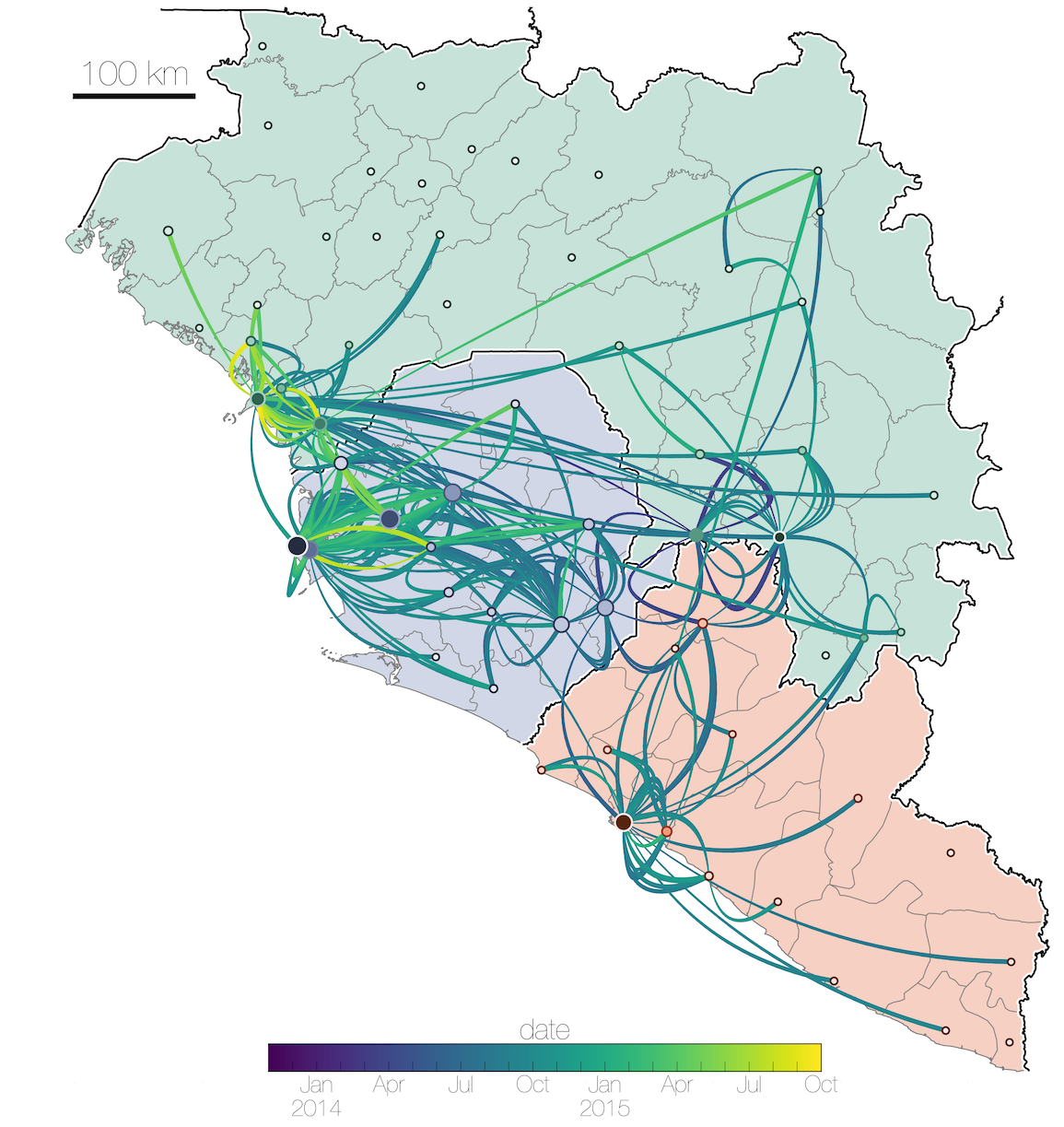
Factors influencing migration rates
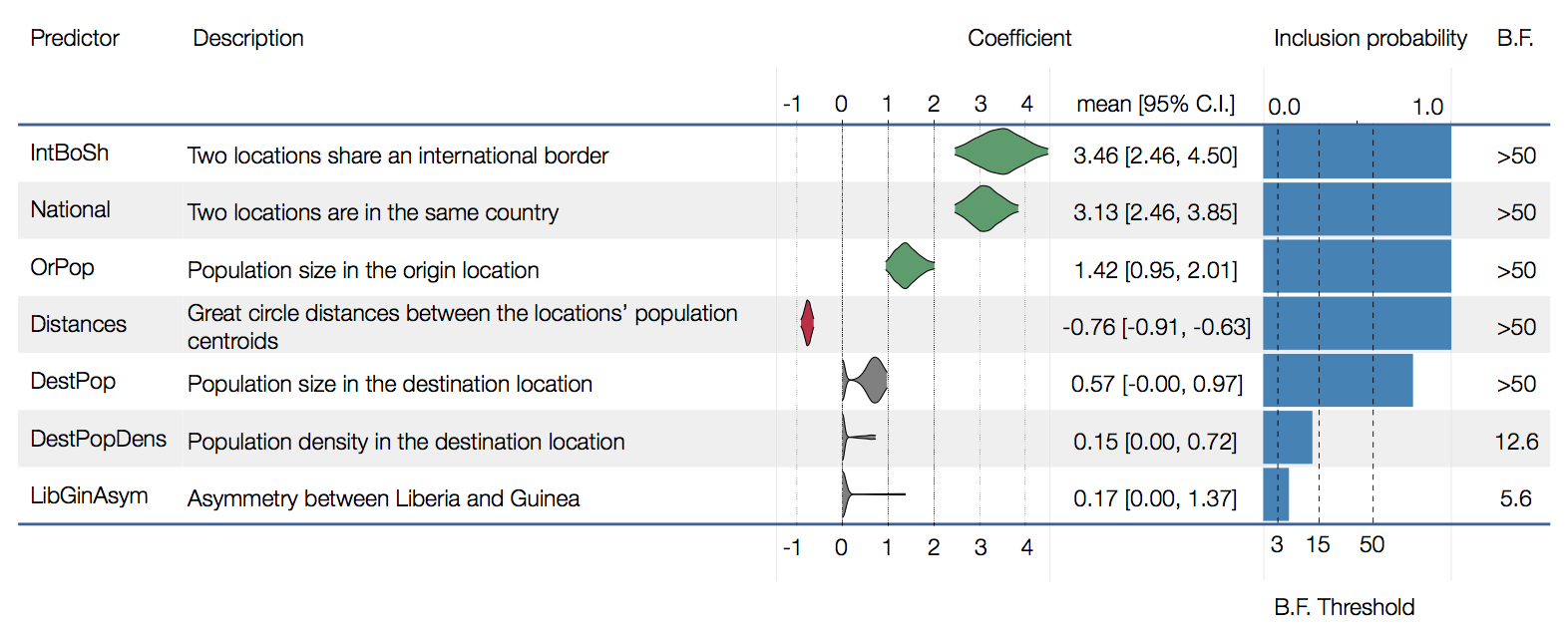
Effect of borders on migration rates
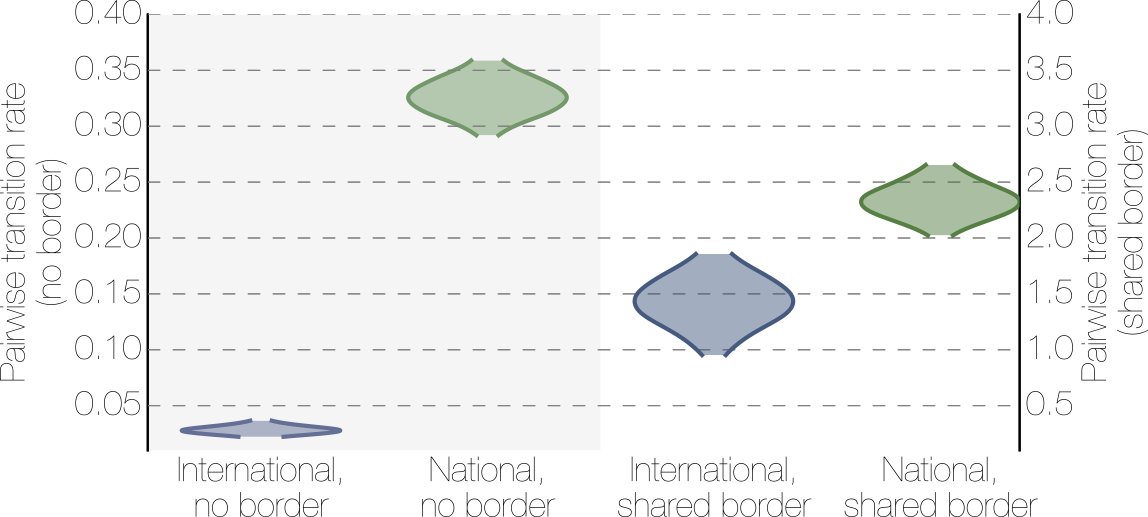
Spatial structure at the country level
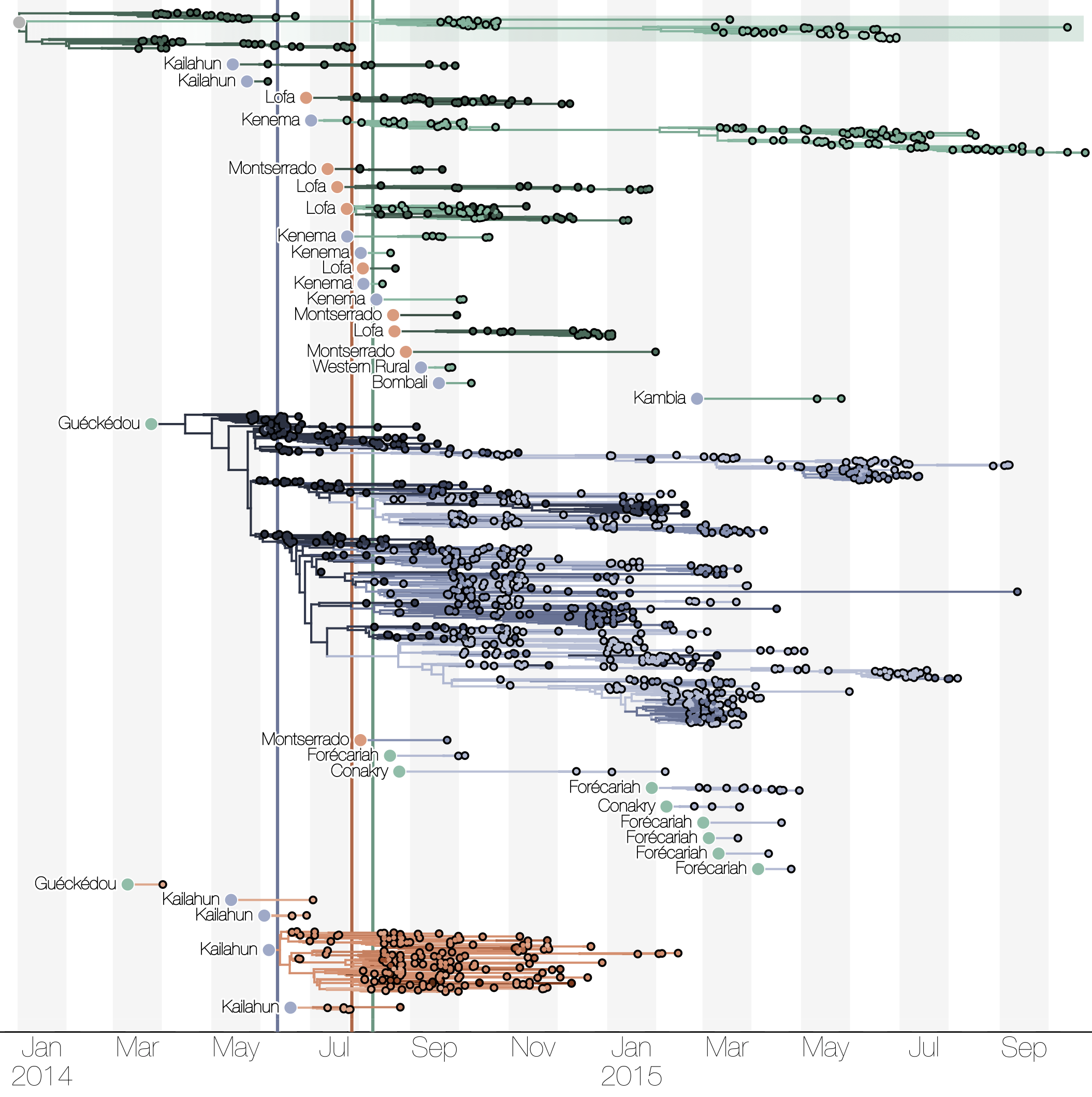
Substantial mixing at the regional level
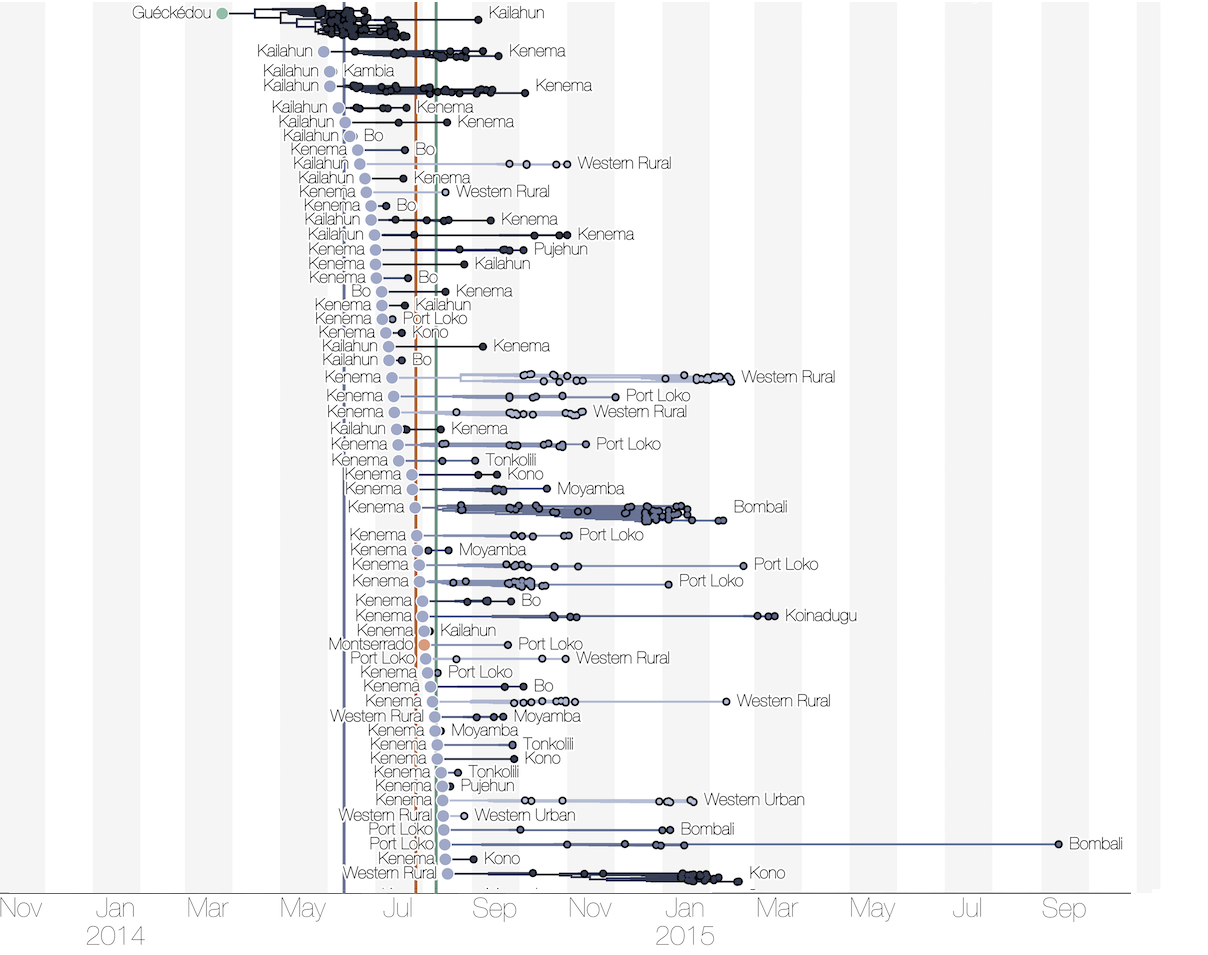
Regional outbreaks due to multiple introductions
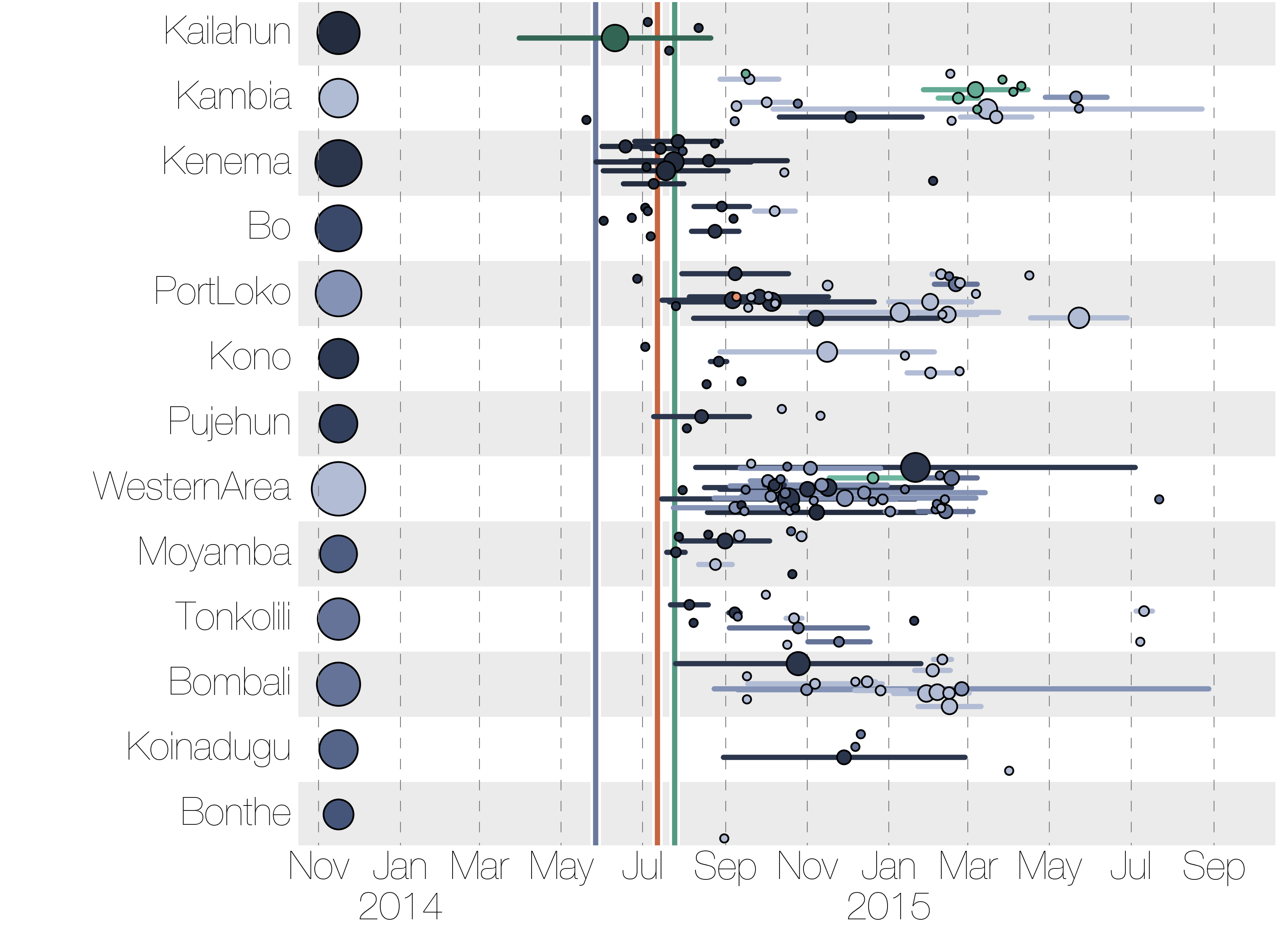
Each introduction results in a minor outbreak
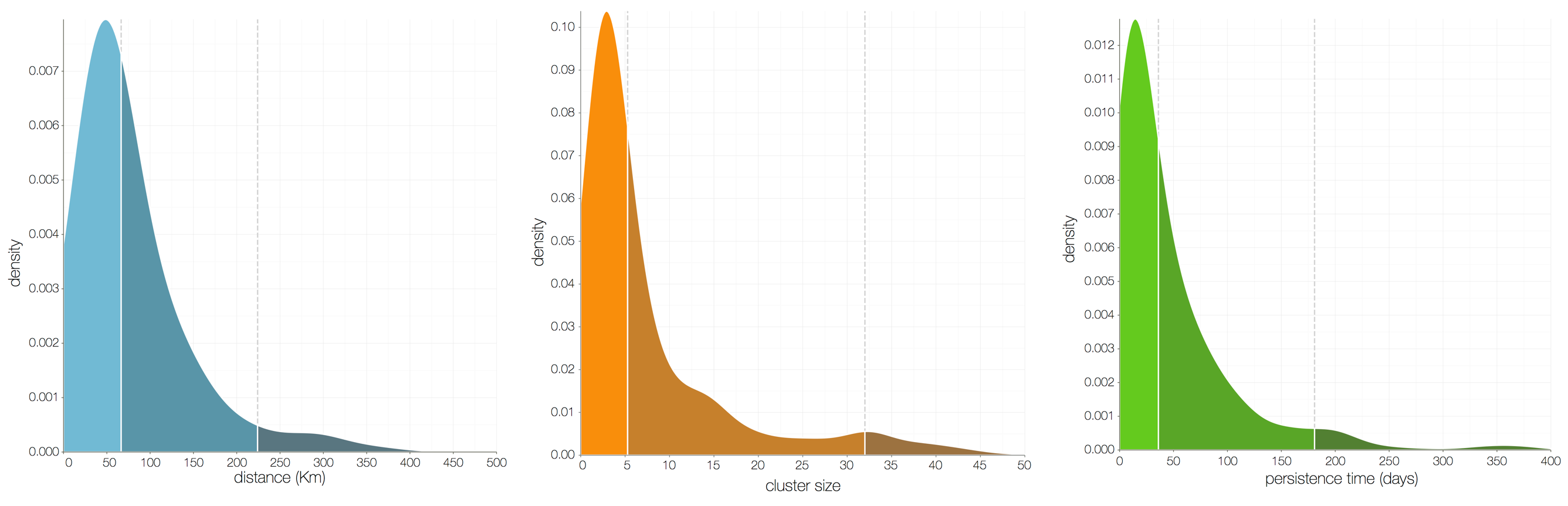
Zika
Zika's arrival and spread in the Americas
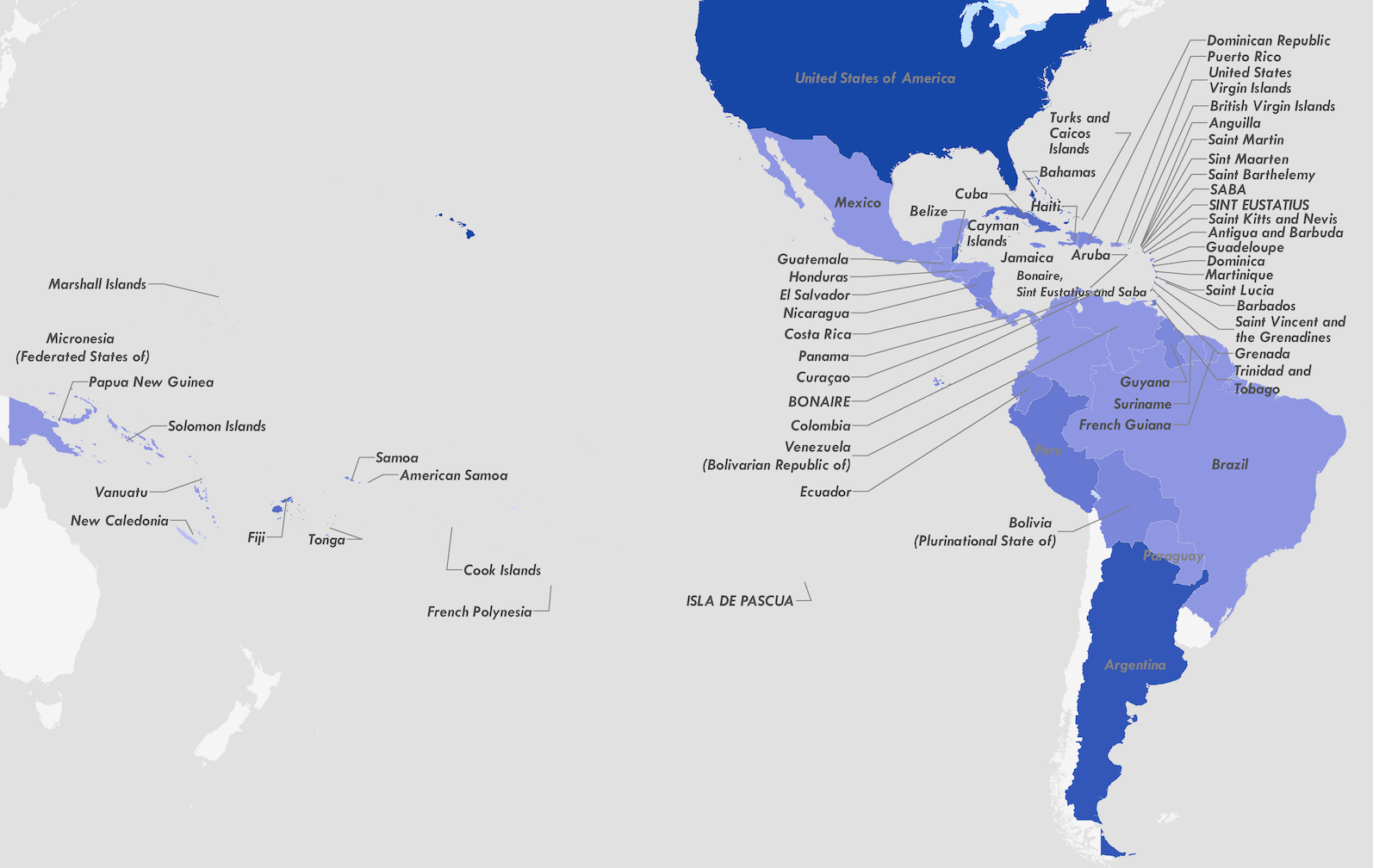
Establishment and cryptic transmission of Zika virus in Brazil and the Americas
with Nuno Faria, Nick Loman, Oli Pybus, Luiz Alcantara, Ester Sabino, Josh Quick,
![]() Alli Black,
Ingra Morales, Julien Thézé, Marcio Nunes, Jacqueline de Jesus,
Alli Black,
Ingra Morales, Julien Thézé, Marcio Nunes, Jacqueline de Jesus,
Marta Giovanetti, Moritz Kraemer, Sarah Hill and many others

Road trip through northeast Brazil to collect samples and sequence
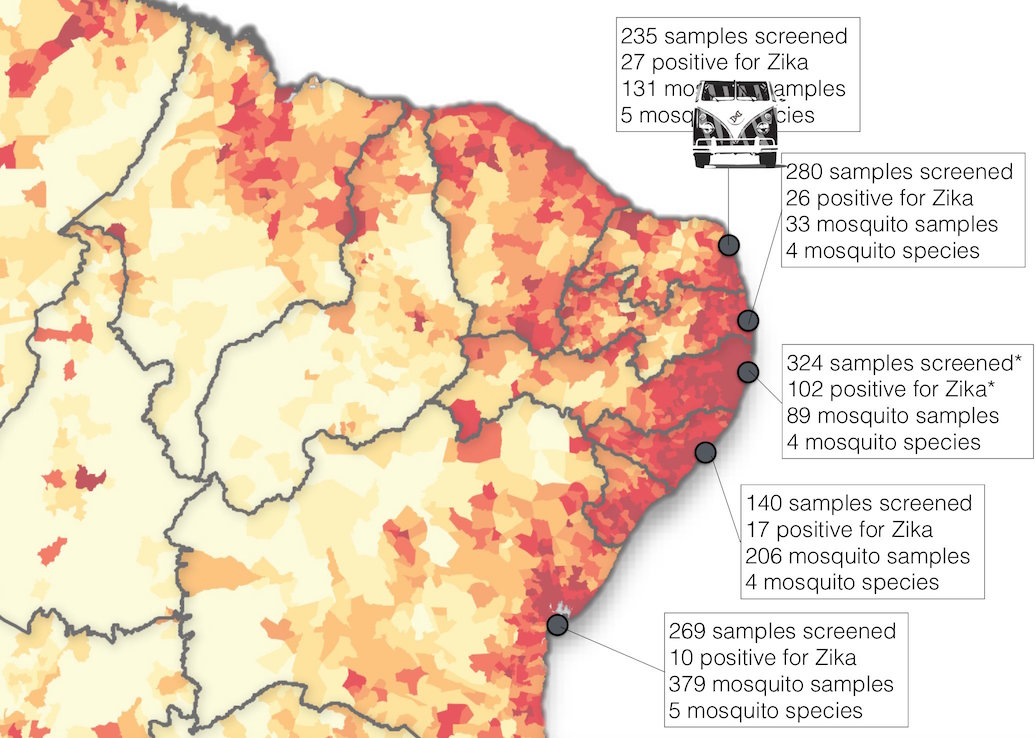
Case reports and diagnostics suggest initiation in northeast Brazil
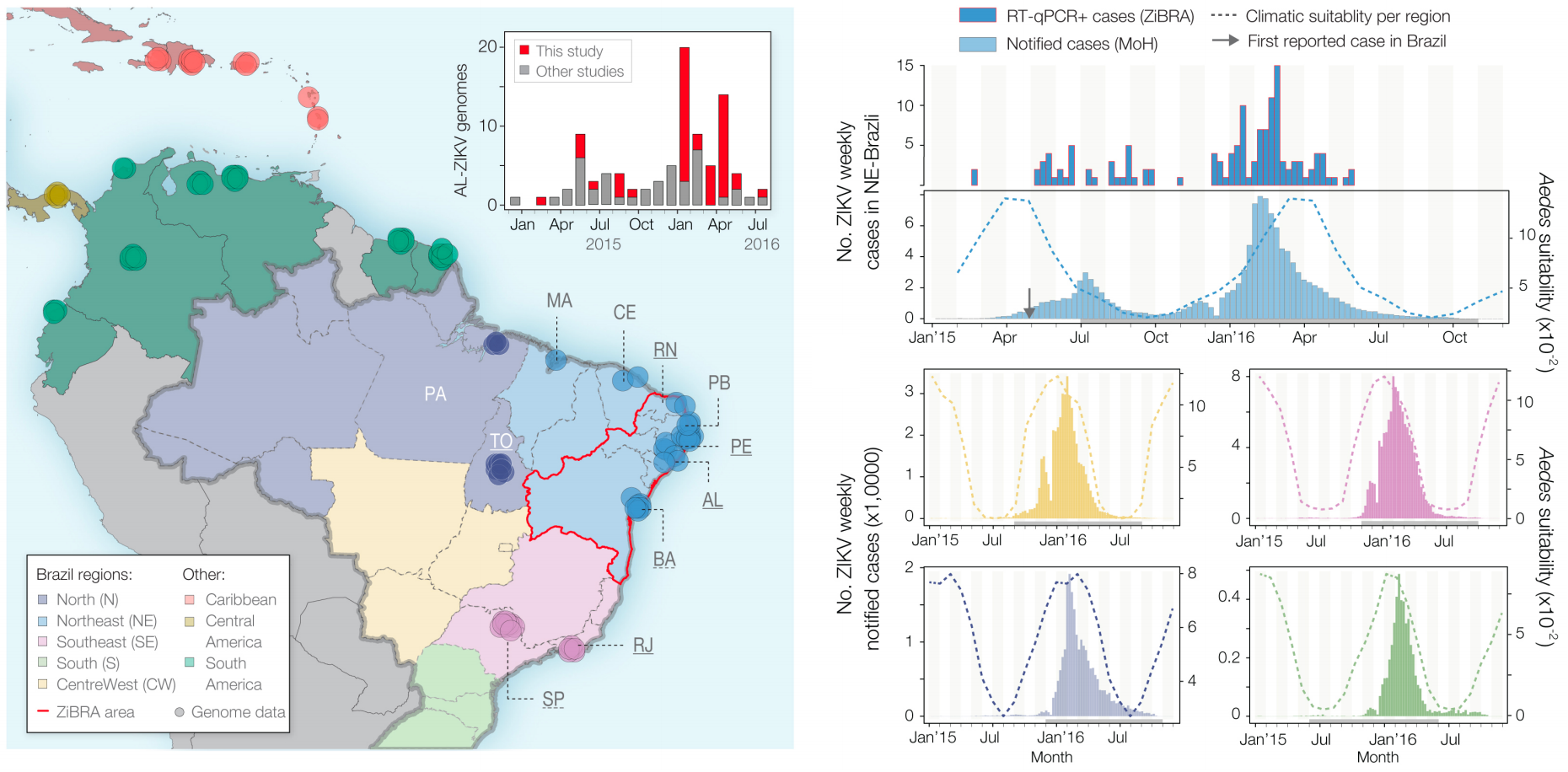
Phylogeny infers an origin in northeast Brazil
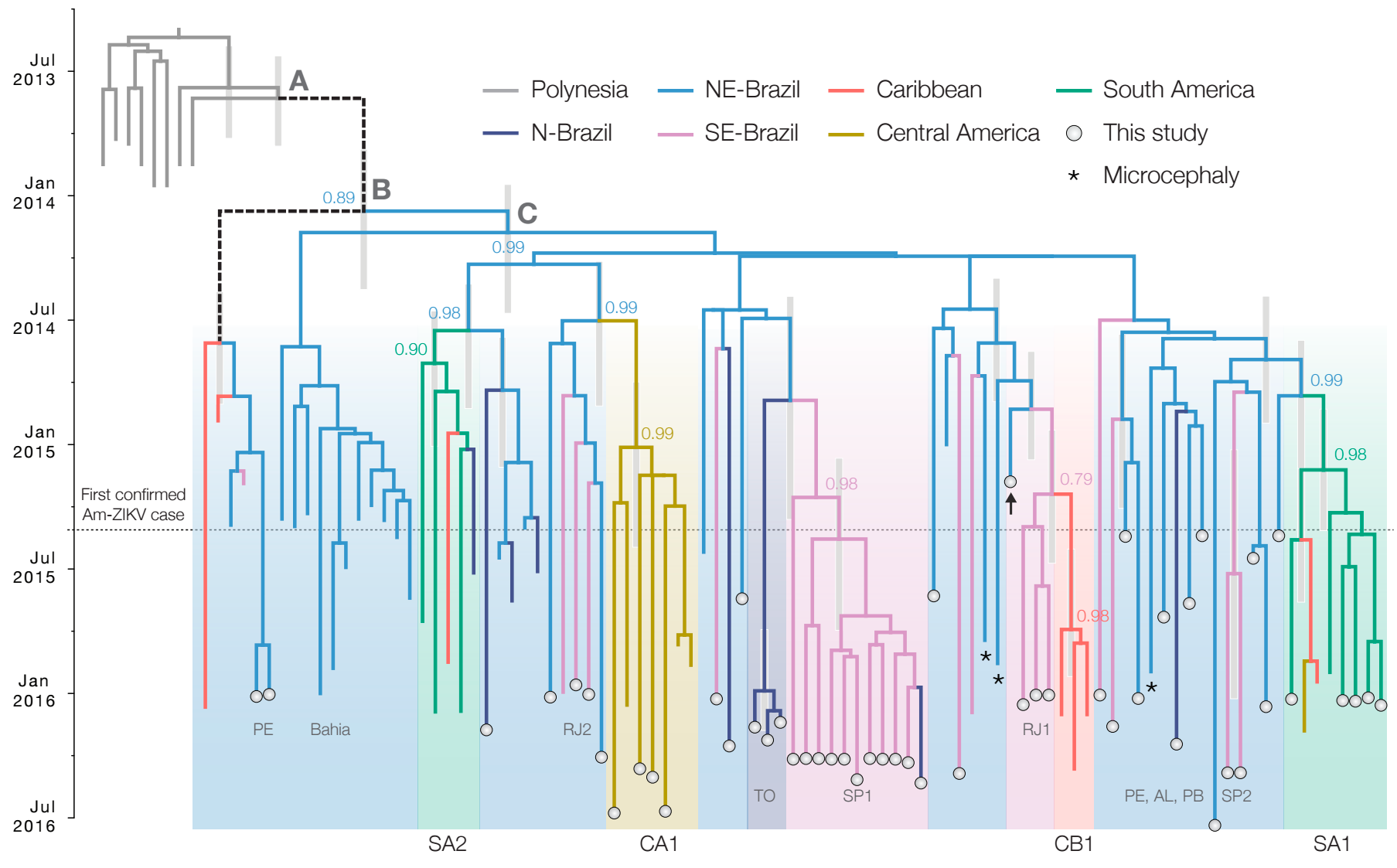
Genomic epidemiology reveals multiple introductions of Zika virus into the United States
with Nathan Grubaugh, Kristian Andersen, Jason Ladner, Gustavo Palacios, Sharon Isern, Oli Pybus,
Moritz Kraemer, ![]() Gytis Dudas,
Amanda Tan, Karthik Gangavarapu, Michael Wiley, Stephen White,
Julien Thézé, Scott Michael, Leah Gillis, Pardis Sabeti, and many others
Gytis Dudas,
Amanda Tan, Karthik Gangavarapu, Michael Wiley, Stephen White,
Julien Thézé, Scott Michael, Leah Gillis, Pardis Sabeti, and many others
Outbreak of locally-acquired infections focused in Miami-Dade county
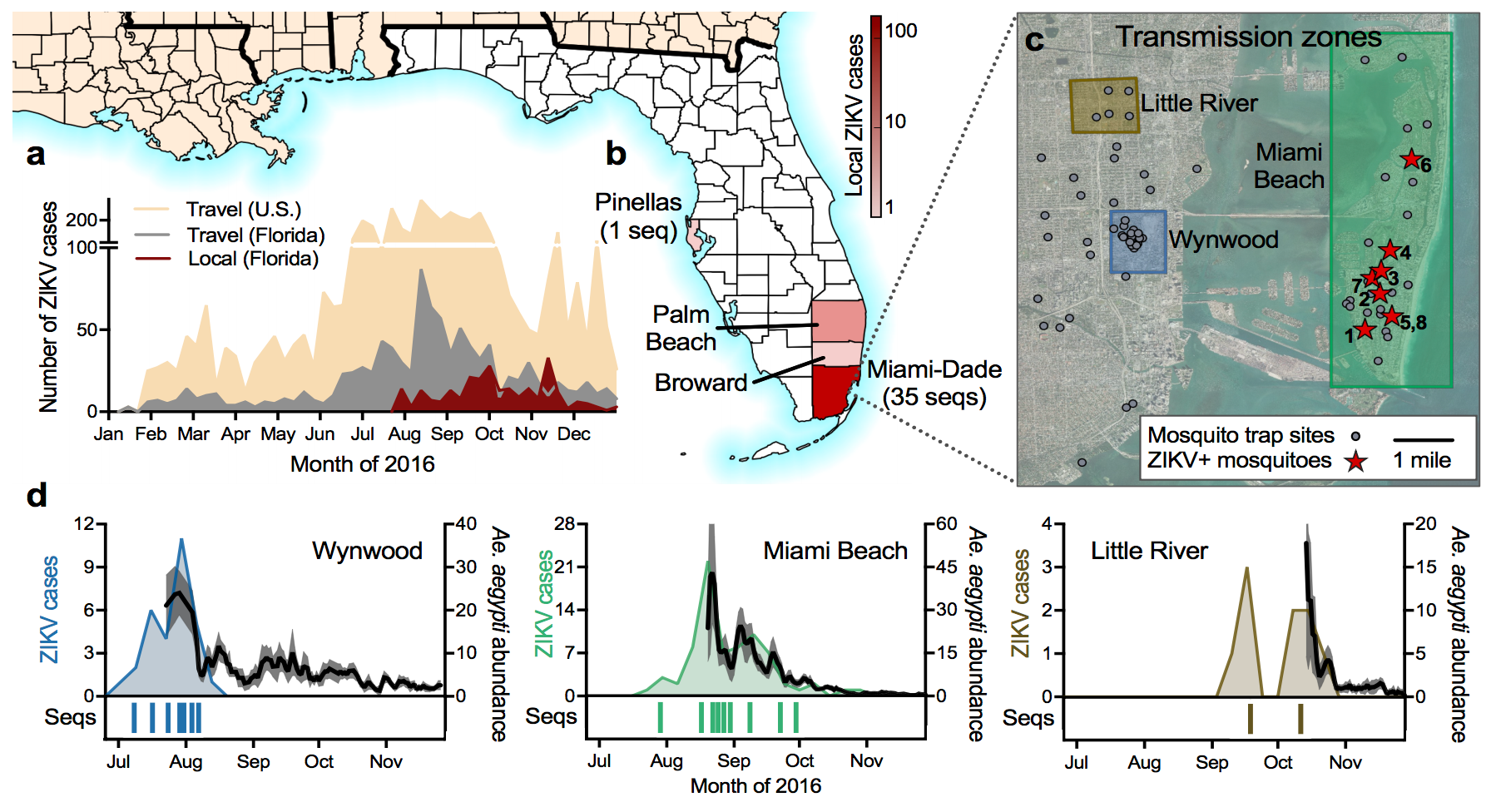
Phylogeny shows introductions from the Caribbean and a surprising degree of clustering
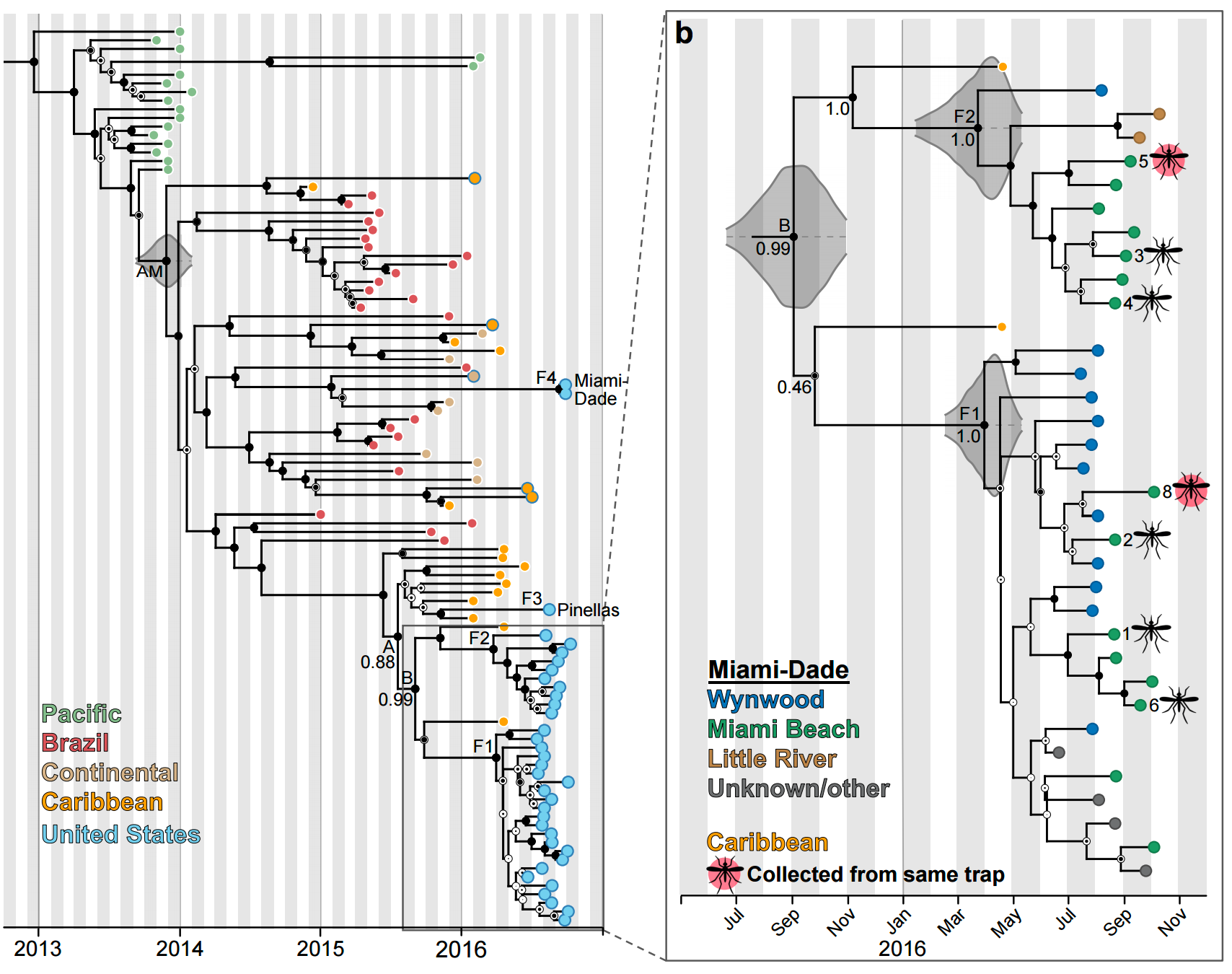
Flow of infected travelers greatest from Caribbean
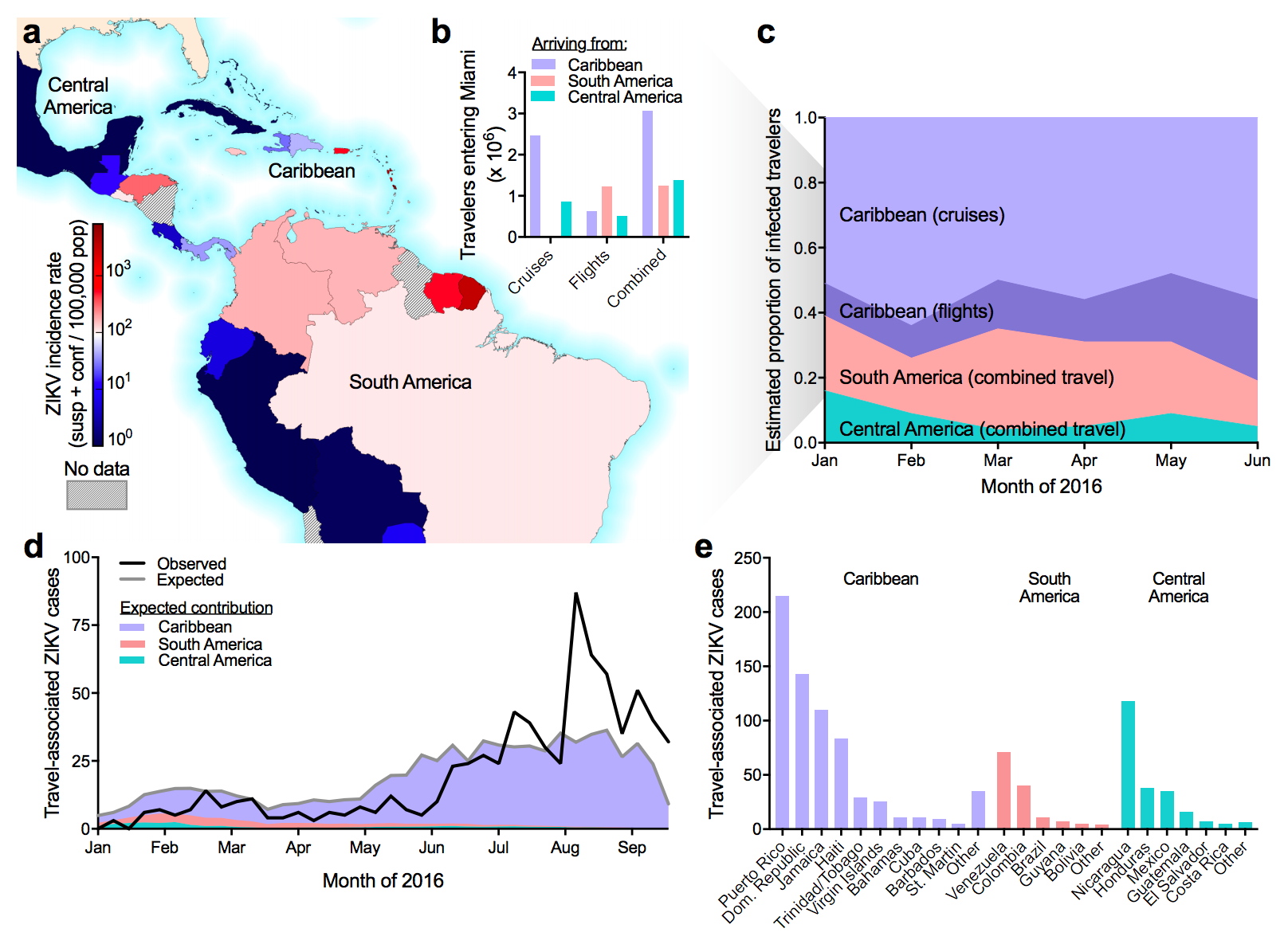
Clustering suggests fewer, longer transmission chains and higher R0
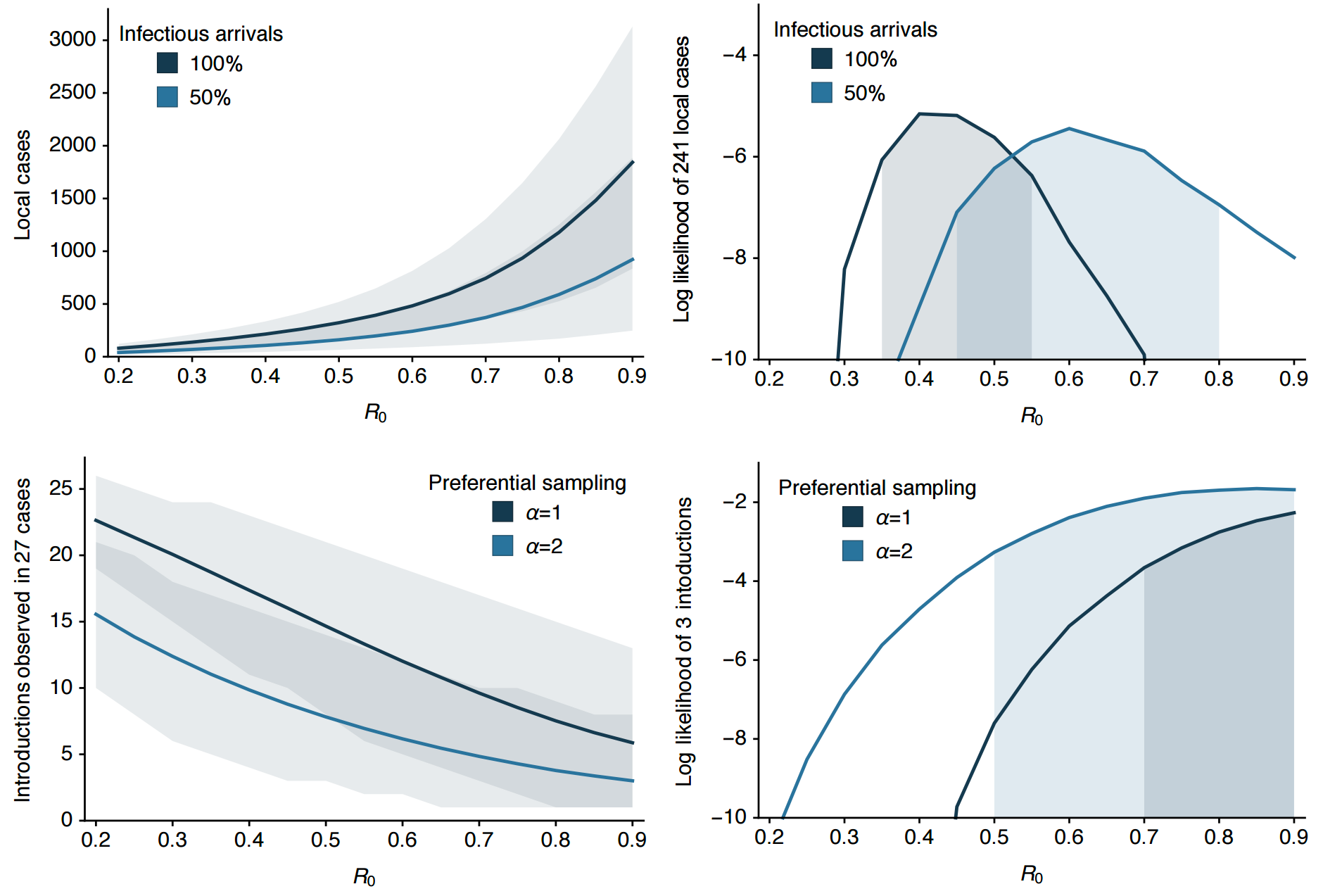
Genomic epidemiology of Zika in the US Virgin Islands
with ![]() Alli Black,
Alli Black,
![]() Barney Potter,
Barney Potter,
![]() Gytis Dudas,
Esther Ellis, Brett Ellis,
Gytis Dudas,
Esther Ellis, Brett Ellis,
Kristian Andersen, Nathan Grubaugh, Leora Feldstein, and others
(and special thanks to Adam Geballe)
Preliminary analysis of 31 genomes shows multiple introductions to USVI
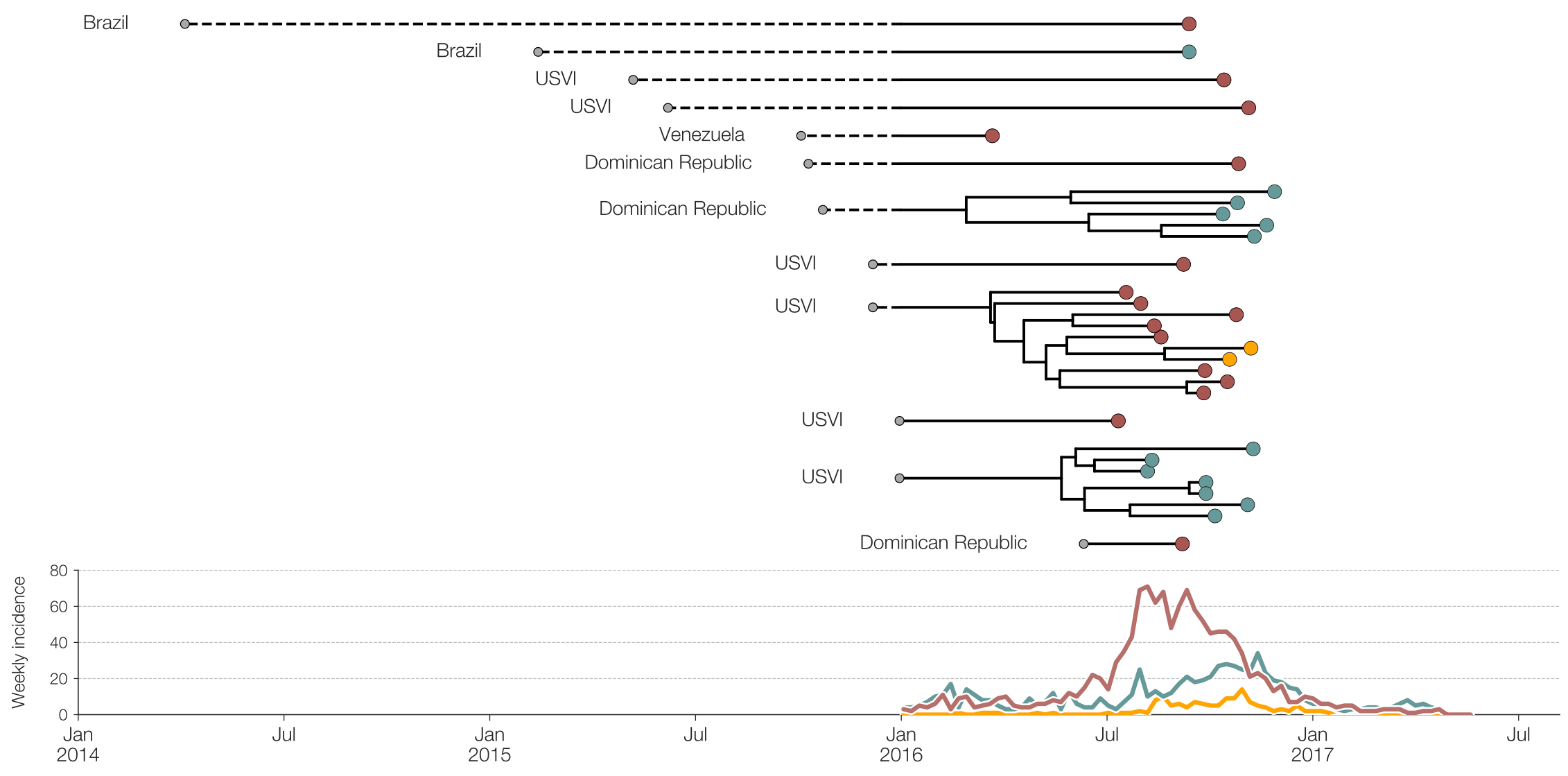
Actionable inferences
Genomic analyses were mostly done in a retrospective manner

Key challenges to making genomic epidemiology actionable
- Timely analysis and sharing of results critical
- Dissemination must be scalable
- Integrate many data sources
- Results must be easily interpretable and queryable
Nextstrain
Project to conduct real-time molecular epidemiology and evolutionary analysis of emerging epidemics
with
![]() Richard Neher,
Richard Neher,
![]() James Hadfield,
James Hadfield,
![]() Colin Megill,
Colin Megill,
![]() Sidney Bell,
Sidney Bell,
![]() John Huddleston,
John Huddleston,
![]() Barney Potter,
Barney Potter,
![]() Charlton Callender,
Charlton Callender,
![]() Emma Hodcroft
Emma Hodcroft
Nextstrain architecture
All code open source at github.com/nextstrain
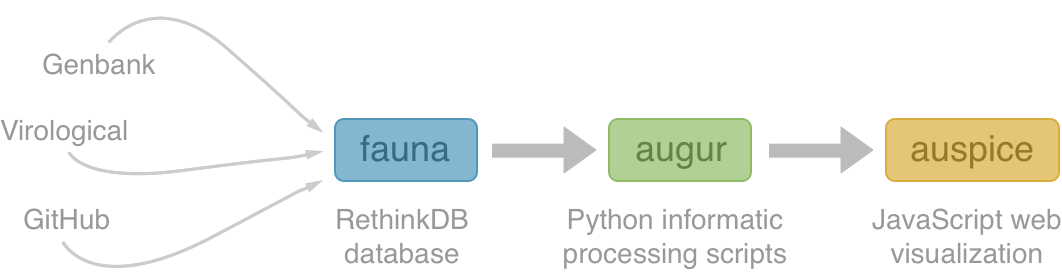
Fauna
Rethink database of virus and titer data
- Harmonizes data from different sources
- Integrates different types of data (serology, sequences, case details)
- Provides an interface for downstream analysis
Augur
Build scripts to align sequences, build trees and annotate
- Flexible build scripts to incorporate different viruses and analyses
- Constructs time-resolved phylogenies
- Annotates with geographic transitions and mutation events
Example augur pipeline for 1600 Ebola genomes
- Align with MAFFT (34 min)
- Build ML tree with RAxML (54 min)
- Temporally resolve tree and geographic ancestry with TreeTime (16 min)
- Total pipeline (1 hr 46 min)
Auspice
Web visualization of resulting trees
- Interactive data exploration and filtering
- Framework through React / D3
- Connects phylogeny, geography and genotypes
nextstrain.org
Rapid on-the-ground sequencing by Ian Goodfellow, Matt Cotten and colleagues
Build out pipelines for different pathogens,
improve databasing and lower
bioinformatics bar
Acknowledgements
Bedford Lab:
 Alli Black,
Alli Black,
 Sidney Bell,
Sidney Bell,
 Gytis Dudas,
Gytis Dudas,
 John Huddleston,
John Huddleston,
 Barney Potter,
Barney Potter,
 James Hadfield,
James Hadfield,
 Louise Moncla
Louise Moncla
Influenza: WHO Global Influenza Surveillance Network, GISAID, Richard Neher, Barney Potter, John Huddleston, Dave Wentworth, Becky Garten, Jackie Katz, Vivien Dugan, Xiyan Xu, Elizabeth Neuhaus, Sujatha Seenu
Ebola: Gytis Dudas, Andrew Rambaut, Luiz Carvalho, Philippe Lemey, Marc Suchard, Andrew Tatem
Zika: Nick Loman, Nuno Faria, Oli Pybus, Josh Quick, Kristian Andersen, Nathan Grubaugh, Alli Black, Jason Ladner, Gustavo Palacios, Sharon Isern, Gytis Dudas, Barney Potter, Esther Ellis
Nextstrain: Richard Neher, James Hadfield, Colin Megill, Sidney Bell, Charlton Callender, Barney Potter, John Huddleston, Emma Hodcroft






